Abstract
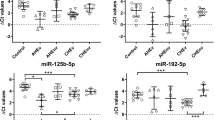
Hepatitis C virus (HCV) infection affects approximately 3 % of the world population. HCV targets hepatic tissue, and most infected patients develop a chronic infection. Currently, studies have demonstrated an association between HCV-RNA replication and miR-122, the most abundant microRNA in the liver. Our aim was to evaluate liver and serum expression of miR-122 in patients infected with HCV genotypes 1 and 3, and to identify possible associations between miR-122 expression and lipid profiles, HCV viral load, apolipoproteins and liver enzymes. MicroRNAs were isolated from blood and liver tissue, and miR-122 expression was quantified by real-time PCR. HCV viral load was quantified by real-time PCR and HCV genotype, and serum biomarkers were obtained from medical report. The levels of miR-122 were higher in liver than those in blood from individuals infected with HCV genotypes 1 and 3 (p < 0.0001). The tissue levels of miR-122 were higher in subjects infected with HCV genotype 3 (6.22-fold, p < 0.001). A positive correlation was observed between the blood and hepatic levels of miR-122 in patients infected with HCV genotype 1 (r = 0.302, p = 0.026); in these patients, an inverse correlation was observed between serum apolipoprotein A-II (ApoA-II) levels and the blood (r = −0.330; p = 0.014) and hepatic (r = −0.311; p = 0.020) levels of miR-122. In patients infected with HCV genotype 3, there was a positive correlation between the hepatic miR-122 and the high-density lipoprotein—HDL (r = 0.412, p = 0.036) and insulin (r = 0.478, p = 0.044). Lipid metabolism proteins and miR-122 expression levels have different relations in HCV-3- and HCV-1-infected patients.




Similar content being viewed by others
References
Shepard CW, Finelli L, Alter MJ (2005) Global epidemiology of hepatitis C virus infection. Lancet Infect Dis 5:558–567
Thomas DL (2013) Cure of hepatitis C virus infection without interferon alfa: scientific basis and current clinical evidence. Top Antivir Med 21:152–156
Cheng JC, Yeh YJ, Tseng CP, Hsu SD, Chang YL, Sakamoto N et al (2012) Let-7b is a novel regulator of hepatitis C virus replication. Cell Mol Life Sci 69:2621–2633
Jopling CL, Yi M, Lancaster AM, Lemon SM, Sarnow P (2005) Modulation of hepatitis C virus RNA abundance by a liver-specific MicroRNA. Science 309:1577–1581
Murakami Y, Aly HH, Tajima A, Inoue I, Shimotohno K (2009) Regulation of the hepatitis C virus genome replication by miR-199a. J Hepatol 50:453–460
Erson AE, Petty EM (2008) MicroRNAs in development and disease. Clin Genet 74:296–306
Chang J, Nicolas E, Marks D, Sander C, Lerro A, Buendia MA et al (2004) miR-122, a mammalian liver-specific microRNA, is processed from hcr mRNA and may downregulate the high affinity cationic amino acid transporter CAT-1. RNA Biol 1:106–113
Girard M, Jacquemin E, Munnich A, Lyonnet S, Henrion-Caude A (2008) miR-122, a paradigm for the role of microRNAs in the liver. J Hepatol 48:648–656
Hou W, Tian Q, Zheng J, Bonkovsky HL (2010) MicroRNA-196 represses Bach1 protein and hepatitis C virus gene expression in human hepatoma cells expressing hepatitis C viral proteins. Hepatology 51:1494–1504
Esau C, Davis S, Murray SF, Yu XX, Pandey SK, Pear M et al (2006) miR-122 regulation of lipid metabolism revealed by in vivo antisense targeting. Cell Metab 3:87–98
Rubbia-Brandt L, Quadri R, Abid K, Giostra E, Malé PJ, Mentha G et al (2000) Hepatocyte steatosis is a cytopathic effect of hepatitis C virus genotype 3. J Hepatol 33:106–115
Douam F, Lavillette D, Cosset FL (2015) The mechanism of HCV entry into host cells. Prog Mol Biol Transl Sci 129:63–107
Andre P, Komurian-Pradel F, Deforges S, Perret M, Berland JL, Sodoyer M et al (2002) Characterization of low- and very-low-density hepatitis C virus RNA-containing particles. J Virol 76:6919–6928
Gastaminza P, Cheng G, Wieland S, Zhong J, Liao W, Chisari FV (2008) Cellular determinants of hepatitis C virus assembly, maturation, degradation, and secretion. J Virol 82:2120–2129
Jiang J, Luo G (2009) Apolipoprotein E but not B is required for the formation of infectious hepatitis C virus particles. J Virol 83:12680–12691
