Abstract
Key message
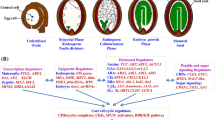
Overview of seed size control.
Human and livestock nutrition is largely based on calories derived from seeds, in particular cereals and legumes. Unveiling the control of seed size is therefore of remarkable importance in the frame of developing new strategies for crop improvement. The networks controlling the development of the seed coat, the endosperm and the embryo, as well as their interplay, have been described in Arabidopsis thaliana. In this review, we provide a comprehensive description of the current knowledge regarding the molecular mechanisms controlling seed size in Arabidopsis.



Similar content being viewed by others
References
Adams S, Vinkenoog R, Spielman M et al (2000) Parent-of-origin effects on seed development in Arabidopsis thaliana require DNA methylation. Development 127:2493–2502
Adamski N, Anastasiou E, Eriksson S et al (2009) Local maternal control of seed size by KLUH/CYP78A5-dependent growth signaling. Proc Natl Acad Sci USA 106:20115–20120. doi:10.1073/pnas.0907024106
Allen RS, Li J, Stahle MI et al (2007) Genetic analysis reveals functional redundancy and the major target genes of the Arabidopsis miR159 family. Proc Natl Acad Sci USA 104:16371–16376. doi:10.1073/pnas.0707653104
Anastasiou E, Kenz S, Gerstung M et al (2007) Control of plant organ size by KLUH/CYP78A5-dependent intercellular signaling. Dev Cell 13:843–856. doi:10.1016/j.devcel.2007.10.001
Appelhagen I, Thiedig K, Nordholt N et al (2014) Update on transparent testa mutants from Arabidopsis thaliana: characterisation of new alleles from an isogenic collection. Planta 240:955–970. doi:10.1007/s00425-014-2088-0
Aw SJ, Hamamura Y, Chen Z et al (2010) Sperm entry is sufficient to trigger division of the central cell but the paternal genome is required for endosperm development in Arabidopsis. Development 137:2683–2690. doi:10.1242/dev.052928
Bartrina I, Otto E, Strnad M et al (2011) Cytokinin regulates the activity of reproductive meristems, flower organ size, ovule formation, and thus seed yield in Arabidopsis thaliana. Plant Cell 23:69–80. doi:10.1105/tpc.110.079079
Bauer MJ, Fischer RL (2011) Genome demethylation and imprinting in the endosperm. Curr Opin Plant Biol 14:162–167. doi:10.1016/j.pbi.2011.02.006
Becker MG, Hsu S-W, Harada JJ, Belmonte MF (2014) Genomic dissection of the seed. Front Plant Sci 5:464. doi:10.3389/fpls.2014.00464
Beeckman T, De Rycke R, Viane R, Inzé D (2000) Histological study of seed coat development in Arabidopsis thaliana. J Plant Res 113:139–148. doi:10.1007/PL00013924
Bencivenga S, Colombo L, Masiero S (2011) Cross talk between the sporophyte and the megagametophyte during ovule development. Sex Plant Reprod 24:113–121. doi:10.1007/s00497-011-0162-3
Berger F, Grini PE, Schnittger A (2006) Endosperm: an integrator of seed growth and development. Curr Opin Plant Biol 9:664–670. doi:10.1016/j.pbi.2006.09.015
Bishopp A, Benková E, Helariutta Y (2011) Sending mixed messages: auxin-cytokinin crosstalk in roots. Curr Opin Plant Biol 14:10–16. doi:10.1016/j.pbi.2010.08.014
Boisnard-Lorig C, Colon-Carmona A, Bauch M et al (2001) Dynamic analyses of the expression of the HISTONE:YFP fusion protein in arabidopsis show that syncytial endosperm is divided in mitotic domains. Plant Cell 13:495–509
