Abstract
Purpose
To set up a method by use of gene expression data to predict the prognosis of breast cancer patients on the basis of genes as few as possible, but maintaining the accuracy of prediction, we reanalyze the data from van’t Veer et al. (Nature 415:530–536, 2002) and van de Vijver et al. (N Engl J Med 347:1999–2009, 2002).
Methods
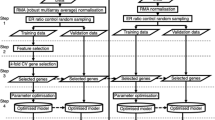
A three-step method based on re-sampling strategy is employed to select the prognostic genes. And based on these genes, a predictive approach is established. Validation sets are used to testify the predictive power of the prognostic genes.
Results
We have discovered 13 genes as the most informative ones to predict the clinical outcomes of breast cancer patients with lymph node-negative. The validation results show the robust performances of these genes. And the results of further analysis illustrate the significant association of the prediction to the time of metastases and overall survival.
Conclusion
Our predictive approach is useful in prognosis prediction for breast cancer patients with lymph node-negative. The gene markers provide valuable information for the progression of breast cancer and suggest potential target genes for treating the cancer.


Similar content being viewed by others
References
Becker M, Sommer A, Kratzschmar JR, Seidel H, Pohlenz HD, Fichtner I (2005) Distinct gene expression patterns in a tamoxifen-sensitive human mammary carcinoma xenograft and its tamoxifen-resistant subline MaCa 3366/TAM. Mol Cancer Ther 4:151–168
Bhattacharjee A, Richards WG, Staunton J, Li C, Monti S, Vasa P, Ladd C, Beheshti J, Bueno R, Gillette M, Loda M, Weber G, Mark EJ, Lander ES, Wong W, Johnson BE, Golub TR, Sugarbaker DJ, Meyerson M (2001) Classification of human lung carcinomas by mRNA expression profiling reveals distinct adenocarcinoma subclasses. Proc Natl Acad Sci USA 98:13790–13795
Butt AJ, Dickson KA, McDougall F, Baxter RC (2003) Insulin-like growth factor-binding protein-5 inhibits the growth of human breast cancer cells in vitro and in vivo. J Biol Chem 278:29676–29685
Cromer A, Carles A, Millon R, Ganguli G, Chalmel F, Lemaire F, Young J, Dembele D, Thibault C, Muller D, Poch O, Abecassis J, Wasylyk B (2004) Identification of genes associated with tumorigenesis and metastatic potential of hypopharyngeal cancer by microarray analysis. Oncogene 23:2484–2498
Carr JA, Havstad S, Zarbo RJ, Divine G, Mackowiak P, Velanovich V (2000) The association of HER-2/neu amplification with breast cancer recurrence. Arch Surg 135:1469–1474
Egawa C, Miyoshi Y, Taguchi T, Tamaki Y, Noguchi S (2002) High BRCA2 mRNA expression predicts poor prognosis in breast cancer patients. Int J Cancer 98:879–882
Fogli A, Rodriguez D, Eymard-Pierre E, Bouhour F, Labauge P, Meaney BF, Zeesman S, Kaneski CR, Schiffmann R, Boespflug-Tanguy O (2003) Ovarian failure related to eukaryotic initiation factor 2B mutations. Am J Hum Genet 72:1544–1550
Glanzer JG, Eberwine JH (2004) Expression profiling of small cellular samples in cancer: less is more. Br J Cancer 90:1111–1114
Golub TR, Slonim DK, Tamayo P, Huard C, Gaasenbeek M, Mesirov JP, Coller H, Loh ML, Downing JR, Caligiuri MA, Bloomfiel CD, Lander ES (1999) Molecular classification of cancer: class discovery and class prediction by gene expression monitoring. Science 286:531–537
Gruvberger S, Ringnér M, Chen Y, Panavally S, Saal LH, Borg Å, Fernö M (2001) Estrogen receptor status in breast cancer is associated with remarkably distinct gene expression pattern. Cancer Res 61:5979–5984
