Abstract
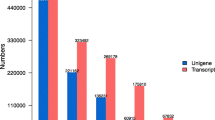
Antarctic hairgrass (Deschampsia antarctica Desv.) is the only natural grass species in the maritime Antarctic. It has been studied as an extremophile that has successfully adapted to marginal land with the harshest environment for terrestrial plants. However, limited genetic research has focused on this species due to the lack of genomic resources. Here, we present the first de novo assembly of its transcriptome by massive parallel sequencing and its expression profile using D. antarctica grown under various stress conditions. Total sequence reads generated by pyrosequencing were assembled into 60,765 unigenes (28,177 contigs and 32,588 singletons). A total of 29,173 unique protein-coding genes were identified based on sequence similarities to known proteins. The combined results from all three stress conditions indicated differential expression of 3,110 genes. Quantitative reverse transcription polymerase chain reaction showed that several well-known stress-responsive genes encoding late embryogenesis abundant protein, dehydrin 1, and ice recrystallization inhibition protein were induced dramatically and that genes encoding U-box-domain-containing protein, electron transfer flavoprotein-ubiquinone, and F-box-containing protein were induced by abiotic stressors in a manner conserved with other plant species. We identified more than 2,000 simple sequence repeats that can be developed as functional molecular markers. This dataset is the most comprehensive transcriptome resource currently available for D. antarctica and is therefore expected to be an important foundation for future genetic studies of grasses and extremophiles.




Similar content being viewed by others
Abbreviations
- AOX:
-
Alternative oxidase
- DEG:
-
Differentially expressed gene
- DHN:
-
Dehydrin
- EC:
-
Enzyme commission
- EST:
-
Expressed sequence tags
- GC:
-
Guanine–cytosine
- GI:
-
GIGANTEA
- GO:
-
Gene ontology
- IRIP:
-
Ice recrystallization inhibition protein
- IVD:
-
Isovaleryl Co-A dehydrogenase
- KEGG:
-
Kyoto Encyclopedia of Genes and Genomes
- LEA:
-
Late embryogenesis abundant protein
- PEPC:
-
Phosphoenolpyruvate carboxylase
- ProDH:
-
Proline dehydrogenase
- qPCR:
-
Quantitative reverse transcription polymerase chain reaction
- SSR:
-
Simple sequence repeat
- TIM:
-
Inner membrane translocase subunit
References
Alberdi M, Bravo LA, Gutierrez A, Gidekel M, Corcuera LJ (2002) Ecophysiology of Antarctic vascular plants. Physiol Plant 115:479–486
Bravo LA, Griffith M (2005) Characterization of antifreeze activity in Antarctic plants. J Exp Bot 56:1189–1196
Bravo LA, Ulloa N, Zuñiga GE, Casanova A, Corcuera LJ, Alberdi M (2001) Cold resistance in Antarctic angiosperms. Physiol Plant 111:55–65
Cao S, Ye M, Jiang S (2005) Involvement of GIGANTEA gene in the regulation of the cold stress response in Arabidopsis. Plant Cell Rep 24:683–690
Chou HH, Holmes MH (2001) DNA sequence quality trimming and vector removal. Bioinformatics 17:1093–1104
Chwedorzewska KJ (2009) Terrestrial Antarctic ecosystems in the changing world: an overview. Pol Polar Res 30:263–276
Conesa A, Götz S, García-Gómez JM, Terol J, Talón M, Robles M (2005) Blast2GO: a universal tool for annotation, visualization and analysis in functional genomics research. Bioinformatics 21:3674–3676
