Abstract
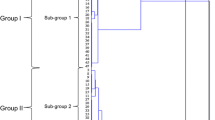
The aim of the present study was to genetically characterize Fasciola hepatica strains from diverse ecogeographical regions (America and Europe), susceptible and resistant to Triclabendazole, using the random amplified polymorphic DNA fragments (RAPDs-PCR) technique to elucidate genetic variability between the different isolates. Ten different oligonucleotide primers of 10 bases with GC content varying from 50–70 % were used. A polymerase chain reaction (PCR) was carried out in 25 μl of total volume. Duplicate PCR reactions on each individual template DNA were performed to test the reproducibility of the individual DNA bands. The size of the RAPD-PCR fragments was determined by the reciprocal plot between the delay factors (Rf) versus the logarithm of molecular weight ladder. The phenogram obtained showed three main clusters, the major of which contained European Strains (Cullompton and Sligo) showing a genetic distance of 27.2 between them. The American strains (Cedive and Cajamarca) on the other hand formed each their distinctive group but clearly maintaining a closer genetic relationship among them than that to their European counterparts, with which showed a distance of 33.8 and 37.8, respectively. This polymorphism would give this species enhanced adaptability against the host, as well as the environment. The existence of genetically different populations of F. hepatica could allow, against any selection pressure, natural or artificial (for use fasciolicides products and/or control measures), one or more populations of F. hepatica to be able to survive and create resistance or adaptability to such selective pressure.



Similar content being viewed by others
References
Ai L, Weng Y, Elsheikha H et al (2011) Genetic diversity and relatedness of Fasciola spp. isolates from different hosts and geographic regions revealed by analysis of mitochondrial DNA sequences. Vet Parasitol 181:329–334
AVMA (2001) 2000 Report of the AVMA Panel on Euthanasia. J Am Vet Med Assoc 218:669–696
Boray JC (1994) Disease of domestic animals caused by flukes. FAO, Rome
Campbell CS, Alice LA, Wright WA (1999) Comparisons of within-population genetic variation in sexual and agamospermous Amelanchier (Rosaceae) using RAPD markers. Plant Syst Evol 215:157–167
Elliott T, Muller A, Brockwell Y et al (2014) Evidence for high genetic diversity of NAD1 and COX1 mitochondrial haplotypes among triclabendazole resistant and susceptible populations and field isolates of Fasciola hepatica (liver fluke) in Australia. Vet Parasitol 200:90–96
Fairweather I (2011) Liver fluke isolates: a question of provenance. Vet Parasitol 176:1–8
Fletcher H, Hoey E, Orr N, Trudgett A, Fairweather I, Robinson M (2004) The occurrence and significance of triploidy in the liver fluke, Fasciola hepatica. Parasitology 128:69–72
Gasser RB (1999) PCR-based technology in veterinary parasitology. Vet Parasitol 84:229–258
Gasser RB (2006) Molecular tools—advances, opportunities and prospects. Vet Parasitol 136:69–89
Gomes MA, Melo MN, Macedo AM, Furst C, Silva EF (2000) RAPD in the analysis of isolates of Entamoeba histolytica. Acta Trop 75:71–77
Olaechea F, Lovera V, Larroza M, Raffo F, Cabrera R (2011) Resistance of Fasciola hepatica against triclabendazole in cattle in Patagonia (Argentina). Vet Parasitol 178:364–366
Ortiz P, Scarcella S, Cerna C et al (2013) Resistance of Fasciola hepatica against Triclabendazole in cattle in Cajamarca (Peru): a clinical trial and an in vivo efficacy test in sheep. Vet Parasitol 128:69–72
Overend DJ, Bowen FL (1995) Resistance of Fasciola hepatica to triclabendazole. Aust Vet J 22:275–276
Pavel AB, Vasile CI (2012) PyElph-a software tool for gel images analysis and phylogenetics. BMC Bioinforma 13:9
Sambrook J, Fritsch EF, Maniatis T (1989) Molecular cloning, vol 2. Cold spring harbor laboratory press, New York
Sanabria R, Ceballos L, Moreno L, Romero J, Lanusse C, Alvarez L (2013) Identification of a field isolate of Fasciola hepatica resistant to albendazole and susceptible to triclabendazole. Vet Parasitol 193:105–110
Solana HD, Rodriguez JA, Lanusse CE (2001) Comparative metabolism of albendazole and albendazole sulphoxide by different helminth parasites. Parasitol Res 87:275–280
Staub JE, Serquen FC, Gupta M (1996) Genetic markers, map construction, and their application in plant breeding. Hortic Sci 31:729–741
Teofanova D, Kantzoura V, Walker S et al (2011) Genetic diversity of liver flukes Fasciola hepatica from Eastern Europe. Infect Genet Evol 11:109–115
Vilas R, Vázquez-Prieto S, Paniagua E (2012) Contrasting patterns of population genetic structure of Fasciola hepatica from cattle and sheep: implications for the evolution of anthelmintic resistance. Infect Genet Evol 12:45–52
Walker S, Johnston C, Hoey E et al (2011) Population dynamics of the liver fluke, Fasciola hepatica: the effect of time and spatial separation on the genetic diversity of fluke populations in the Netherlands. Parasitology 138:215–223
Williams JG, Kubelik AR, Livak KJ, Rafalski JA, Tingey SV (1990) DNA polymorphisms amplified by arbitrary primers are useful as genetic markers. Nucleic Acids Res 18:6531–6535
Conflict of interest
The authors declare that they have no conflict of interest.
Declaration of animal welfare
All the authors of the present manuscript declare that animal procedures and management protocols were approved by the Ethics Committee according to Animal Welfare Policy (Act 087/02) of the Faculty of Veterinary Medicine, Universidad Nacional del Centro de la Provincia de Buenos Aires (UNCPBA), Tandil, Argentina (http://www.vet.unicen.edu.ar), and to the internationally accepted animal welfare guidelines (AVMA 2001). The experiments comply with the current laws of the country in which they were performed.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Scarcella, S., Miranda-Miranda, E., Solana, M.V. et al. Approach to molecular characterization of different strains of Fasciola hepatica using random amplified polymorphic DNA polymerase chain reaction. Parasitol Res 114, 1341–1345 (2015). https://doi.org/10.1007/s00436-015-4310-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00436-015-4310-9