Abstract
Purpose
As the premalignant lesion of human esophageal adenocarcinoma (EAC), Barrett’s esophagus (BE) is characterized by intestinal metaplasia in the normal esophagus (NE). Gene expression profiling with microarray and serial analysis of gene expression (SAGE) may help us understand the potential molecular mechanism of human BE.
Methods
We analyzed three microarray datasets (two cDNA arrays and one oligonucleotide array) and one SAGE dataset with statistical tools, significance analysis of microarrays (SAM) and SAGE(Poisson), to identify individual genes differentially expressed in BE. Gene set enrichment analysis (GSEA) was used to identify a priori defined sets of genes that were differentially expressed. These gene sets were grouped according to either certain signaling pathways (GSEA curated), or the presence of consensus binding sequences of known transcription factors (GSEA motif). Immunohistochemical staining (IHC) was used to validate differential gene expression.
Results
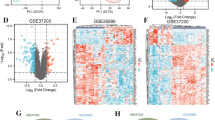
Both SAM and SAGE(Poisson) identified 68 differentially expressed genes (55 BE genes and 13 NE genes) with an arbitrary cutoff ratio (≥4-fold). With IHC on matched pairs of NE and BE tissues from 6 patients, these genes were grouped into 6 categories: category I (25 genes only expressed in BE), category II (5 genes only expressed in NE), category III (8 genes expressed more in BE than in NE), and category IV (2 genes expressed more in NE than in BE). Differential expression of the remaining genes was not confirmed by IHC either due to false discovery (category V), or lack of proper antibodies (category VI). Besides individual genes, the TGFβ pathway and several transcription factors (CDX2, HNF1, and HNF4) were identified by GSEA as enriched pathways and motifs in BE. Apart from 9 target genes known to be up-regulated in BE, IHC staining confirmed up-regulation of 19 additional CDX1 and CDX2 target genes in BE.
Conclusion
Our data suggested an important role of CDX1 and CDX2 in the development of BE. The IHC-confirmed gene list will lead to future studies on the molecular mechanism of BE.




Similar content being viewed by others
Abbreviations
- BE:
-
Barrett’s esophagus
- EAC:
-
Esophageal adenocarcinoma
- FDR:
-
False discovery rate
- GenMapp:
-
Gene Map Annotator and Pathway Profiler
- GO:
-
Gene ontology
- GSEA:
-
Gene set enrichment analysis
- IHC:
-
Immunohistochemical staining
- IM:
-
Intestinal metaplasia
- NE:
-
Normal esophagus
- SAGE:
-
Serial analysis of gene expression
- SAM:
-
Significance analysis of microarrays
References
Chen X, Yang CS. Esophageal adenocarcinoma: a review and perspectives on the mechanism of carcinogenesis and chemoprevention. Carcinogenesis. 2001;22:1119–29.
Jankowski JA, Harrison RF, Perry I, Balkwill F, Tselepis C. Barrett’s metaplasia. Lancet. 2000;356:2079–85.
Fitzgerald RC. Barrett’s oesophagus and oesophageal adenocarcinoma: how does acid interfere with cell proliferation and differentiation? Gut. 2005;54(Suppl 1):i21–6.
Krishnadath KK. Novel findings in the pathogenesis of esophageal columnar metaplasia or Barrett’s esophagus. Curr Opin Gastroenterol. 2007;23:440–5.
van Baal JW, Krishnadath KK. High throughput techniques for characterizing the expression profile of Barrett’s esophagus. Dis Esophagus. 2008;21:634–40.
Barrett MT, Yeung KY, Ruzzo WL, Hsu L, Blount PL, Sullivan R, et al. Transcriptional analyses of Barrett’s metaplasia and normal upper GI mucosae. Neoplasia. 2002;4:121–8.
Fox CA, Sapinoso LM, Zhang H, Zhang W, McLeod HL, Petroni GR, et al. Altered expression of TFF-1 and CES-2 in Barrett’s esophagus and associated adenocarcinomas. Neoplasia. 2005;7:407–16.