Brocard IM, Lynch TJ, Finkelstein RR (2002) Regulation and role of the Arabidopsis abscisic acid-insensitive 5 gene in abscisic acid, sugar, and stress response. Plant Physiol 129:1533–1543. doi:10.1104/pp.005793
Chen X (2004) A microRNA as a translational repressor of APETALA2 in Arabidopsis flower development. Science 303:2022–2025. doi:10.1126/science.1088060
Cheng Y, Dai X, Zhao Y (2007) Auxin synthesized by the YUCCA flavin monooxygenases is essential for embryogenesis and leaf formation in Arabidopsis. Plant Cell 19:2430–2439. doi:10.1105/tpc.107.053009
Cheng ZJ, Zhao XY, Shao XX et al (2014) Abscisic acid regulates early seed development in Arabidopsis by ABI5-mediated transcription of SHORT HYPOCOTYL UNDER BLUE1. Plant Cell 26:1053–1068. doi:10.1105/tpc.113.121566
Choe S, Tanaka A, Noguchi T et al (2000) Lesions in the sterol delta reductase gene of Arabidopsis cause dwarfism due to a block in brassinosteroid biosynthesis. Plant J 21:431–443
Choi Y, Gehring M, Johnson L et al (2002) DEMETER, a DNA glycosylase domain protein, is required for endosperm gene imprinting and seed viability in arabidopsis. Cell 110:33–42
Conlon I, Raff M (1999) Size control in animal development. Cell 96:235–244
Day SJ, Lawrence PA (2000) Measuring dimensions: the regulation of size and shape. Development 127:2977–2987
Day RC, Herridge RP, Ambrose BA, Macknight RC (2008) Transcriptome analysis of proliferating Arabidopsis endosperm reveals biological implications for the control of syncytial division, cytokinin signaling, and gene expression regulation. Plant Physiol 148:1964–1984. doi:10.1104/pp.108.128108
Debeaujon I, Nesi N, Perez P et al (2003) Proanthocyanidin-accumulating cells in Arabidopsis testa: regulation of differentiation and role in seed development. Plant Cell 15:2514–2531. doi:10.1105/tpc.014043
Deslauriers SD, Larsen PB (2010) FERONIA is a key modulator of brassinosteroid and ethylene responsiveness in Arabidopsis hypocotyls. Mol Plant 3:626–640. doi:10.1093/mp/ssq015
Dilkes BP, Spielman M, Weizbauer R et al (2008) The maternally expressed WRKY transcription factor TTG2 controls lethality in interploidy crosses of Arabidopsis. PLoS Biol 6:2707–2720. doi:10.1371/journal.pbio.0060308
Doughty J, Aljabri M, Scott RJ (2014) Flavonoids and the regulation of seed size in Arabidopsis. Biochem Soc Trans 42:364–369. doi:10.1042/BST20140040
Du L, Li N, Chen L et al (2014) The ubiquitin receptor DA1 regulates seed and organ size by modulating the stability of the ubiquitin-specific protease UBP15/SOD2 in Arabidopsis. Plant Cell 26:665–677. doi:10.1105/tpc.114.122663
Duan Q, Kita D, Li C et al (2010) FERONIA receptor-like kinase regulates RHO GTPase signaling of root hair development. Proc Natl Acad Sci USA 107:17821–17826. doi:10.1073/pnas.1005366107
Erdmann R, Gramzow L, Melzer R et al (2010) GORDITA (AGL63) is a young paralog of the Arabidopsis thaliana B(sister) MADS box gene ABS (TT16) that has undergone neofunctionalization. Plant J 63:914–924. doi:10.1111/j.1365-313X.2010.04290.x
Erilova A, Brownfield L, Exner V et al (2009) Imprinting of the polycomb group gene MEDEA serves as a ploidy sensor in Arabidopsis. PLoS Genet 5:e1000663. doi:10.1371/journal.pgen.1000663
Escobar-Restrepo J-M, Huck N, Kessler S et al (2007) The FERONIA receptor-like kinase mediates male-female interactions during pollen tube reception. Science 317:656–660. doi:10.1126/science.1143562
Fang W, Wang Z, Cui R et al (2012) Maternal control of seed size by EOD3/CYP78A6 in Arabidopsis thaliana. Plant J 70:929–939. doi:10.1111/j.1365-313X.2012.04907.x
