Abstract
Subjects
Little is known about the pathology and pathogenesis of the Moyamoya disease (MMD). To better understand the molecular processes involved in MMD gene expression analysis of Moyamoya artery tissue to a control artery, this study was conducted.
Methods
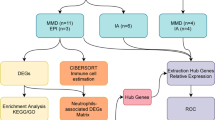
Tissue samples of two MMD and two non-MMD were profiled using oligonucleotide microarrays. Gene ontology classification of the differentially expressed genes was analyzed, and regulatory functional networks and pathways were identified with a network-based computational pathway analysis tool. Analysis of MMD and control tissue revealed 104 differentially expressed genes.
Results
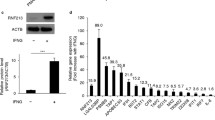
The two major significantly associated gene ontology terms was cellular development and cellular movement. Further network-based analysis showed significant interaction between RNF213 downstream gene networks and the top 3 score gene networks of MMD. Three major nodes of this network were evident in the merged network and were showing interactions with downstream network of RNF213.
Conclusions
Our results demonstrate that cellular development and cellular movement in MMD are the key role of mechanisms.



Similar content being viewed by others
References
Suzuki J, Kodama N (1983) Moyamoya disease—a review. Stroke 14:104–109
Suzuki J, Takaku A (1969) Cerebrovascular “Moyamoya” disease. Disease showing abnormal net-like vessels in base of brain. Arch Neurol 20:288–299
Kuriyama S, Kusaka Y, Fujimura M, Wakai K, Tamakoshi A, Hashimoto S, Tsuji I, Inaba Y, Yoshimoto T (2008) Prevalence and clinicoepidemiological features of Moyamoya disease in Japan: findings from a nationwide epidemiological survey. Stroke 39:42–47
Baba T, Houkin K, Kuroda S (2008) Novel epidemiological features of Moyamoya disease. J Neurol Neurosurg Psychiatry 79:900–904
Ikeda H, Sasaki T, Yoshimoto T, Fukui M, Arinami T (1999) Mapping of a familial Moyamoya disease gene to chromosome 3p24.2–p26. Am J Hum Genet 64:533–537
Nanba R, Tada M, Kuroda S, Houkin K, Iwasaki Y (2005) Sequence analysis and bioinformatics analysis of chromosome 17q25 in familial Moyamoya disease. Childs Nerv Syst 21:62–68
Inoue TK, Ikezaki K, Sasazuki T, Matsushima T, Fukui M (2000) Linkage analysis of Moyamoya disease on chromosome 6. J Child Neurol 15:179–182
Sakurai K, Horiuchi Y, Ikeda H, Ikezaki K, Yoshimoto T, Fukui M, Arinami T (2004) A novel susceptibility locus for Moyamoya disease on chromosome 8q23. J Hum Genet 49:278–281
Kamada F, Aoki Y, Narisawa A, Abe Y, Komatsuzaki S, Kikuchi A, Kanno J, Niihori T, Ono M, Ishii N, Owada Y, Fujimura M, Mashimo Y, Suzuki Y, Hata A, Tsuchiya S, Tominaga T, Matsubara Y, Kure S (2011) A genome-wide association study identifies RNF213 as the first Moyamoya disease gene. J Hum Genet 56:34–40
Smith JL (2009) Understanding and treating Moyamoya disease in children. Neurosurg Focus 26:E4
Soriano SG, Cowan DB, Proctor MR, Scott RM (2002) Levels of soluble adhesion molecules are elevated in the cerebrospinal fluid of children with Moyamoya syndrome. Neurosurgery 50:544–549
Takahashi A, Sawamura Y, Houkin K, Kamiyama H, Abe H (1993) The cerebrospinal fluid in patients with Moyamoya disease (spontaneous occlusion of the circle of Willis) contains high level of basic fibroblast growth factor. Neurosci Lett 160:214–216
Yamamoto M, Aoyagi M, Tajima S, Wachi H, Fukai N, Matsushima Y, Yamamoto K (1997) Increase in elastin gene expression and protein synthesis in arterial smooth muscle cells derived from patients with Moyamoya disease. Stroke 28:1733–1738
Yoshimoto T, Houkin K, Takahashi A, Abe H (1996) Angiogenic factors in Moyamoya disease. Stroke 27:2160–2165
Malek AM, Connors S, Robertson RL, Folkman J, Scott RM (1997) Elevation of cerebrospinal fluid levels of basic fibroblast growth factor in moyamoya and central nervous system disorders. Pediatr Neurosurg 27:182–189
Yamamoto M, Aoyagi M, Fukai N, Matsushima Y, Yamamoto K (1998) Differences in cellular responses to mitogens in arterial smooth muscle cells derived from patients with Moyamoya disease. Stroke 29:1188–1193
Masuda J, Ogata J, Yutani C (1993) Smooth muscle cell proliferation and localization of macrophages and T cells in the occlusive intracranial major arteries in Moyamoya disease. Stroke 24:1960–1967
Acknowledgments
The authors would like to thank David Huang and staff of the Department of Neurosurgery, Tokyo Women’s Medical University for their guidance in preparing this paper.
Declaration of funding source
This study was supported by research funds of the Department of Neurosurgery, Tokyo Women’s Medical University.
Conflict of interest
The authors have no personal financial or institutional interest in any of the materials or devices described in this article.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Okami, N., Aihara, Y., Akagawa, H. et al. Network-based gene expression analysis of vascular wall of juvenile Moyamoya disease. Childs Nerv Syst 31, 399–404 (2015). https://doi.org/10.1007/s00381-014-2605-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00381-014-2605-2