Bedossa P, Poynard T (1996) An algorithm for the grading of activity in chronic hepatitis C. The METAVIR cooperative study group. Hepatology 24:289–293
Schmittgen TD, Livak KJ (2008) Analyzing real-time PCR data by the comparative C(T) method. Nat Protoc 3:1101–1108
Morita K, Taketomi A, Shirabe K, Umeda K, Kayashima H, Ninomiya M et al (2011) Clinical significance and potential of hepatic microRNA-122 expression in hepatitis C. Liver Int 31:474–484
Horie T, Baba O, Kuwabara Y, Yokode M, Kita T, Kimura T et al (2014) MicroRNAs and lipoprotein metabolism. J Atheroscler Thromb 21:17–22
Fukuhara T, Wada M, Nakamura S, Ono C, Shiokawa M, Yamamoto S et al (2014) Amphipathic α-helices in apolipoproteins are crucial to the formation of infectious hepatitis C virus particles. PLoS Pathog 10:e1004534
Sabile A, Perlemuter G, Bono F, Kohara K, Demaugre F, Kohara M et al (1999) Hepatitis C virus core protein binds to apolipoprotein AII and its secretion is modulated by fibrates. Hepatology 30:1064–1076
Shibata C, Kishikawa T, Otsuka M, Ohno M, Yoshikawa T, Takata A et al (2013) Inhibition of microRNA122 decreases SREBP1 expression by modulating suppressor of cytokine signaling 3 expression. Biochem Biophys Res Commun 438:230–235
Schaefer EA, Chung RT (2013) HCV and host lipids: an intimate connection. Semin Liver Dis 33:358–368
Hsu SH, Wang B, Kota J, Yu J, Costinean S, Kutay H et al (2012) Essential metabolic, anti-inflammatory, and anti-tumorigenic functions of miR-122 in liver. J Clin Invest 122:2871–2883
Tsai WC, Hsu SD, Hsu CS, Lai TC, Chen SJ, Shen R et al (2012) Micro-RNA 122 plays a critical role in liver homeostasis and hepatocarcinogenesis. J Clin Invest 122:2884–2897
Cermelli S, Ruggieri A, Marrero JA, Ioannou GN, Beretta L (2011) Circulating microRNAs in patients with chronic hepatitis C and non-alcoholic faty liver disease. PLoS one 6:e23937
Boštjančič E, Bandelj E, Luzar B, Poljak M, Glavač D (2015) Hepatic expression of miR-122, miR-126, miR-136 and miR-181ª and their correlation to histopathological and clinical characteristics of patients with hepatitis C. J Viral Hepat 22:146–157
Cheung O, Puri P, Eicken C, Contos MJ, Mirshahi F, Maher JW et al (2008) Nonalcoholic steatohepatitis is associated with altered hepatic MicroRNA expression. Hepatology 48:1810–1820
Jackel-Cram C, Qiao L, Xiang Z, Brownlie R, Zhou Y, Babiuk L et al (2010) Hepatitis C virus genotype-3a core protein enhances sterol regulatory element-binding protein-1 activity through the phosphoinositide 3-kinase–Akt-2 pathway. J Gen Virol 91:1388–1395
Abid K, Pazienza V, de Gottardi A, Rubbia-Brandt L, Conne B, Pugnale P et al (2005) An in vitro model of hepatitis C virus genotype 3a-associated triglycerides accumulation. J Hepatol 42:744–751
Hourioux C, Patient R, Morin A, Blanchard E, Moreau A, Trassard S et al (2007) The genotype 3-specific hepatitis C virus core protein residue phenylalanine 164 increases steatosis in an in vitro cellular model. Gut 56:1302–1308
Pazienza V, Clément S, Pugnale P, Conzelmann S, Pascarella S, Mangia A et al (2009) Gene expression profile of Huh-7 cells expressing hepatitis C virus genotype 1b or 3a core proteins. Liver Int 29:661–669
Piodi A, Chouteau P, Lerat H, Hezode C, Pawlotsky JM (2008) Morphological changes in intracellular lipid droplets induced by different hepatitis C virus genotype core sequences and relationship with steatosis. Hepatology 48:16–27
Acknowledgments
FAPESP São Paulo Research Foundation (2012/50332-7 and 2012/15706-3) and Alves de Queiroz Family Fund for Research. João R. R. Pinho receives fellowship from CNPq (Bolsista de Produtividade em Pesquisa do CNPq—Nível 2).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Rights and permissions
About this article
Cite this article
Oliveira, K.G., Malta, F.M., Nastri, A.C.S.S. et al. Increased hepatic expression of miRNA-122 in patients infected with HCV genotype 3. Med Microbiol Immunol 205, 111–117 (2016). https://doi.org/10.1007/s00430-015-0431-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00430-015-0431-0