Ferguson-Smith AC (2011) Genomic imprinting: the emergence of an epigenetic paradigm. Nat Rev Genet 12:565–575. doi:10.1038/nrg3032
Feuillet C, Leach JE, Rogers J et al (2011) Crop genome sequencing: lessons and rationales. Trends Plant Sci 16:77–88. doi:10.1016/j.tplants.2010.10.005
Figueiredo DD, Köhler C (2014) Signalling events regulating seed coat development. Biochem Soc Trans 42:358–363. doi:10.1042/BST20130221
Finkelstein R (2004) Plant hormones: biosynthesis, signal transduction, action!. Kluwer, Dordrecht, pp 549–573
Finkelstein RR, Lynch T (2000) The Arabidopsis abscisic acid response gene ABI5 encodes a basic leucine zipper transcription factor. Plant Cell 12:599–609. doi:10.1105/tpc.12.4.599
Finkelstein RR, Gampala SSL, Rock CD (2002) Abscisic acid signaling in seeds and seedlings. Plant Cell 14(Suppl):S15–S45
Friml J, Vieten A, Sauer M et al (2003) Efflux-dependent auxin gradients establish the apical–basal axis of Arabidopsis. Nature 426:147–153. doi:10.1038/nature02085
Fujioka S, Li J, Choi YH et al (1997) The Arabidopsis deetiolated2 mutant is blocked early in brassinosteroid biosynthesis. Plant Cell 9:1951–1962. doi:10.1105/tpc.9.11.1951
Garcia D, Saingery V, Chambrier P et al (2003) Arabidopsis haiku mutants reveal new controls of seed size by endosperm. Plant Physiol 131:1661–1670. doi:10.1104/pp.102.018762
Garcia D, Fitz Gerald JN, Berger F (2005) Maternal control of integument cell elongation and zygotic control of endosperm growth are coordinated to determine seed size in Arabidopsis. Plant Cell 17:52–60. doi:10.1105/tpc.104.027136
Gehring M, Huh JH, Hsieh T-F et al (2006) DEMETER DNA glycosylase establishes MEDEA polycomb gene self-imprinting by allele-specific demethylation. Cell 124:495–506. doi:10.1016/j.cell.2005.12.034
González-Guzmán M, Apostolova N, Bellés JM et al (2002) The short-chain alcohol dehydrogenase ABA2 catalyzes the conversion of xanthoxin to abscisic aldehyde. Plant Cell 14:1833–1846
Grossniklaus U, Vielle-Calzada JP, Hoeppner MA, Gagliano WB (1998) Maternal control of embryogenesis by MEDEA, a polycomb group gene in Arabidopsis. Science 280:446–450
Guitton A-E, Page DR, Chambrier P et al (2004) Identification of new members of Fertilisation Independent Seed Polycomb Group pathway involved in the control of seed development in Arabidopsis thaliana. Development 131:2971–2981. doi:10.1242/dev.01168
Guo H, Ye H, Li L, Yin Y (2009) A family of receptor-like kinases are regulated by BES1 and involved in plant growth in Arabidopsis thaliana. Plant Signal Behav 4:784–786. doi:10.1073/pnas.0812346106
Haig D, Westoby R (1989) Parent-specific gene-expression and the triploid endosperm. Am Nat 134:147–155
Haig D, Westoby R (1991) Genomic imprinting in endosperm: its effects on seed development in crosses between species, and between different ploidies of the same species, and its implications for the evolution of apomixis. Philos Trans Biol Sci 333:1–13
Hamann T, Benkova E, Bäurle I et al (2002) The Arabidopsis BODENLOS gene encodes an auxin response protein inhibiting MONOPTEROS-mediated embryo patterning. Genes Dev 16:1610–1615. doi:10.1101/gad.229402
Harper JL, Lovell PH, Moore KG (1970) The shapes and sizes of seeds. Annu Rev Ecol Syst 1:327–356. doi:10.1146/annurev.es.01.110170.001551