Gomes LI, Esteves GH, Carvalho AF, Cristo EB, Hirata R Jr, Martins WK, et al. Expression profile of malignant and nonmalignant lesions of esophagus and stomach: differential activity of functional modules related to inflammation and lipid metabolism. Cancer Res. 2005;65:7127–36.
Greenawalt DM, Duong C, Smyth GK, Ciavarella ML, Thompson NJ, Tiang T, et al. Gene expression profiling of esophageal cancer: comparative analysis of Barrett’s esophagus, adenocarcinoma, and squamous cell carcinoma. Int J Cancer. 2007;120:1914–21.
Hao Y, Triadafilopoulos G, Sahbaie P, Young HS, Omary MB, Lowe AW. Gene expression profiling reveals stromal genes expressed in common between Barrett’s esophagus and adenocarcinoma. Gastroenterology. 2006;131:925–33.
Helm J, Enkemann SA, Coppola D, Barthel JS, Kelley ST, Yeatman TJ. Dedifferentiation precedes invasion in the progression from Barrett’s metaplasia to esophageal adenocarcinoma. Clin Cancer Res. 2005;11:2478–85.
Kimchi ET, Posner MC, Park JO, Darga TE, Kocherginsky M, Karrison T, et al. Progression of Barrett’s metaplasia to adenocarcinoma is associated with the suppression of the transcriptional programs of epidermal differentiation. Cancer Res. 2005;65:3146–54.
Ostrowski J, Mikula M, Karczmarski J, Rubel T, Wyrwicz LS, Bragoszewski P, et al. Molecular defense mechanisms of Barrett’s metaplasia estimated by an integrative genomics. J Mol Med. 2007;85:733–43.
Ostrowski J, Rubel T, Wyrwicz LS, Mikula M, Bielasik A, Butruk E, et al. Three clinical variants of gastroesophageal reflux disease form two distinct gene expression signatures. J Mol Med. 2006;84:872–82.
Pohler E, Craig AL, Cotton J, Lawrie L, Dillon JF, Ross P, et al. The Barrett’s antigen anterior gradient-2 silences the p53 transcriptional response to DNA damage. Mol Cell Proteomics. 2004;3:534–47.
Selaru FM, Zou T, Xu Y, Shustova V, Yin J, Mori Y, et al. Global gene expression profiling in Barrett’s esophagus and esophageal cancer: a comparative analysis using cDNA microarrays. Oncogene. 2002;21:475–8.
van Baal JW, Milano F, Rygiel AM, Bergman JJ, Rosmolen WD, van Deventer SJ, et al. A comparative analysis by SAGE of gene expression profiles of Barrett’s esophagus, normal squamous esophagus, and gastric cardia. Gastroenterology. 2005;129:1274–81.
Wang S, Zhan M, Yin J, Abraham JM, Mori Y, Sato F, et al. Transcriptional profiling suggests that Barrett’s metaplasia is an early intermediate stage in esophageal adenocarcinogenesis. Oncogene. 2006;25:3346–56.
Xu Y, Selaru FM, Yin J, Zou TT, Shustova V, Mori Y, et al. Artificial neural networks and gene filtering distinguish between global gene expression profiles of Barrett’s esophagus and esophageal cancer. Cancer Res. 2002;62:3493–7.
van Baal JW, Diks SH, Wanders RJ, Rygiel AM, Milano F, Joore J, et al. Comparison of kinome profiles of Barrett’s esophagus with normal squamous esophagus and normal gastric cardia. Cancer Res. 2006;66:11605–12.
Tusher VG, Tibshirani R, Chu G. Significance analysis of microarrays applied to the ionizing radiation response. Proc Natl Acad Sci USA. 2001;98:5116–21.
Cai L, Huang H, Blackshaw S, Liu JS, Cepko C, Wong WH. Clustering analysis of SAGE data using a Poisson approach. Genome Biol. 2004;5:R51.
Lewin K, Appelman H. Barrett’s esophagus, columnar dysplasia, and adenocarcinoma of the esophagus. In: Appelman KLaH, editor. Tumors of the Esophagus and Stomach. Washington: AFIP; 1995. p. 99–144.