Hao X, Sun B, Hu L, Lahdesmaki H, Dunmire V, Feng Y, Zhang SW, Wang H, Wu C, Wang H, Fuller GN, Symmans WF, Shmulevich I, Zhang W (2004) Differential gene and protein expression in primary breast malignancies and their lymph node metastases as revealed by combined cDNA microarray and tissue microarray analysis. Cancer 100:1110–1122
Huang E, Cheng SH, Dressman H, Pittman J, Tsou MH, Horng CF, Bild A, Iversen ES, Liao M, Chen CM, West M, Nevins JR, Huang AT (2003) Gene expression predictors of breast cancer outcomes. Lancet 361:1590–1596
Lakhani SR, Ashworth A (2001) Microarray and histopathological analysis of tumours: the future and the past? Nat Rev Cancer 1:151–157
Leegwater PA, Pronk JC, van der Knaap MS (2003) Leukoencephalopathy with vanishing white matter: from magnetic resonance imaging pattern to five genes. J Child Neurol 18:639–645
Ma XJ, Salunga R, Tuggle JT, Gaudet J, Enright E, McQuary P, Payette T, Pistone M, Stecker K, Zhang BM, Zhou YX, Varnholt H, Smith B, Gadd M, Chatfield E, Kessler J, Baer TM, Erlander MG, Sgroi DC (2003) Gene expression profiles of human breast cancer progression. Proc Natl Acad Sci USA 100:5974–5979
McCaig C, Perks CM, Holly JM (2002) Intrinsic actions of IGFBP-3 and IGFBP-5 on Hs578T breast cancer epithelial cells: inhibition or accentuation of attachment and survival is dependent upon the presence of fibronectin. J Cell Sci 115:4293–4303
Nagahata T, Onda M, Emi M, Nagai H, Tsumagar K, Fujimoto T, Hirano A, Sato T, Nishikawa K, Akiyama F, Sakamoto G, Kasumi F, Miki Y, Tanaka T, Tsunoda T (2004) Expression profiling to predict postoperative prognosis for estrogen receptor-negative breast cancers by analysis of 25,344 genes on a cDNA microarray. Cancer Sci 95:218–225
Olofsson A, Willén H, Göransson M, Engström K, Meis-Kindblom JM, Stenman G, Kindblom LG, Åman P (2004) Abnormal expression of cell cycle regulators in FUS-CHOP carrying liposarcomas. Int J Oncol 25:1349–1355
Onda M, Emi M, Nagai H, Nagahata T, Tsumagari K, Fujimoto T, Akiyama F, Sakamoto G, Makita M, Kasumi F, Miki Y, Tanaka T, Tsunoda T, Nakamura Y (2004) Gene expression patterns as marker for 5-year postoperative prognosis of primary breast cancers. J Cancer Res Clin Oncol 130:537–545
Perou CM, Jeffrey SS, van de Rijn M, Rees CA, Eisen MB, Ross DT, Pergamenschikov A, Williams CF, Zhu SX, Lee JCF, Lashkari D, Shalon D, Brown PO, Botstein D (1999) Distinctive gene expression patterns in human mammary epithelial cells and breast cancers. Proc Natl Acad Sci USA 96:9212–9217
Richardson JP, Mohammad SS, Pavitt GD (2004) Mutations causing childhood ataxia with central nervous system hypomyelination reduce eukaryotic initiation factor 2B complex formation and activity. Mol Cell Biol 24:2352–2363
Singh D, Febbo PG, Ross K, Jackson DG, Manola J, Ladd C, Tamayo P, Renshaw AA, D’Amicl AV, Riche JP, Lander ES, Loda M, Kantoff PW, Golub TR, Sellers WR (2002) Gene expression correlates of clinical prostate cancer behavior. Cancer Cell 1:203–209
Sørlie T, Perou CM, Tibshirani R, Aas T, Geisler S, Johnsen H, Hastie T, Eisen MB, van de Rijn M, Jeffrey SS, Thorsen T, Quist H, Matese JC, Brown PO, Botstein D, Lønning PE, Børresen-Dale AL (2001) Gene expression patterns of breast carcinomas distinguish tumor subclasses with clinical implication. Proc Natl Acad Sci USA 98:10869–10874
Sørlie T, Tibshirani R, Parker J, Hastie T, Marron JS, Nobel A, Deng S, Johnsen H, Pesich R, Geisler S, Demeter J, Perou CM, Lønning PE, Brown PO, Børresen-Dale AL, Botstein D (2003) Repeated observation of breast tumor subtypes in independent gene expression data sets. Proc Natl Acad Sci USA 600:8418–8423