Hategan L, Godza B, Kozma-Bognar L et al (2014) Differential expression of the brassinosteroid receptor-encoding BRI1 gene in Arabidopsis. Planta 239:989–1001. doi:10.1007/s00425-014-2031-4
Haughn G, Chaudhury A (2005) Genetic analysis of seed coat development in Arabidopsis. Trends Plant Sci 10:472–477. doi:10.1016/j.tplants.2005.08.005
Hehenberger E, Kradolfer D, Köhler C (2012) Endosperm cellularization defines an important developmental transition for embryo development. Development 139:2031–2039. doi:10.1242/dev.077057
Higuchi M, Pischke MS, Mähönen AP et al (2004) In planta functions of the Arabidopsis cytokinin receptor family. Proc Natl Acad Sci USA 101:8821–8826. doi:10.1073/pnas.0402887101
Hsieh T-F, Shin J, Uzawa R et al (2011) Regulation of imprinted gene expression in Arabidopsis endosperm. Proc Natl Acad Sci USA 108:1755–1762. doi:10.1073/pnas.1019273108
Hughes R, Spielman M, Schruff MC et al (2008) Yield assessment of integument-led seed growth following targeted repair of auxin response factor 2. Plant Biotechnol J 6:758–769. doi:10.1111/j.1467-7652.2008.00359.x
Ingouff M, Jullien PE, Berger F (2006) The female gametophyte and the endosperm control cell proliferation and differentiation of the seed coat in Arabidopsis. Plant Cell 18:3491–3501. doi:10.1105/tpc.106.047266
Jahnke S, Scholten S (2009) Epigenetic resetting of a gene imprinted in plant embryos. Curr Biol 19:1677–1681. doi:10.1016/j.cub.2009.08.053
Jenik PD, Barton MK (2005) Surge and destroy: the role of auxin in plant embryogenesis. Development 132:3577–3585. doi:10.1242/dev.01952
Jiang W-B, Lin W-H (2013) Brassinosteroid functions in Arabidopsis seed development. Plant Signal Behav 8:1–5. doi:10.4161/psb.25928
Jiang W-B, Huang H-Y, Hu Y-W et al (2013) Brassinosteroid regulates seed size and shape in Arabidopsis. Plant Physiol. doi:10.1104/pp.113.217703
Jofuku KD, den Boer BG, Van Montagu M, Okamuro JK (1994) Control of Arabidopsis flower and seed development by the homeotic gene APETALA2. Plant Cell 6:1211–1225
Jofuku KD, Omidyar PK, Gee Z, Okamuro JK (2005) Control of seed mass and seed yield by the floral homeotic gene APETALA2. Proc Natl Acad Sci USA 102:3117–3122. doi:10.1073/pnas.0409893102
Johnson CS, Kolevski B, Smyth DR (2002) TRANSPARENT TESTA GLABRA2, a trichome and seed coat development gene of Arabidopsis, encodes a WRKY transcription factor. Plant Cell 14:1359–1375. doi:10.1105/tpc.001404
Jullien PE, Susaki D, Yelagandula R et al (2012) DNA methylation dynamics during sexual reproduction in Arabidopsis thaliana. Curr Biol 22:1825–1830. doi:10.1016/j.cub.2012.07.061
Kang X, Ni M (2006) Arabidopsis SHORT HYPOCOTYL UNDER BLUE1 contains SPX and EXS domains and acts in cryptochrome signaling. Plant Cell 18:921–934. doi:10.1105/tpc.105.037879
Kang I-H, Steffen JG, Portereiko MF et al (2008) The AGL62 MADS domain protein regulates cellularization during endosperm development in Arabidopsis. Plant Cell 20:635–647. doi:10.1105/tpc.107.055137
Kang X, Li W, Zhou Y, Ni M (2013) A WRKY transcription factor recruits the SYG1-like protein SHB1 to activate gene expression and seed cavity enlargement. PLoS Genet 9:e1003347. doi:10.1371/journal.pgen.1003347
Kanno Y, Jikumaru Y, Hanada A et al (2010) Comprehensive hormone profiling in developing Arabidopsis seeds: examination of the site of ABA biosynthesis, ABA transport and hormone interactions. Plant Cell Physiol 51:1988–2001. doi:10.1093/pcp/pcq158