Liu T, Zhang X, So CK, Wang S, Wang P, Yan L, et al. Regulation of Cdx2 expression by promoter methylation, and effects of Cdx2 transfection on morphology and gene expression of human esophageal epithelial cells. Carcinogenesis. 2007;28:488–96.
Wong NA, Wilding J, Bartlett S, Liu Y, Warren BF, Piris J, et al. CDX1 is an important molecular mediator of Barrett’s metaplasia. Proc Natl Acad Sci USA. 2005;102:7565–70.
Yang Q, Bermingham NA, Finegold MJ, Zoghbi HY. Requirement of Math1 for secretory cell lineage commitment in the mouse intestine. Science. 2001;294:2155–8.
Eda A, Osawa H, Satoh K, Yanaka I, Kihira K, Ishino Y, et al. Aberrant expression of CDX2 in Barrett’s epithelium and inflammatory esophageal mucosa. J Gastroenterol. 2003;38:14–22.
Silberg DG, Furth EE, Taylor JK, Schuck T, Chiou T, Traber PG. CDX1 protein expression in normal, metaplastic, and neoplastic human alimentary tract epithelium. Gastroenterology. 1997;113:478–86.
Chen X, Qin R, Liu B, Ma Y, Su Y, Yang CS, et al. Multilayered epithelium in a rat model and human Barrett’s esophagus: similar expression patterns of transcription factors and differentiation markers. BMC Gastroenterol. 2008;8:1.
Milano F, van Baal JW, Buttar NS, Rygiel AM, de Kort F, DeMars CJ, et al. Bone morphogenetic protein 4 expressed in esophagitis induces a columnar phenotype in esophageal squamous cells. Gastroenterology. 2007;132:2412–21.
Slack JM, Tosh D. Transdifferentiation and metaplasia—switching cell types. Curr Opin Genet Dev. 2001;11:581–6.
Brabender J, Marjoram P, Salonga D, Metzger R, Schneider PM, Park JM, et al. A multigene expression panel for the molecular diagnosis of Barrett’s esophagus and Barrett’s adenocarcinoma of the esophagus. Oncogene. 2004;23:4780–8.
Glickman JN, Blount PL, Sanchez CA, Cowan DS, Wongsurawat VJ, Reid BJ, et al. Mucin core polypeptide expression in the progression of neoplasia in Barrett’s esophagus. Hum Pathol. 2006;37:1304–15.
Jovov B, Van Itallie CM, Shaheen NJ, Carson JL, Gambling TM, Anderson JM, et al. Claudin-18: a dominant tight junction protein in Barrett’s esophagus and likely contributor to its acid resistance. Am J Physiol Gastrointest Liver Physiol. 2007;293:G1106–13.
Kumble S, Omary MB, Fajardo LF, Triadafilopoulos G. Multifocal heterogeneity in villin and Ep-CAM expression in Barrett’s esophagus. Int J Cancer. 1996;66:48–54.
Madsen J, Nielsen O, Tornoe I, Thim L, Holmskov U. Tissue localization of human trefoil factors 1, 2, and 3. J Histochem Cytochem. 2007;55:505–13.
Mitas M, Almeida JS, Mikhitarian K, Gillanders WE, Lewin DN, Spyropoulos DD, et al. Accurate discrimination of Barrett’s esophagus and esophageal adenocarcinoma using a quantitative three-tiered algorithm and multimarker real-time reverse transcription-PCR. Clin Cancer Res. 2005;11:2205–14.
van Baal JW, Bozikas A, Pronk R, Ten Kate FJ, Milano F, Rygiel AM, et al. Cytokeratin and CDX-2 expression in Barrett’s esophagus. Scand J Gastroenterol. 2008;43:132–40.
Wong NA, Warren BF, Piris J, Maynard N, Marshall R, Bodmer WF. EpCAM and gpA33 are markers of Barrett’s metaplasia. J Clin Pathol. 2006;59:260–3.
Christie KN, Thomson C. The distribution of carbonic anhydrase II in human, pig and rat oesophageal epithelium. Histochem J. 2000;32:753–7.
