Abstract
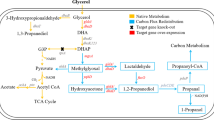
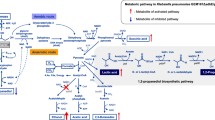
In the present study, we developed an efficient method of 1,3-propanediol (1,3-PD) production from glycerol by genetic engineering of Klebsiella pneumoniae AK mutant strains. The proposed approach eliminated by-product formation and IPTG induction resulted in maximal production of 1,3-PD. A series of recombinant strains was designed to constitutively express the dhaB and/or dhaT genes, using the bacteriophage T5 PDE20 promoter and the rho-independent transcription termination signal of the Rahnella aquatilis levansucrase gene. Among these strains, AK/pConT expressing dhaT alone gave the highest yield of 1,3-PD. Fed-batch fermentation resulted in efficient production of 1,3-PD from either pure or crude glycerol, without by-product formation.





Similar content being viewed by others
References
Nakamura CE, Whited GM (2003) Metabolic engineering for the microbial production of 1,3-propanediol. Curr Opin Biotechnol 14:454–459
Biebl H, Menzel K, Zeng AP, Deckwer WD (1999) Microbial production of 1,3-propanediol. Appl Microbiol Biotechnol 52:289–297
Saxena RK, Anand P, Saran S, Isar J (2009) Microbial production of 1,3-propanediol: recent developments and emerging opportunities. Biotechnol Adv 27:895–913
Homann T, Tag C, Biebl H, Deckwer WD, Schink B (1990) Fermentation of glycerol to 1,3-propanediol by Klebsiella and Citrobacter strains. Appl Microbiol Biotechnol 33:121–126
Daniel R, Stuertz K, Gottschalk G (1995) Biochemical and molecular characterization of the oxidative branch of glycerol utilization by Citrobacter freundii. J Bacteriol 177:392–401
El-Ziney MG, Debevere JM (1998) The effect of reuterin on Listeria monocytogenes and Escherichia coli O157:h7 in milk and cottage cheese. J Food Prot 61:1275–1280
Marçal D, Rêgo AT, Carrondo MA, Enguita FJ (2009) 1,3-Propanediol dehydrogenase from Klebsiella pneumoniae: decameric quaternary structure and possible subunit cooperativity. J Bacteriol 191:1143–1151
Bhatia SK, Kurian JV (2008) Biological characterization of sorona polymer from corn-derived 1,3-propanediol. Biotechnol Lett 30:619–623
Ma J, Rao Z, Xu L, Liao X, Fang H, Zhuge B, Zhuge J (2009) Expression of dha operon required for 1,3-PD formation in Escherichia coli and Saccharomyces cerevisiae. Curr Microbiol 60:191–198
Gellissen G (2000) Heterologous protein production in methylotrophic yeasts. Appl Microbiol Biotechnol 54:741–750
Sohn JH (1997) Characterization of telomere-associated ARSs (autonomously replicating sequences) and their use for multiple gene integration in Hansenula polymorpha. PhD thesis. Korea Advanced Institute of Science and Technology, Taejon, Korea
Heo JH, Hong WK, Cho EY, Kim MW, Kim JY, Kim CH, Rhee SK, Kang HA (2003) Properties of the Hansenula polymorpha-derived constitutive GAP promoter, assessed using an HSA reporter gene. FEMS Yeast Res 4:175–184
Hill J, Donald KA, Griffiths DE (1991) DMSO-enhanced whole cell yeast transformation. Nucleic Acids Res 9:5791
Hong WK, Kim CH, Heo SY, Luo LH, Oh BR, Seo JW (2010) Enhanced production of ethanol from glycerol by engineered Hansenula polymorpha expressing pyruvate decarboxylase and aldehyde dehydrogenase genes from Zymomonas mobilis. Biotechnol Lett 32:1077–1082
Gonzalez-Pajuelo M, Meynial-Salles I, Mendes F, Soucaille P, Vasconcelos I (2006) Microbial conversion of glycerol to 1,3-propanediol: physiological comparison of a natural producer, Clostridium butyricum VPI 3266, and an engineered strain, Clostridium acetobutylicum DG1(pSPD5). Appl Environ Microbiol 72:96–101
Olivier C, Aline LF (2001) Primers and a specific DNA probe for detecting lactic acid bacteria producing 3-hydroxypropionaldehyde from glycerol in spoiled ciders. J Food Prot 64:833–837
Sulzenbacher G, Alvarez K, Van Den Heuvel RH, Versluis C, Spinelli S, Campanacci V, Valencia C, Cambillau C, Eklund H, Tegoni M (2004) Crystal structure of E. coli alcohol dehydrogenase YqhD: evidence of a covalently modified NADP coenzyme. J Mol Biol 342:489–502
Oh BR, Seo JW, Choi MH, Kim CH (2008) Optimization of culture conditions for 1,3-propanediol production from crude glycerol by Klebsiella pneumoniae using response surface methodology. Biotechnol Bioprocess Eng 13:524–532
Seo MY, Seo JW, Heo SY, Baek JO, Rairakhwada D, Oh BR, Seo PS, Choi MH, Kim CH (2009) Elimination of by-product formation during production of 1,3-propanediol in Klebsiella pneumoniae by inactivation of glycerol oxidative pathway. Appl Microbiol Biotechnol 84:527–534
Seo JW, Seo MY, Oh BR, Heo SY, Baek JO, Rairakhwada D, Luo LH, Hong WK, Kim CH (2010) Identification and utilization of a 1,3-propanediol oxidoreductase isoenzyme for production of 1,3-propanediol from glycerol in Klebsiella pneumoniae. Appl Microbiol Biotechnol 85:659–666
Seo JW, Hong WK, Rairakhwada R, Seo PS, Choi MH, Song KB, Rhee SK, Kim CH (2009) An efficient plasmid vector for constitutive high-level expression of foreign genes in Escherichia coli. Biotechnol Lett 31:877–881
Xu YZ, Guo NN, Zheng ZM, Ou XJ, Liu HJ, Liu DH (2009) Metabolism in 1,3-propanediol fed-batch fermentation by a d-lactate deficient mutant of Klebsiella pneumoniae. Biotechnol Bioeng 104:965–972
Zhuge B, Zhang C, Fang H, Zhuge J, Permaul K (2010) Expression of 1,3-propanediol oxidoreductase and its isoenzyme in Klebsiella pneumoniae for bioconversion of glycerol into 1,3-propanediol. Appl Microbiol Biotechnol 87:2177–2184
Acknowledgments
This subject was supported by Korea Ministry of Environment as “Converging technology project” and by the New & Renewable Energy Technology Development Program of the Korea Institute of Energy Technology Evaluation and Panning (KETEP) Grant (2010T100100690) funded by the Korea government Ministry of Knowledge Economy.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Oh, BR., Seo, JW., Heo, SY. et al. Efficient production of 1,3-propanediol from glycerol upon constitutive expression of the 1,3-propanediol oxidoreductase gene in engineered Klebsiella pneumoniae with elimination of by-product formation. Bioprocess Biosyst Eng 36, 757–763 (2013). https://doi.org/10.1007/s00449-013-0901-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00449-013-0901-y