Kesavan M, Song JT, Seo HS (2013) Seed size: a priority trait in cereal crops. Physiol Plant 147:113–120. doi:10.1111/j.1399-3054.2012.01664.x
Khan D, Millar JL, Girard IJ, Belmonte MF (2014) Transcriptional circuitry underlying seed coat development in Arabidopsis. Plant Sci 219–220:51–60. doi:10.1016/j.plantsci.2014.01.004
Kinoshita T, Miura A, Choi Y et al (2004) One-way control of FWA imprinting in Arabidopsis endosperm by DNA methylation. Science 303:521–523. doi:10.1126/science.1089835
Kiyosue T, Ohad N, Yadegari R et al (1999) Control of fertilization-independent endosperm development by the MEDEA polycomb gene in Arabidopsis. Proc Natl Acad Sci USA 96:4186–4191
Klucher KM, Chow H, Reiser L, Fischer RL (1996) The AINTEGUMENTA gene of Arabidopsis required for ovule and female gametophyte development is related to the floral homeotic gene APETALA2. Plant Cell 8:137–153. doi:10.1105/tpc.8.2.137
Kradolfer D, Hennig L, Köhler C (2013) Increased maternal genome dosage bypasses the requirement of the FIS polycomb repressive complex 2 in Arabidopsis seed development. PLoS Genet 9:e1003163. doi:10.1371/journal.pgen.1003163
Krizek BA (2009) Making bigger plants: key regulators of final organ size. Curr Opin Plant Biol 12:17–22. doi:10.1016/j.pbi.2008.09.006
Kunieda T, Mitsuda N, Ohme-Takagi M et al (2008) NAC family proteins NARS1/NAC2 and NARS2/NAM in the outer integument regulate embryogenesis in Arabidopsis. Plant Cell 20:2631–2642. doi:10.1105/tpc.108.060160
Lafon-Placette C, Köhler C (2014) Embryo and endosperm, partners in seed development. Curr Opin Plant Biol 17:64–69. doi:10.1016/j.pbi.2013.11.008
Langridge P, Fleury D (2011) Making the most of “omics” for crop breeding. Trends Biotechnol 29:33–40. doi:10.1016/j.tibtech.2010.09.006
Le BH, Cheng C, Bui AQ et al (2010) Global analysis of gene activity during Arabidopsis seed development and identification of seed-specific transcription factors. Proc Natl Acad Sci USA 107:8063–8070. doi:10.1073/pnas.1003530107
Leyser O (2005) Auxin distribution and plant pattern formation: how many angels can dance on the point of PIN? Cell 121:819–822. doi:10.1016/j.cell.2005.06.005
Li N, Li Y (2014) Ubiquitin-mediated control of seed size in plants. Front Plant Sci 5:332. doi:10.3389/fpls.2014.00332
Li H, Johnson P, Stepanova A et al (2004) Convergence of signaling pathways in the control of differential cell growth in Arabidopsis. Dev Cell 7:193–204. doi:10.1016/j.devcel.2004.07.002
Li Y, Zheng L, Corke F et al (2008) Control of final seed and organ size by the DA1 gene family in Arabidopsis thaliana. Genes Dev 22:1331–1336. doi:10.1101/gad.463608
Li J, Nie X, Tan JLH, Berger F (2013) Integration of epigenetic and genetic controls of seed size by cytokinin in Arabidopsis. Proc Natl Acad Sci USA 110:15479–15484. doi:10.1073/pnas.1305175110
Locascio A, Roig-Villanova I, Bernardi J, Varotto S (2014) Current perspectives on the hormonal control of seed development in Arabidopsis and maize: a focus on auxin. Front Plant Sci 5:412. doi:10.3389/fpls.2014.00412
Luo M, Bilodeau P, Dennis ES et al (2000) Expression and parent-of-origin effects for FIS2, MEA, and FIE in the endosperm and embryo of developing Arabidopsis seeds. Proc Natl Acad Sci USA 97:10637–10642. doi:10.1073/pnas.170292997
Luo M, Dennis ES, Berger F et al (2005) MINISEED3 (MINI3), a WRKY family gene, and HAIKU2 (IKU2), a leucine-rich repeat (LRR) KINASE gene, are regulators of seed size in Arabidopsis. Proc Natl Acad Sci USA 102:17531–17536. doi:10.1073/pnas.0508418102
