Abstract
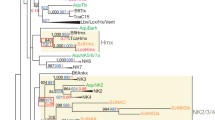
Sponges are among the earliest diverging lineage within the metazoan phyla. Although their adult morphology is distinctive, at several stages of development, they possess characteristics found in more complex animals. The T-box family of transcription factors is an evolutionarily ancient gene family known to be involved in the development of structures derived from all germ layers in the bilaterian animals. There is an incomplete understanding of the role that T-box transcription factors play in normal sponge development or whether developmental pathways using the T-box family share similarities between parazoan and eumetazoan animals. To address these questions, we present data that identify several important T-box genes in marine and freshwater sponges, place these genes in a phylogenetic context, and reveal patterns in how these genes are expressed in developing sponges. Phylogenetic analyses demonstrate that sponges have members of at least two of the five T-box subfamilies (Brachyury and Tbx2/3/4/5) and that the T-box genes expanded and diverged in the poriferan lineage. Our analysis of signature residues in the sponge T-box genes calls into question whether “true” Brachyury genes are found in the Porifera. Expression for a subset of the T-box genes was elucidated in larvae from the marine demosponge, Halichondria bowerbanki. Our results show that sponges regulate the timing and specificity of gene expression for T-box orthologs across larval developmental stages. In situ hybridization reveals distinct, yet sometimes overlapping expression of particular T-box genes in free-swimming larvae. Our results provide a comparative framework from which we can gain insights into the evolution of developmentally important pathways.




Similar content being viewed by others
References
Abascal F, Zardoya R, Posada D (2005) ProtTest: selection of best-fit models of protein evolution. Bioinformatics 21:2104–2105
Adell T, Müller WEG (2005) Expression pattern of the Brachyury and Tbx2 homologues from the sponge Suberites domuncula. Biol Cell 97:641–650
Adell T, Grebenjuk VA, Wiens M, Müller WEG (2003) Isolation and characterization of two T-box genes from sponges, the phylogenetically oldest metazoan taxon. Dev Genes Evol 213:421–434
Bielen H, Oberleitner S, Marcellini S, Gee L, Lemaire P, Bode HR, Rupp R, Technau U (2007) Divergent functions of two ancient Hydra Brachyury paralogues suggest specific roles for their C-terminal domains in tissue fate induction. Development 134:4187–4197
Curtis KM, Gomez LA, Rios C, Garbayo E, Raval AP, Perez-Pinzon MA, Schiller PC (2010) EF1a and RPL13a represent normalization genes suitable for RT-qPCR analysis of bone marrow derived mesenchymal stem cells. BMC Mol Biol 11:61
Dereeper A, Guignon V, Blanc G, Audic S, Buffet S, Chevenet F, Dufayard JF, Guindon S, Lefort V, Lescot M, Claverie JM, Gascuel O (2008) Phylogeny.fr: robust phylogenetic analysis for the non-specialist. Nuc Acid Res 36:465–469
Edgar RC (2004) MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res 32:1792–1797
Fell P, Jacob W (1979) Reproduction and development of Halichondria sp. in the Mystic estuary, Connecticut. Biol Bull 155:62–75
Felsenstein J (2004) Inferring phylogenies. Sinauer, Sunderland
Funayama N, Nakatsukasa M, Hayashi T, Agata K (2005) Isolation of the choanocyte in the fresh water sponge, Ephydatia fluviatilis and its lineage marker, Ef annexin. Dev Growth Differ 47:243–253
Guindon S, Gascuel O (2003) A simple, fast, and accurate algorithm to estimate large phylogenies by Maximum Likelihood. Syst Biol 52:696–704
Herrmann BG, Labeit S, Poustka A, King TR, Lehrach H (1990) Cloning of the T gene required for mesodermal formation in the mouse. Nature 343:617–622