Dalchau N, Baek SJ, Briggs HM, Robertson FC, Dodd AN, Gardner MJ, Stancombe MA, Haydon MJ, Stan GB, Goncalves JM, Webb AA (2011) The circadian oscillator gene GIGANTEA mediates a long-term response of the Arabidopsis thaliana circadian clock to sucrose. Proc Natl Acad Sci USA 108:5104–5109
Däschner K, Couée I, Binder S (2001) The mitochondrial isovaleryl-coenzyme A dehydrogenase of Arabidopsis oxidizes intermediates of leucine and valine catabolism. Plant Physiol 126:601–612
Du Z, Zhou X, Ling Y, Zhang Z, Su Z (2010) agriGO: a GO analysis toolkit for the agricultural community. Nucleic Acids Res 38:W64–W70
Edgington ES (1995) Randomization tests, 3rd edn. Marcel-Dekker, New York
Edward JA, Lewis Smith RI (1988) Photosynthesis and respiration of Colobanthus quitensis and Deschampsia antarctica from the maritime Antarctic. BAS Bull 81:43–63
Gielwanowska I, Szczuka E, Bednara J, Gorecki R (2005) Anatomical features and ultrastructure of Deschampsia antarctica (Poaceae) leaves from different growing habitats. Ann Bot 96:1109–1119
Gill SS, Tuteja N (2010) Reactive oxygen species and antioxidant machinery in abiotic stress tolerance in crop plants. Plant Physiol Biochem 48:909–930
González MC, Sánchez R, Cejudo FJ (2003) Abiotic stresses affecting water balance induce phosphoenolpyruvate carboxylase expression in roots of wheat seedlings. Planta 216:985–992
Hong S-Y, Lee S, Seo P, Yang M-S, Park C-M (2010) Identification and molecular characterization of a Brachypodium distachyon GIGANTEA gene: functional conservation in monocot and dicot plants. Plant Mol Biol 72:485–497
Iordachescu M, Imai R (2008) Trehalose biosynthesis in response to abiotic stresses. J Integr Plant Biol 50:1223–1229
John UP, Polotnianka RM, Sivakumaran KA, Chew O, Mackin L, Kuiper MJ, Talbot JP, Nugent GD, Mautord J, Schrauf GE, Spangenberg GC (2009) Ice recrystallization inhibition proteins (IRIPs) and freeze tolerance in the cryophilic Antarctic hair grass Deschampsia antarctica E. Desv. Plant Cell Environ 32:336–348
Kajala K, Covshoff S, Karki S, Woodfield H, Tolley BJ, Dionora MJA, Mogul RT, Mabilangan AE, Danila FR, Hibberd JM, Quick WP (2011) Strategies for engineering a two-celled C4 photosynthetic pathway into rice. J Exp Bot 62:3001–3010
Kanehisa M, Goto S (2000) KEGG: Kyoto encyclopedia of genes and genomes. Nucleic Acids Res 28:27–30
Kreps J, Wu Y, Chang H, Zhu T, Wang X, Harper J (2002) Transcriptome changes for Arabidopsis in response to salt, osmotic, and cold stress. Plant Physiol 130:2129–2141
Lara A, Pérez-Trabado G, Villalobos D, Díaz-Moreno S, Cantón F, Claros M (2007) A web tool to discover full-length sequences: full-lengther. In: Corchado E, Corchado J, Abraham A (eds) Innovations hybrid intelligent systems. Springer, Berlin, pp 361–368
Lee H, Cho HH, Kim I-C, Yim JH, Lee HK, Lee YK (2008) Expressed sequence tag analysis of Antarctic hairgrass Deschampsia antarctica from King George Island, Antarctica. Mol Cells 25:258–264
Lee H, Kim JH, Park M, Kim I-C, Yim JH, Lee HK (2010) Reference genes validation for qPCR normalization in Deschampsia antarctica during abiotic stresses. Antarct Sci 22:477–484
Lewis Smith RI (1994) Vascular plants as bioindicators of regional warming in Antarctica. Oecologia 99:322–328
Lister R, Chew O, Lee MN, Heazlewood JL, Clifton R, Parker KL, Millar AH, Whelan J (2004) A transcriptomic and proteomic characterization of the Arabidopsis mitochondrial protein import apparatus and its response to mitochondrial dysfunction. Plant Physiol 134:777–789