Lur HS, Setter TL (1993) Role of auxin in maize endosperm development (timing of nuclear dna endoreduplication, zein expression, and cytokinin). Plant Physiol 103:273–280
Makarevich G, Villar CBR, Erilova A, Köhler C (2008) Mechanism of PHERES1 imprinting in Arabidopsis. J Cell Sci 121:906–912. doi:10.1242/jcs.023077
McCarty DR (1995) Genetic control and integration of maturation and germination pathways in seed development. Annu Rev Plant Physiol Plant Mol Biol 46:71–93. doi:10.1146/annurev.pp.46.060195.000443
Mizzotti C, Mendes MA, Caporali E et al (2012) The MADS box genes SEEDSTICK and ARABIDOPSIS B(sister) play a maternal role in fertilization and seed development. Plant J. doi:10.1111/j.1365-313X.2011.04878.x
Morley-Smith ER, Pike MJ, Findlay K et al (2008) The transport of sugars to developing embryos is not via the bulk endosperm in oilseed rape seeds. Plant Physiol 147:2121–2130. doi:10.1104/pp.108.124644
Nesi N, Debeaujon I, Jond C et al (2002) The TRANSPARENT TESTA16 locus encodes the ARABIDOPSIS BSISTER MADS domain protein and is required for proper development and pigmentation of the seed coat. Plant Cell 14:2463–2479
Nishimura C, Ohashi Y, Sato S et al (2004) Genetic analysis of Arabidopsis Histidine Kinase genes encoding cytokinin receptors reveals their overlapping biological functions in the regulation of shoot and root growth in Arabidopsis thaliana. Plant Cell 16:1365–1377. doi:10.1105/tpc.021477
Nowack MK, Grini PE, Jakoby MJ et al (2006) A positive signal from the fertilization of the egg cell sets off endosperm proliferation in angiosperm embryogenesis. Nat Genet 38:63–67. doi:10.1038/ng1694
Nowack MK, Shirzadi R, Dissmeyer N et al (2007) Bypassing genomic imprinting allows seed development. Nature 447:312–315. doi:10.1038/nature05770
Nowack MK, Ungru A, Bjerkan KN et al (2010) Reproductive cross-talk: seed development in flowering plants. Biochem Soc Trans 38:604–612. doi:10.1042/BST0380604
Ohto M-A, Fischer RL, Goldberg RB et al (2005) Control of seed mass by APETALA2. Proc Natl Acad Sci USA 102:3123–3128. doi:10.1073/pnas.0409858102
Ohto M, Floyd SK, Fischer RL et al (2009) Effects of APETALA2 on embryo, endosperm, and seed coat development determine seed size in Arabidopsis. Sex Plant Reprod 22:277–289. doi:10.1007/s00497-009-0116-1
Okamuro JK, Szeto W, Lotys-Prass C, Jofuku KD (1997) Photo and hormonal control of meristem identity in the Arabidopsis flower mutants apetala2 and apetala1. Plant Cell 9:37–47. doi:10.1105/tpc.9.1.37
Okushima Y, Mitina I, Quach HL, Theologis A (2005) AUXIN RESPONSE FACTOR 2 (ARF2): a pleiotropic developmental regulator. Plant J 43:29–46. doi:10.1111/j.1365-313X.2005.02426.x
Peer WA, Murphy AS (2007) Flavonoids and auxin transport: modulators or regulators? Trends Plant Sci 12:556–563
Pinyopich A, Ditta GS, Savidge B et al (2003) Assessing the redundancy of MADS-box genes during carpel and ovule development. Nature 424:85–88. doi:10.1038/nature01741
Prasad K, Zhang X, Tobón E, Ambrose BA (2010) The Arabidopsis B-sister MADS-box protein, GORDITA, represses fruit growth and contributes to integument development. Plant J 62:203–214. doi:10.1111/j.1365-313X.2010.04139.x
Riechmann JL, Meyerowitz EM (1998) The AP2/EREBP family of plant transcription factors. Biol Chem 379:633–646
