Abstract
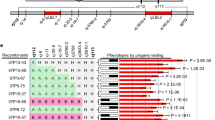
A gene underlying a quantitative trait locus (QTL) controlling plant height on chromosome 1 (QTLph1) in rice (Oryza sativa L.) was identified using the candidate-gene strategy. First, the function of a targeted gene was analyzed using near isogenic lines (NILs) in which the chromosomal region of a targeted QTL was substituted with that of another line. Second, for physiological information, the candidate gene was selected in the annotation data by the genome sequencing. Physiological analyses of an NIL-expressing QTLph1 (NIL6) suggested that the targeted gene controls plant height by enabling higher amounts of sucrose to be translocated in leaves. The results indicated that the gene for sucrose phosphate synthase (SPS; EC 2.4.1.14), the major limiting enzyme for sucrose synthesis, is a candidate gene for QTLph1 among the annotation results of the region of QTLph1. The higher level of SPS transcripts and the activity of SPS in NIL6 compared to control plants, and the fact that the relative SPS activity per SPS protein content was almost the same between NIL6 and Nipponbare suggested that the higher plant height in NIL6 compared to Nipponbare was due to the high SPS activity in NIL6. In agreement with this hypothesis, transgenic rice plants with a maize SPS gene that had about 3 times the SPS activity of that in Nipponbare (control plants) were significantly taller than Nipponbare from the early growth stage. From these results and the physiological data from NIL6, we concluded that SPS is the targeted gene underlying QTLph1.




Similar content being viewed by others
Abbreviations
- cFBPase:
-
cytosolic fructose-1,6-bisphosphate
- LOD:
-
likelihood odds ratio
- NIL:
-
near isogenic line
- QTL:
-
quantitative trait locus
- QTLph1:
-
quantitative trait locus controlling plant height on chromosome 1
- RGA1:
-
heterotrimeric G protein
- SPS:
-
sucrose phosphate synthase
References
Altschul SF, Madden TL, Schäffer AA, Zhang J, Zhang Z, Miller W, Lipman DJ (1997) Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res 25: 3389–3402
Ashikari M, Wu J, Yano M, Sasaki T, Yoshimura A (1999) Rice gibberellin-insensitive dwarf mutant gene Dwarf 1 encodes the α-subunit of GTP-binding protein. Proc Natl Acad Sci USA 96:10284–10289
Bergmeyer HU, Bert E (1974) Methods for determination of metabolites: Carbohydrate metabolism: sucrose. In: Bergmeyer HU (ed) Methods of enzymatic analysis. Academic Press, New York, pp 1176–1179
de Vries P (1989) Assimilation and dissimilation of carbon. In: Baños L (ed) Simulation of ecophysiological processes of growth in several annual crops. IRRI, Manila, Philippines, pp 27–72
Federspiel N (2000) Deciphering a weed. Genomic sequencing of Arabidopsis. Plant Physiol 124:1456–1459
Foyer CH, Galtier N (1996) Source-sink interaction and communication in leaves. In: Zmaski E, Schafter AA (eds) Photoassimilate distribution in plants and crops: source–sink relations. Dekker, New York, pp 311–340
Frary A, Nesbitt TC, Frary A, Grandillo S, van der Knaap E, Cong B, Liu J, Meller J, Elber R, Alpert KB, Tanksley SD (2000) fw2.2: A quantitative trait locus key to the evolution of tomato fruit size. Science 289:85–88
Frewen BE, Chen THH, Howe GT, Davis J, Rohde A, Boerjan W, Bradshaw HD Jr (2000) Quantitative trait loci and candidate gene mapping of bud set and bud flush in Populus. Genetics 154:837–845
