Abstract
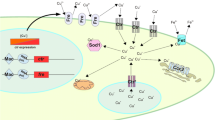
Copper (Cu) is an essential metal for all living organisms, although it is toxic in excess. Filamentous fungus must acquire copper from its environment for growth. Despite its essentiality for growth, the mechanisms that maintain copper homeostasis are not fully understood in filamentous fungus. To gain insights into copper homeostasis, we investigated the roles of a copper transcription factor Afmac1 in the life-threatening fungus Aspergillus fumigatus, a homolog of the yeast MAC1. We observed that the Afmac1 deletion mutant exhibited not only significantly slower growth, but also incomplete conidiation including a short chain of conidia and defective melanin. Moreover, the expressions of the copper transporters, ctrA1, ctrA2, and ctrC, and metalloreductases, Afu8g01310 and fre7, were repressed in ∆Afmac1 cells, while those expressions were induced under copper depletion conditions in wild-type. The expressions of pksP and wetA, which are, respectively, involved in biosynthesis of conidia-specific melanin and the late stage of conidiogenesis, were decreased in the ∆Afmac1 strain under minimal media condition. Taken together, these results indicate that copper acquisition through AfMac1 functions in growth as well as conidiation.






Similar content being viewed by others
References
Bailey TL, Elkan C (1994) Fitting a mixture model by expectation maximization to discover motifs in biopolymers. Proc Int Conf Intell Syst Mol Biol 2:28–36
Bodey GP, Vartivarian S (1989) Aspergillosis. Eur J Clin Microbiol Infect Dis 8:413–437
Carroll A, Sweigard J, Valent B (1994) Improved vectors for selecting resistance to hygromycin. Fungal Genet Newsl 41:22
Catlett N, Lee B-N, Yoder O, Turgeon BG (2003) Split-marker recombination for efficient targeted deletion of fungal genes. Fungal Genet Newsl 50:9–11
Ding C, Yin J, Tovar EM, Fitzpatrick DA, Higgins DG, Thiele DJ (2011) The copper regulon of the human fungal pathogen Cryptococcus neoformans H99. Mol Microbiol 81:1560–1576. doi:10.1111/j.1365-2958.2011.07794.x
Ding C, Festa RA, Chen YL, Espart A, Palacios O, Espin J, Capdevila M, Atrian S, Heitman J, Thiele DJ (2013) Cryptococcus neoformans copper detoxification machinery is critical for fungal virulence. Cell Host Microbe 13:265–276. doi:10.1016/j.chom.2013.02.002
Ding C, Festa RA, Sun TS, Wang ZY (2014) Iron and copper as virulence modulators in human fungal pathogens. Mol Microbiol 93:10–23. doi:10.1111/mmi.12653
Festa RA, Thiele DJ (2011) Copper: an essential metal in biology. Curr Biol 21:R877–R883. doi:10.1016/j.cub.2011.09.040
Georgatsou E, Alexandraki D (1999) Regulated expression of the Saccharomyces cerevisiae Fre1p/Fre2p Fe/Cu reductase related genes. Yeast 15:573–584. doi:10.1002/(SICI)1097-0061(199905)15:7<573::AID-YEA404>3.0.CO;2-7
Graden JA, Winge DR (1997) Copper-mediated repression of the activation domain in the yeast Mac1p transcription factor. Proc Natl Acad Sci U S A 94:5550–5555
Griffith GW, Easton GL, Detheridge A, Roderick K, Edwards A, Worgan HJ, Nicholson J, Perkins WT (2007) Copper deficiency in potato dextrose agar causes reduced pigmentation in cultures of various fungi. FEMS Microbiol Lett 276:165–171. doi:10.1111/j.1574-6968.2007.00923.x
Gross C, Kelleher M, Iyer VR, Brown PO, Winge DR (2000) Identification of the copper regulon in Saccharomyces cerevisiae by DNA microarrays. J Biol Chem 275:32310–32316. doi:10.1074/jbc.M005946200
Hagiwara D, Takahashi-Nakaguchi A, Toyotome T, Yoshimi A, Abe K, Kamei K, Gonoi T, Kawamoto S (2013) NikA/TcsC histidine kinase is involved in conidiation, hyphal morphology, and responses to osmotic stress and antifungal chemicals in Aspergillus fumigatus. PLoS One 8:e80881. doi:10.1371/journal.pone.0080881
Jamison McDaniels CP, Jensen LT, Srinivasan C, Winge DR, Tullius TD (1999) The yeast transcription factor Mac1 binds to DNA in a modular fashion. J Biol Chem 274:26962–26967