Riefler M, Novak O, Strnad M, Schmülling T (2006) Arabidopsis cytokinin receptor mutants reveal functions in shoot growth, leaf senescence, seed size, germination, root development, and cytokinin metabolism. Plant Cell 18:40–54. doi:10.1105/tpc.105.037796
Roszak P, Köhler C (2011) Polycomb group proteins are required to couple seed coat initiation to fertilization. Proc Natl Acad Sci USA 108:20826–20831. doi:10.1073/pnas.1117111108
Schoft VK, Chumak N, Choi Y et al (2011) Function of the DEMETER DNA glycosylase in the Arabidopsis thaliana male gametophyte. Proc Natl Acad Sci USA 108:8042–8047. doi:10.1073/pnas.1105117108
Schruff MC, Spielman M, Tiwari S et al (2006) The AUXIN RESPONSE FACTOR 2 gene of Arabidopsis links auxin signalling, cell division, and the size of seeds and other organs. Development 133:251–261. doi:10.1242/dev.02194
Schuettengruber B, Cavalli G (2009) Recruitment of polycomb group complexes and their role in the dynamic regulation of cell fate choice. Development 136:3531–3542. doi:10.1242/dev.033902
Scott RJ, Spielman M, Bailey J, Dickinson HG (1998) Parent-of-origin effects on seed development in Arabidopsis thaliana. Development 125:3329–3341
Scott R, Tratt J, Bolbol A (2013) Polyploid and hybrid genomics. Wiley, Oxford
Sun X, Shantharaj D, Kang X, Ni M (2010) Transcriptional and hormonal signaling control of Arabidopsis seed development. Curr Opin Plant Biol 13:611–620. doi:10.1016/j.pbi.2010.08.009
Takahashi N, Nakazawa M, Shibata K et al (2005) shk1-D, a dwarf Arabidopsis mutant caused by activation of the CYP72C1 gene, has altered brassinosteroid levels. Plant J 42:13–22. doi:10.1111/j.1365-313X.2005.02357.x
Terasaka K, Blakeslee JJ, Titapiwatanakun B et al (2005) PGP4, an ATP binding cassette P-glycoprotein, catalyzes auxin transport in Arabidopsis thaliana roots. Plant Cell 17:2922–2939. doi:10.1105/tpc.105.035816
Tiwari S, Spielman M, Schulz R et al (2010) Transcriptional profiles underlying parent-of-origin effects in seeds of Arabidopsis thaliana. BMC Plant Biol 10:72. doi:10.1186/1471-2229-10-72
Tsukaya H (2006) Mechanism of leaf-shape determination. Annu Rev Plant Biol 57:477–496. doi:10.1146/annurev.arplant.57.032905.105320
Turk EM, Fujioka S, Seto H et al (2003) CYP72B1 inactivates brassinosteroid hormones: an intersection between photomorphogenesis and plant steroid signal transduction. Plant Physiol 133:1643–1653. doi:10.1104/pp.103.030882
Ulmasov T, Hagen G, Guilfoyle TJ (1999) Activation and repression of transcription by auxin-response factors. Proc Natl Acad Sci USA 96:5844–5849
Vandenbussche F, Van Der Straeten D (2004) Shaping the shoot: a circuitry that integrates multiple signals. Trends Plant Sci 9:499–506. doi:10.1016/j.tplants.2004.08.002
Vanstraelen M, Benková E (2012) Hormonal interactions in the regulation of plant development. Annu Rev Cell Dev Biol 28:463–487. doi:10.1146/annurev-cellbio-101011-155741
Varshney RK, Nayak SN, May GD, Jackson SA (2009) Next-generation sequencing technologies and their implications for crop genetics and breeding. Trends Biotechnol 27:522–530. doi:10.1016/j.tibtech.2009.05.006
Vert G, Walcher CL, Chory J, Nemhauser JL (2008) Integration of auxin and brassinosteroid pathways by Auxin Response Factor 2. Proc Natl Acad Sci USA 105:9829–9834. doi:10.1073/pnas.0803996105
Wabnik K, Robert HS, Smith RS, Friml J (2013) Modeling framework for the establishment of the apical-basal embryonic axis in plants. Curr Biol 23:2513–2518. doi:10.1016/j.cub.2013.10.038