Fujisawa Y, Kato T, Ohki S, Ishikawa A, Kitano H, Sasaki T, Asahi T, Iwasaki Y (1999) Suppression of the heterotrimeric G protein causes abnormal morphology, including dwarfism, in rice. Proc Natl Acad Sci USA 96:7575–7580
Gale MD, Devos KM (1998) Comparative genetics in the grasses. Proc Natl Acad Sci USA 95:1971–1974
Galtier N, Foyer CH, Huber J, Voelker TA, Huber SC (1993) Effects of elevated sucrose-phosphate synthase activity on photosynthesis, assimilate partitioning and growth in tomato (Lycopersicon esculentum var UC82B). Plant Physiol 101:535–543
Goff SA, Ricke D, Lan TH et al. (2002) A draft sequence of the rice genome (Oryza sativa L. ssp. japonica). Science 296:92–100
Gifford RM, Evans LT (1981) Photosynthesis, carbon partitioning and yield. Annu Rev Plant Physiol 32:485–509
Hirochika H (1997) Retrotransposons of rice: their regulation and use for genome analysis. Plant Mol Biol 35:231–240
Huber SC, Huber JL (1996) Role and regulation of sucrose-phosphate synthase in higher plants. Annu Rev Plant Physiol Plant Mol Biol 47:431–444
Ishimaru K, Yano M, Aoki N, Ono K, Hirose T, Lin SY, Monna L, Sasaki T, Ohsugi R (2001a) Toward the mapping of physiological and agronomic characters on a rice function map: QTL analysis and comparison between QTLs and expressed sequence tags. Theor Appl Genet 102:792–800
Ishimaru K, Kobayashi N, Ono K, Yano M, Ohsugi R (2001b) Are contents of Rubisco, soluble protein and nitrogen in flag leaves of rice controlled by the same genetics? J Exp Bot 52:1827–1833
Ishimaru K, Shirota K, Higa M, Kawamitsu Y (2001c) Identification of quantitative trait loci for adaxial and abaxial stomatal frequencies in Oryza sativa. Plant Physiol Biochem 39:173–177
Kende H, van der Knaap E, Cho HT (1998) Deepwater rice: a model plant to study stem elongation. Plant Physiol 118:1105–1110
Lincoln S, Daly M, Lander E (1993) Mapping genes controlling quantitative traits with MAPMAKER/QTL 1.1: a tutorial and reference manual, 2nd edn. Whitehead Institute Technical Report, Cambridge, UK
Lunn JE, Hatch MD (1997) The role of sucrose-phosphate synthase in the control of photosynthate partitioning in Zea mays leaves. Aust J Plant Physiol 24:1–8
Mann CC (1999) Crop sciences seek a new revolution. Science 283:310–314
Monna L, Kitazawa N, Yoshino R, Suzuki J, Masuda H, Maehara Y, Tanji M, Sato M, Nasu S, Minobe Y (2002) Positional cloning of rice semidwarfing gene, sd-1: rice “green revolution gene” encodes mutant enzyme involved in gibberellin synthesis. DNA Res. 9:11–7
Niklas KJ, Enquist BJ (2000) Invariant scaling relationships for interspecific plant biomass production rates and body size. Proc Natl Acad Sci USA 98:2922–2927
Ono K, Ishimaru K, Aoki N, Takahashi S, Ozawa K, Ohkawa Y, Ohsugi R (1999) Characterization of a maize sucrose-phosphate synthase protein and its effect on carbon partitioning in transgenic rice plant. Plant Prod Sci 2:172–177
Prioul J, Pelleschi S, Sene M, Theevenot C, Causse M, de Vienne D, Leonardi A (1999) From QTLs for enzyme activity to candidate genes in maize. J Exp Bot 50: 1281–1288
Sari-Gorla M, Krajewski P, Di Fonzo N, Villa M, Frova C (1999) Genetic analysis of drought tolerance in maize by molecular markers. II. Plant height and flowering. Theor Appl Genet 99:289–295
Sasaki T, Matsumoto T, Yamamoto K et al. (2002) The genome sequence and structure of rice chromosome 1. Nature 420:312–316