Sotiriou C, Neo SY, McShane LM, Korn EL, Long PM, Jazaeri A, Martiat P, Fox SB, Harris AL, Liu ET (2003) Breast cancer classification and prognosis based on gene expression profiles from a population-based study. Proc Natl Acad Sci USA 600:10393–10398
Subrahmanyam YVBK, Yamaga S, Prashar Y, Lee HH, Hoe NP, Kluger Y, Gerstein M, Goguen JD, Newburger PE, Weissman SM (2001) RNA expression patterns change dramatically in human neutrophils exposed to bacteria. Blood 97:2457–2468
Swami S, Raghavachari N, Muller UR, Bao YP, Feldman D (2003) Vitamin D growth inhibition of breast cancer cells: gene expression patterns assessed by cDNA microarray. Breast Cancer Res Treat 80:49–62
van de Vijver MJ, He YD, van’t Veer LJ, Dai H, Hart AAM, Voskuil DW, Schreiber GJ, Peterse JL, Roberts C, Marton MJ, Parrish M, Atsma D, Witteveen A, Glas A, Delahaye L, van Der Velde T, Bartelink H, Rodenhuis S, Rutgers ET, Friend SH, Bernards R (2002) A gene-expression signature as a predictor of survival in breast cancer. N Engl J Med 347:1999–2009
van’t Veer LJ, Dai H, van de Vijver M, He YD, Har AAM, Mao M, Peterse HL, van der Kooy K, Marton MJ, Witteveen AT, Schreiber GJ, Kerkhoven RM, Roberts C, Linsley PS, Bernards R, Friend SH (2002) Gene expression profiling predicts clinical outcome of breast cancer. Nature 415:530–536
Virtanen C, Ishikawa Y, Honjoh D, Kimura M, Shimane M, Miyoshi T, Nomura H, Jones MH (2002) Integrated classification of lung tumors and cell lines by expression profiling. Proc Natl Acad Sci USA 99:12357–12362
Wang E, Miller LD, Ohnmacht GA, Mocellin S, Perez-Diez A, Petersen D, Zhao Y, Simon R, Powell JI, Asaki E, Alexander HR, Duray PH, Herlyn M, Restifo NP, Liu ET, Rosenberg SA, Marincola FM (2002) Prospective molecular profiling of melanoma metastases suggests classifiers of immune responsiveness. Cancer Res 62:3581–3586
Wang Y, Klijin JGM, Zhang Y, Sieuwerts AM, Look MP, Yang F, Talantov D, Timmermans M, Meijer-van Gelder ME, Yu J, Jatkoe T, Berns EMJJ, Atkins D, Foekens JA (2005) Gene-expression profiles to predict distant metastasis of lymph-node-negative primary breast cancer. Lancet 365:671–679
West M, Blanchette C, Dressman H, Huang E, Ishida S, Spang R, Zuzan H, Olson JA Jr, Marks JR, Nevin JR (2001) Predicting the clinical status of human breast cancer by using gene expression profiles. Proc Natl Acad Sci USA 98:11462–11467
Yeoh EJ, Ross ME, Shurtleff SA, Williams WK, Patel D, Mahfouz R, Behm FG, Raimondi SC, Relling MV, Patel A, Cheng C, Campana D, Wilkins D, Zhou X, Li J, Liu H, Pui CH, Evans WE, Naeve C, Wong L, Downing JR (2002) Classification, subtype discovery, and prediction of outcome in pediatric acute lymphoblastic leukemia by gene expression profiling. Cancer Cell 1:133–143
Yeung KY, Bumgarner RE, Raftery AE (2005) Bayesian model averaging: development of an improved multi-class, gene selection and classification tool for microarray data. Bioinformatics 21:2394–2402
Zucman-Rossi J, Batzer MA, Stoneking M, Delattre O, Thomas G (1997) Interethnic polymorphism of EWS intron 6: genome plasticity mediated by Alu retroposition and recombination. Hum Genet 99:357–363
Acknowledgment
We are indebted to Hongyue Dai in Rosetta Inpharmatics for his helpful suggestion and revising of the manuscript.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Jiang, D., Zhao, N. A clinical prognostic prediction of lymph node-negative breast cancer by gene expression profiles. J Cancer Res Clin Oncol 132, 579–587 (2006). https://doi.org/10.1007/s00432-006-0108-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00432-006-0108-6