Hill A, Boll W, Ries C, Warner L, Osswalt M, Hill M, Noll M (2010) Origin of Pax and Six gene families in sponges: single PaxB and Six1/2 orthologs in Chalinula loosanoffi. Dev Biol 343:106–123
Holland PW, Koschorz B, Holland LZ, Herrmann BG (1995) Conservation of Brachyury (T) genes in amphioxus and vertebrates: developmental and evolutionary implications. Development 121:4283–4291
Horton N, Mahadevan NR, Minguillon C, Osoegawa K, Rokhsar DS, Ruvinsky I, de Jong PJ, Logan MP, Gibson-Brown JJ (2008) Conservation of linkage and evolution of developmental function within the Tbx2/3/4/5 subfamily of T-box genes: implications for the origin of vertebrate limbs. Dev Genes Evol 218:613–628
Huelsenbeck JP, Ronquist F (2001) MRBAYES: Bayesian inference of phylogeny. Bioinformatics 17:754–755
King N, Westbrook MJ, Young SL, Kuo A, Abedin M, Chapman J, Fairclough S, Hellsten U, Isogai Y, Letunic I, Marr M, Pincus D, Putnam N, Rokas A, Wright KJ, Zuzow R, Dirks W, Good M, Goodstein D, Lemons D, Li W, Lyons JB, Morris A, Nichols S, Richter DJ, Salamov A, Sequencing JGI, Bork P, Lim WA, Manning G, Miller WT, McGinnis W, Shapiro H, Tjian R, Grigoriev IV, Rokhsar D (2008) The genome of the choanoflagellate Monosiga brevicollis and the origin of metazoans. Nature 451:783–788
Larroux C, Fahey B, Liubicich D, Hinman VF, Gauthier M, Gongora M, Green K, Worheide G, Leys S, Degnan BM (2006) Developmental expression of transcription factor genes in a demosponge: insights into the origin of metazoan multicellularity. Evol Dev 8(2):150–173
Larroux C, Luke GN, Koopman P, Rokhsar D, Shimeld SM, Degnan BM (2008) Genesis and expansion of metazoan transcription factor gene classes. Mol Biol Evol 25(5):980–996
Leys SP (2004) Gastrulation in sponges. In: Stern C (ed) Gastrulation. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, pp 23–31
Manuel M, Le Parco Y, Borchiellini C (2004) Comparative analysis of Brachyury T-domains, with the characterization of two new sponge sequences from a hexactinellid and a calcisponge. Gene 340:291–301
Martinelli C, Spring J (2003) Distinct expression patterns of the two T-box homologues Brachyury and Tbx2/3 in the placozoan Trichoplax adhaerens. Dev Genes Evol 231:492–499
Martinelli C, Spring J (2005) T-box and homeobox genes from ctenophore Pleurobrachia pileus: comparison of Brachyury, Tbx2/3 and Tlx in basal metazoans and bilaterians. FEBS Lett 579:5024–5028
McCurley AT, Callard GV (2008) Characterization of housekeeping genes in zebrafish: male–female differences and effects of tissue type, developmental stage and chemical treatment. BMC Mol Biol 9:102
Mikhailov KV, Konstantinova AV, Nikitin MA, Troshin PV, Rusin LY, Lyubetsky VA, Panchin YV, Mylnikov AP, Moroz LL, Kuman S, Aleoshin VV (2009) The origin of Metazoa: a transition from temporal to spatial cell differentiation. Bioessays 31:758–768
Müller CW, Herrmann BG (1997) Crystallographic structure of the T domain-DNA complex of the Brachyury transcription factor. Nature 289:884–888
Naiche LA, Harrelson Z, Kelly RG, Papaioannou VE (2005) T-box genes in vertebrate development. Annu Rev Genet 39:219–239
Nichols SA, Dirks W, Pearse JS, King N (2006) Early evolution of animal cell signaling and adhesion genes. Proc Natl Acad Sci USA 103:12451–12456
Papaioannou VE (2001) T-box genes in development: from hydra to humans. Int Rev Cytol 207:1–70
Pick KS, Philippe H, Schreiber F, Erpenbeck D, Jackson DJ, Wrede P, Wiens M, Alié A, Morgenstern B, Manuel M, Wörheide G (2010) Improved phylogenomic taxon sampling noticeably affects non-bilaterian relationships. Mol Biol Evol 27(9):1983–1987
Putnam NH, Srivastava M, Hellsten U, Dirks B, Chapman J, Salamov A, Terry A, Shapiro H, Lindquist E, Kapitonov VV, Jurka J, Genikhovick G, Grigoriev IV, Lucas SM, Steele RE, Finnerty JR, Technau U, Martindale MQ, Rokhsar DS (2007) Sea anemone genome reveals ancestral eumetazoan gene repertoire and genomic organization. Science 317:86–94