Xia SH, Hu LP, Hu H, Ying WT, Xu X, Cai Y, et al. Three isoforms of annexin I are preferentially expressed in normal esophageal epithelia but down-regulated in esophageal squamous cell carcinomas. Oncogene. 2002;21:6641–8.
Mobasheri A, Wray S, Marples D. Distribution of AQP2 and AQP3 water channels in human tissue microarrays. J Mol Histol. 2005;36:1–14.
Chu PG, Weiss LM. Keratin expression in human tissues and neoplasms. Histopathology. 2002;40:403–39.
Gerber JK, Richter T, Kremmer E, Adamski J, Hofler H, Balling R, et al. Progressive loss of PAX9 expression correlates with increasing malignancy of dysplastic and cancerous epithelium of the human oesophagus. J Pathol. 2002;197:293–7.
Ihrie RA, Marques MR, Nguyen BT, Horner JS, Papazoglu C, Bronson RT, et al. Perp is a p63-regulated gene essential for epithelial integrity. Cell. 2005;120:843–56.
South AP. Plakophilin 1: an important stabilizer of desmosomes. Clin Exp Dermatol. 2004;29:161–7.
Nishimori T, Tomonaga T, Matsushita K, Oh-Ishi M, Kodera Y, Maeda T, et al. Proteomic analysis of primary esophageal squamous cell carcinoma reveals downregulation of a cell adhesion protein, periplakin. Proteomics. 2006;6:1011–8.
Mori-Akiyama Y, van den Born M, van Es JH, Hamilton SR, Adams HP, Zhang J, et al. SOX9 is required for the differentiation of paneth cells in the intestinal epithelium. Gastroenterology. 2007;133:539–46.
Niimi T, Nagashima K, Ward JM, Minoo P, Zimonjic DB, Popescu NC. Claudin-18, a novel downstream target gene for the T/EBP/NKX2.1 homeodomain transcription factor, encodes lung- and stomach-specific isoforms through alternative splicing. Mol Cell Biol. 2001;21:7380–90.
Haveri H, Westerholm-Ormio M, Lindfors K, Maki M, Savilahti E, Andersson LC, et al. Transcription factors GATA-4 and GATA-6 in normal and neoplastic human gastrointestinal mucosa. BMC Gastroenterol. 2008;8:9.
Calon A, Gross I, Lhermitte B, Martin E, Beck F, Duclos B, et al. Different effects of the Cdx1 and Cdx2 homeobox genes in a murine model of intestinal inflammation. Gut. 2007;56:1688–95.
Barros R, Pereira B, Duluc I, Azevedo M, Mendes N, Camilo V, et al. Key elements of the BMP/SMAD pathway co-localize with CDX2 in intestinal metaplasia and regulate CDX2 expression in human gastric cell lines. J Pathol. 2008.
Acknowledgments
We thank Dr. Li Cai, Department of Biomedical Engineering, Rutgers University, Piscataway, NJ, in helping in SAGE(Poisson) analysis; Dr. Vasilis Vasiliou, Molecular Toxicology and Environmental Health Sciences Program, Department of Pharmaceutical Sciences, University of Colorado Health Sciences Center, Denver, CO, for providing the anti-ALDH3A1 used in this study; Drs. Danielle M. Greenawalt and Wayne A. Phillips, Centre for Cancer Genomics and Predictive Medicine, Peter MacCallum Cancer Centre, Victoria, Australia, for providing original data and information regarding pathology; Dr. Jerzy Ostrowski, Department of Gastroenterology, Medical Center for Postgraduate Education, Maria Sklodowska-Curie Memorial Cancer Center and Institute of Oncology, Warsaw, Poland, for providing original data and information regarding pathology; Dr. William R. Otto, Histopathology Unit, London Research Institute, Cancer Research UK, London, UK, for staining our tissue sections for GDDR. This study was supported by NIH grants U56 CA092077 and P20 MD000175.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Wang, J., Qin, R., Ma, Y. et al. Differential gene expression in normal esophagus and Barrett’s esophagus. J Gastroenterol 44, 897–911 (2009). https://doi.org/10.1007/s00535-009-0082-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00535-009-0082-2