Margulies M, Egholm M, Altman WE, Attiya S, Bader JS, Bemben LA, Berka J, Braverman MS, Chen YJ, Chen Z, Dewell SB, Du L, Fierro JM, Gomes XV, Godwin BC, He W, Helgesen S, Ho CH, Irzyk GP, Jando SC, Alenquer ML, Jarvie TP, Jirage KB, Kim JB, Knight JR, Lanza JR, Leamon JH, Lefkowitz SM, Lei M, Li J, Lohman KL, Lu H, Makhijani VB, McDade KE, McKenna MP, Myers EW, Nickerson E, Nobile JR, Plant R, Puc BP, Ronan MT, Roth GT, Sarkis GJ, Simons JF, Simpson JW, Srinivasan M, Tartaro KR, Tomasz A, Vogt KA, Volkmer GA, Wang SH, Wang Y, Weiner MP, Yu P, Begley RF, Rothberg JM (2005) Genome sequencing in microfabricated high-density picolitre reactors. Nature 437:376–380
Michael BE, Kaufmann MR (1973) The osmotic potential of polyethylene glycol 6000. Plant Physiol 51:914–916
Morgante M, Hanafey M, Powell W (2002) Microsatellites are preferentially associated with nonrepetitive DNA in plant genomes. Nat Genet 30:194–200
Nanjo T, Kobayashi M, Yoshiba Y, Kakubari Y, Yamaguchi-Shinozaki K, Shinozaki K (1999) Antisense suppression of proline degradation improves tolerance to freezing and salinity in Arabidopsis thaliana. FEBS Lett 461:205–210
Novaes E, Drost DR, Farmerie WG, Pappas GJ Jr, Grattapaglia D, Sederoff RR, Kirst M (2008) High-throughput gene and SNP discovery in Eucalyptus grandis, an uncharacterized genome. BMC Genomics 9:312
O’Leary B, Park J, Plaxton WC (2011) The remarkable diversity of plant PEPC (phosphoenolpyruvate carboxylase): recent insights into the physiological functions and post-translational controls of non-photosynthetic PEPCs. Biochem J 436:15–34
Ozturk ZN, Talamé V, Deyholos M, Michalowski CB, Galbraith DW, Gozukirmizi N, Tuberosa R, Bohnert HJ (2002) Monitoring large-scale changes in transcript abundance in drought- and salt- stressed barley. Plant Mol Biol 48:551–573
Parchman TL, Geist KS, Grahnen JA, Benkman CW, Buerkle CA (2010) Transcriptome sequencing in an ecologically important tree species: assembly, annotation, and marker discovery. BMC Genomics 11:180
Peng JH, Lapitan NL (2005) Characterization of EST-derived microsatellites in the wheat genome and development of eSSR markers. Funct Integr Genomics 5:80–96
Pérez-Torres E, Bascuñán L, Sierra A, Bravo L, Corcuera L (2006) Robustness of activity of Calvin cycle enzymes after high light and low temperature conditions in Antarctic vascular plants. Polar Biol 29:909–916
Pertea G, Huang X, Liang F, Antonescu V, Sultana R, Karamycheva S, Lee Y, White J, Cheung F, Parvizi B, Tsai J, Quackenbush J (2003) TIGR Gene Indices clustering tools (TGICL): a software system for fast clustering of large EST datasets. Bioinformatics 19:651–652
Raines CA (2011) Increasing photosynthetic carbon assimilation in C3 plants to improve crop yield: current and future strategies. Plant Physiol 155:36–42
Rodriguez-Penagos M, Munoz-Clares R (1999) Response of phosphoenolpyruvate carboxylase from maize leaves to moderate water deficit. J Plant Physiol 155:631–638
Rothberg JM, Leamon JH (2008) The development and impact of 454 sequencing. Nat Biotechnol 26:1117–1124
Salvucci ME, Crafts-Brandner SJ (2004) Relationship between the heat tolerance of photosynthesis and the thermal stability of rubisco activase in plants from contrasting thermal environments. Plant Physiol 134:1460–1470
Sánchez R, Flores A, Cejudo FJ (2006) Arabidopsis phosphoenolpyruvate carboxylase genes encode immunologically unrelated polypeptides and are differentially expressed in response to drought and salt stress. Planta 223:901–909