Sato Y, Sentoku N, Miura Y, Hirochika H, Kitano H, Matsuoka M (1999) Loss-of-function mutations in the rice homeobox gene OSH15 affect the architecture of internodes resulting in dwarf plants. EMBO J 18:992–1002
Scott DB, Jin W, Ledford HK, Jung HS, Honma MA (1999) EAF1 regulates vegetative-phase change and flowering time in Arabidopsis. Plant Physiol 120:675–684
Signora L, Galtier N, Skot L, Lucas H, Foyer CH (1998) Over-expression of sucrose synthase in Arabidopsis thaliana results in increased foliar sucrose/starch ratios and favours decreased foliar carbohydrate accumulation in plants after prolonged growth with CO2 enrichment. J Exp Bot 49:669–680
Sussman MR, Amasino RM, Young JC, Krysan PJ, Austin-Phillips S (2000) The Arabidopsis knockout facility at the university of Wisconsin-Madison. Plant Physiol 124:1465–1467
Tanksley SD (1993) Mapping polygenes. Annu Rev Genet 27:205–233
Wadsworth GJ. Redinbaugh MG, Scandalios JG (1988) A procedure for the small-scale isolation of plant RNA suitable for RNA blot analysis. Anal Biochem 172:279–283
Wilson WA, Harrington SE, Woodman WL, Lee M, Sorrells ME, McCouch SR (1999) Inferences on the genome structure of progenitor maize through comparative analysis of rice, maize and the domesticated panicoids. Genetics 153:453–473
Xiao J, Li J, Yuan L, Tanksley SD (1996) Identification of QTLs affecting traits of agronomic importance in a recombinant inbred population derived from a subspecific rice cross. Theor Appl Genet 92:230–244
Xiao J, Li J, Grandillo S, Ahn SN, Yuan L, Tanksley SD, McCouch SR (1998) Identification of trait-improving quantitative trait loci alleles from a wild rice relative, Oryza rufipogon. Genetics 150:899–909
Yamamoto T, Lin H, Sasaki T, Yano M (2000) Identification of heading date quantitative trait locus Hd6 and characterization of its epistatic interactions with Hd2 in rice using advanced backcross progeny. Genetics 154:885–891
Yamanouchi U, Yano M, Lin H, Ashikari M, Yamada K (2002) A rice leaf gene, Spl7, encodes a heat stress transcription factor protein. Proc Natl Acad Sci USA 99:7530–7535
Yano M, Sasaki T (1997) Genetic and molecular dissection of quantitative traits in rice. Plant Mol Biol 35:145–153
Yano M, Katayose Y, Ashikari M, Yamanouchi U, Monna L, Fuse T, Baba T, Yamamoto K, Umehara Y, Nagamura Y, Sasaki T (2000) Hd1, a major photoperiod sensitivity quantitative trait locus in rice, is closely related to the Arabidopsis flowering time gene CONSTANS. Plant Cell 12:2473–2483
Yu J, Hu S, Wang J et al. (2002) A draft sequence of the rice genome (Oryza sativa L. ssp. indica). Science 296:79–92
Zhuang JY, Lin HX, Lu J, Qian HR, Hittalmani S, Huang N, Zheng KL (1997) Analysis of QTL × environment interaction for yield components and plant height in rice. Theor Appl Genet 95:799–808
Acknowledgements
The authors thank Dr. Masahiro Yano for providing NIL seeds, and Prof. Mirella Sari-Gorla and Drs. Enrico Pe, Luca Gianfranceschi, Lisa Monna, Kazuhiko Kobayashi and Haruto Sasaki for their kind suggestions.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Ishimaru, K., Ono, K. & Kashiwagi, T. Identification of a new gene controlling plant height in rice using the candidate-gene strategy. Planta 218, 388–395 (2004). https://doi.org/10.1007/s00425-003-1119-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00425-003-1119-z