Rodriguez-Lanetty M, Phillips WS, Dove S, Hoegh-Guldberg O, Weis VM (2008) Analytical approach for selecting normalizing genes from a cDNA microarray platform to be used in q-RT-PCR assays: a cnidarian case study. J Biochem Biophys Methods 70:985–991
Sakarya O, Armstrong KA, Adamska M, Adamski M, Wang IF, Tidor B, Degnan BM, Oakley KKS (2007) A post-synaptic scaffold at the origin of the animal kingdom. PLoS ONE 2:e506
Scholz CB, Technau U (2003) The ancestral role of Brachyury: expression of NemBra1 in the basal cnidarian Nematostella vectensis (Anthozoa). Dev Genes Evol 212:563–570
Sempere LF, Cole CN, McPeek MA, Peterson KJ (2006) The phylogenetic distribution of metazoan microRNAs: insights into evolutionary complexity and constraint. J Exp Zool B Mol Dev Evol 306(6):575–88
Showell C, Binder O, Conlon F (2004) T-box genes in early embryogenesis. Dev Dyn 229:201–218
Siah A, Dohoo C, McKenna P, Delaporte M, Berthe FCJ (2008) Selecting a set of housekeeping genes for quantitative real-time PCR in normal and tetraploid haemocytes of soft-shell clams, Mya arenaria. Fish Shellfish Immunol 25:202–207
Simpson TL (1984) The cell biology of sponges. Springer, New York
Sperling EA, Peterson KJ, Pisani D (2009) Phylogenetic-signal dissection of nuclear housekeeping genes supports the paraphyly of sponges and the monophyly of Eumetazoa. Mol Biol Evol 26(10):2261–2274
Spring J, Yanze N, Josch C, Middel AM, Winninger B, Schmid V (2002) Conservation of Brachyury, Mef2 and snail in the myogenic lineage of jellyfish: a connection to the mesoderm of Bilateria. Dev Biol 244:372–384
Srivastava M, Begovic E, Chapman J et al (2008) The Trichoplax genome and the nature of placozoans. Nature 454:955–960
Stern C (2004) Gastrulation: from cells to embryo. Cold Spring Harbor Press, Cold Spring Harbor
Strekal TA, McDiffett W (1974) Factors affecting germination, growth, and distribution of the freshwater sponge, Spongilla fragilis Leidy (Porifera). Biol Bull 146:267–278
Technau U, Bode HR (1999) HyBra1, a Brachyury homologue, acts during head formation in Hydra. Development 126:999–1010
Wilson V, Conlon F (2002) The T-box family. Genome Biol 3:3008.1–3008.7
Yamada A, Pang K, Martindale MQ, Tochinai S (2007) Surprisingly complex T-box gene complement in diploblastic metazoans. Evol Dev 9(3):220–230
Yamada A, Martindale MQ, Fukui A, Tochinai S (2010) Highly conserved functions of the Brachyury gene on morphogenetic movements: insight from the early-diverging phylum Ctenophora. Dev Biol 339:212–222
Yüzbaşıoğlu A, Onbaşılar İ, Kocaefe Ç, Özgüç M (2010) Assessment of housekeeping genes for use in normalization of real time PCR in skeletal muscle with chronic degenerative changes. Exp Mol Pathol 88:326–329
Acknowledgements
This work was supported by a grant from the Thomas F. and Kate Miller Jeffress Memorial Trust to A.H. and a Beckman Scholars Program award to the University of Richmond. Special thanks to Scott Nichols for providing T-box sequences for O. carmela.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by M. Martindale
Electronic supplementary materials
Below is the link to the electronic supplementary material.
ESM 1
(DOCX 423 kb)
ESM 2
(DOC 25 kb)
ESM 3
In situ hybridization to illustrate positive controls using a Hb Bar/Bsh probe with cross-section shown in insert where hybridization is specific to inner cell mass. b Hb Actin probe hybridizes throughout larvae (GIF 146 kb)
ESM 3
High Resolution Image (TIFF 7483 kb)
Rights and permissions
About this article
Cite this article
Holstien, K., Rivera, A., Windsor, P. et al. Expansion, diversification, and expression of T-box family genes in Porifera. Dev Genes Evol 220, 251–262 (2010). https://doi.org/10.1007/s00427-010-0344-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00427-010-0344-2