Seki M, Narusaka M, Ishida J, Nanjo T, Fujita M, Oono Y, Kamiya A, Nakajima M, Enju A, Sakurai T, Satou M, Akiyama K, Taji T, Yamaguchi-Shinozaki K, Carninci P, Kawai J, Hayashizaki Y, Shinozaki K (2002) Monitoring the expression profiles of 7000 Arabidopsis genes under drought, cold and high-salinity stresses using a full-length cDNA microarray. Plant J 31:279–292
Shinozaki K, Yamaguchi-Shinozaki K, Seki M (2003) Regulatory network of gene expression in the drought and cold stress responses. Curr Opin Plant Biol 6:410–417
Storey JD (2002) A direct approach to false discovery rates. J R Stat Soc B 64:479–498
Thiel T, Michalek W, Varshney RK, Graner A (2003) Exploiting EST databases for the development and characterization of gene-derived SSR-markers in barley (Hordeum vulgare L.). Theor Appl Genet 106:411–422
Van Aken O, Giraud E, Clifton R, Whelan J (2009) Alternative oxidase: a target and regulator of stress responses. Physiol Plant 137:354–361
Vera JC, Wheat CW, Fescemyer HW, Frilander MJ, Crawford DL, Hanski I, Marden JH (2008) Rapid transcriptome characterization for a nonmodel organism using 454 pyrosequencing. Mol Ecol 17:1636–1647
Verslues PE, Sharma S (2010) Proline metabolism and its implications for plant-environment interaction. Arabidopsis Book 8:e0140
Vidal J, Chollet R (1997) Regulatory phosphorylation of C4 PEP carboxylase. Trends Plant Sci 2:230–237
Wang H-C, Hickey D (2007) Rapid divergence of codon usage patterns within the rice genome. BMC Evol Biol 7:S6
Wang Z, Gerstein M, Snyder M (2009) RNA-Seq: a revolutionary tool for transcriptomics. Nat Rev Genet 10:57–63
Welin BV, Olson A, Nylander M, Palva ET (1994) Characterization and differential expression of dhn/lea/rab-like genes during cold acclimation and drought stress in Arabidopsis thaliana. Plant Mol Biol 26:131–144
Wilhelm BT, Landry J-R (2009) RNA-Seq—quantitative measurement of expression through massively parallel RNA-sequencing. Methods 48:249–257
Wong CE, Li Y, Labbe A, Guevara D, Nuin P, Whitty B, Diaz C, Golding GB, Gray GR, Weretilnyk EA, Griffith M, Moffatt BA (2006) Transcriptional profiling implicates novel interactions between abiotic stress and hormonal responses in Thellungiella, a close relative of Arabidopsis. Plant Physiol 140:1437–1450
Xiong FS, Mueller EC, Day TA (2000) Photosynthetic and respiratory acclimation and growth response of Antarctic vascular plants to contrasting temperature regimes. Am J Bot 87:700–710
Xiong L, Schumaker KS, Zhu J-K (2002) Cell signaling during cold, drought, and salt stress. Plant Cell 14:S165–S183
Zhou S, Sauvé R, Thannhauser TW (2009) Proteome changes induced by aluminium stress in tomato roots. J Exp Bot 60:1849–1857
Zhu JK (2002) Salt and drought stress signal transduction in plants. Annu Rev Plant Biol 53:247–273
Zuñiga GE, Alberdi M, Corcuera LJ (1996) Non-structural carbohydrates in Deschampsia antarctica Desv. from South Shetland Islands, maritime Antarctic. Environ Exp Bot 36:393–399
Acknowledgments
This work was supported by Functional Genomics on Polar Organisms grant (PE12020) funded by Korea Polar Research Institute (KOPRI).
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Lee, J., Noh, E.K., Choi, HS. et al. Transcriptome sequencing of the Antarctic vascular plant Deschampsia antarctica Desv. under abiotic stress. Planta 237, 823–836 (2013). https://doi.org/10.1007/s00425-012-1797-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00425-012-1797-5