Wang J-W, Schwab R, Czech B et al (2008) Dual effects of miR156-targeted SPL genes and CYP78A5/KLUH on plastochron length and organ size in Arabidopsis thaliana. Plant Cell 20:1231–1243. doi:10.1105/tpc.108.058180
Wang A, Garcia D, Zhang H et al (2010) The VQ motif protein IKU1 regulates endosperm growth and seed size in Arabidopsis. Plant J 63:670–679. doi:10.1111/j.1365-313X.2010.04271.x
Weinhofer I, Hehenberger E, Roszak P et al (2010) H3K27me3 profiling of the endosperm implies exclusion of polycomb group protein targeting by DNA methylation. PLoS Genet 6:14. doi:10.1371/journal.pgen.1001152
Werner T, Motyka V, Laucou V et al (2003) Cytokinin-deficient transgenic Arabidopsis plants show multiple developmental alterations indicating opposite functions of cytokinins in the regulation of shoot and root meristem activity. Plant Cell 15:2532–2550. doi:10.1105/tpc.014928
Williams L, Carles CC, Osmont KS, Fletcher JC (2005) A database analysis method identifies an endogenous trans-acting short-interfering RNA that targets the Arabidopsis ARF2, ARF3, and ARF4 genes. Proc Natl Acad Sci USA 102:9703–9708. doi:10.1073/pnas.0504029102
Wolf S, Höfte H (2014) Growth control: a saga of cell walls, ROS, and peptide receptors. Plant Cell 26:1848–1856. doi:10.1105/tpc.114.125518
Xia T, Li N, Dumenil J et al (2013) The ubiquitin receptor DA1 interacts with the E3 ubiquitin ligase DA2 to regulate seed and organ size in Arabidopsis. Plant Cell 25:3347–3359. doi:10.1105/tpc.113.115063
Xiao W, Brown RC, Lemmon BE et al (2006) Regulation of seed size by hypomethylation of maternal and paternal genomes. Plant Physiol 142:1160–1168. doi:10.1104/pp.106.088849
Yang WC, Ye D, Xu J, Sundaresan V (1999) The SPOROCYTELESS gene of Arabidopsis is required for initiation of sporogenesis and encodes a novel nuclear protein. Genes Dev 13:2108–2117
Yang J, Zhang J, Huang Z et al (2002) Correlation of cytokinin levels in the endosperms and roots with cell number and cell division activity during endosperm development in rice. Ann Bot 90:369–377
Yu F, Li J, Huang Y et al (2014) FERONIA receptor kinase controls seed size in Arabidopsis thaliana. Mol Plant 7:920–922. doi:10.1093/mp/ssu010
Zhang Y, Liang W, Shi J et al (2013) MYB56 encoding a R2R3 MYB transcription factor regulates seed size in Arabidopsis thaliana. J Integr Plant Biol 55:1166–1178. doi:10.1111/jipb.12094
Zhou Y, Zhang X, Kang X et al (2009) SHORT HYPOCOTYL UNDER BLUE1 associates with MINISEED3 and HAIKU2 promoters in vivo to regulate Arabidopsis seed development. Plant Cell 21:106–117. doi:10.1105/tpc.108.064972
Acknowledgments
The authors are grateful to Edward Kiegle for assistance in preparing the manuscript. This work was supported by the Consejo Nacional de Ciencia y Tecnología CONACYT–MEXICO and Marie Curie-EVOCODE (to G.O.A); the International European Fellowship-METMADS project and the Centro Nazionale di Ricerca Italiano (Fondo IBF-AR-01-2014Mi) (to I.E.); and a fellowship from the Università degli Studi di Milano (to D.P.).
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by Silvia Coimbra.
A contribution to the special issue ‘From Gametes to Seeds’.
Rights and permissions
About this article
Cite this article
Orozco-Arroyo, G., Paolo, D., Ezquer, I. et al. Networks controlling seed size in Arabidopsis . Plant Reprod 28, 17–32 (2015). https://doi.org/10.1007/s00497-015-0255-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00497-015-0255-5