Jiang N, Sun N, Xiao D, Pan J, Wang Y, Zhu X (2009) A copper-responsive factor gene CUF1 is required for copper induction of laccase in Cryptococcus neoformans. FEMS Microbiol Lett 296:84–90. doi:10.1111/j.1574-6968.2009.01619.x
Jiang N, Liu X, Yang J, Li Z, Pan J, Zhu X (2011) Regulation of copper homeostasis by Cuf1 associates with its subcellular localization in the pathogenic yeast Cryptococcus neoformans H99. FEMS Yeast Res 11:440–448. doi:10.1111/j.1567-1364.2011.00733.x
Jungmann J, Reins HA, Lee J, Romeo A, Hassett R, Kosman D, Jentsch S (1993) MAC1, a nuclear regulatory protein related to Cu-dependent transcription factors is involved in Cu/Fe utilization and stress resistance in yeast. EMBO J 12:5051–5056
Keller G, Bird A, Winge DR (2005) Independent metalloregulation of Ace1 and Mac1 in Saccharomyces cerevisiae. Eukaryot Cell 4:1863–1871. doi:10.1128/EC.4.11.1863-1871.2005
Kim BE, Nevitt T, Thiele DJ (2008) Mechanisms for copper acquisition, distribution and regulation. Nat Chem Biol 4:176–185. doi:10.1038/nchembio.72
Langfelder K, Jahn B, Gehringer H, Schmidt A, Wanner G, Brakhage AA (1998) Identification of a polyketide synthase gene (pksP) of Aspergillus fumigatus involved in conidial pigment biosynthesis and virulence. Med Microbiol Immunol 187:79–89
Langfelder K, Streibel M, Jahn B, Haase G, Brakhage AA (2003) Biosynthesis of fungal melanins and their importance for human pathogenic fungi. Fungal Genet Biol 38:143–158
Mah JH, Yu JH (2006) Upstream and downstream regulation of asexual development in Aspergillus fumigatus. Eukaryot Cell 5:1585–1595. doi:10.1128/EC.00192-06
Martins LJ, Jensen LT, Simon JR, Keller GL, Winge DR (1998) Metalloregulation of FRE1 and FRE2 homologs in Saccharomyces cerevisiae. J Biol Chem 273:23716–23721
Ni M, Yu JH (2007) A novel regulator couples sporogenesis and trehalose biogenesis in Aspergillus nidulans. PLoS One 2:e970. doi:10.1371/journal.pone.0000970
Ory JJ, Griffith CL, Doering TL (2004) An efficiently regulated promoter system for Cryptococcus neoformans utilizing the CTR4 promoter. Yeast 21:919–926. doi:10.1002/yea.1139
Park YS, Lian H, Chang M, Kang CM, Yun CW (2014) Identification of high-affinity copper transporters in Aspergillus fumigatus. Fungal Genet Biol 73:29–38. doi:10.1016/j.fgb.2014.09.008
Pena MM, Koch KA, Thiele DJ (1998) Dynamic regulation of copper uptake and detoxification genes in Saccharomyces cerevisiae. Mol Cell Biol 18:2514–2523
Pena MM, Puig S, Thiele DJ (2000) Characterization of the Saccharomyces cerevisiae high affinity copper transporter Ctr3. J Biol Chem 275:33244–33251. doi:10.1074/jbc.M005392200
Pihet M, Vandeputte P, Tronchin G, Renier G, Saulnier P, Georgeault S, Mallet R, Chabasse D, Symoens F, Bouchara JP (2009) Melanin is an essential component for the integrity of the cell wall of Aspergillus fumigatus conidia. BMC Microbiol 9:177. doi:10.1186/1471-2180-9-177
Pilsyk S, Natorff R, Sienko M, Skoneczny M, Paszewski A, Brzywczy J (2015) The Aspergillus nidulans metZ gene encodes a transcription factor involved in regulation of sulfur metabolism in this fungus and other Eurotiales. Curr Genet 61:115–125. doi:10.1007/s00294-014-0459-5
Puig S, Thiele DJ (2002) Molecular mechanisms of copper uptake and distribution. Curr Opin Chem Biol 6:171–180
Rees EM, Thiele DJ (2004) From aging to virulence: forging connections through the study of copper homeostasis in eukaryotic microorganisms. Curr Opin Microbiol 7:175–184. doi:10.1016/j.mib.2004.02.004
Robinson NJ, Winge DR (2010) Copper metallochaperones. Annu Rev Biochem 79:537–562. doi:10.1146/annurev-biochem-030409-143539
Sakai K, Kinoshita H, Nihira T (2012) Heterologous expression system in Aspergillus oryzae for fungal biosynthetic gene clusters of secondary metabolites. Appl Microbiol Biotechnol 93:2011–2022. doi:10.1007/s00253-011-3657-9
Samanovic MI, Ding C, Thiele DJ, Darwin KH (2012) Copper in microbial pathogenesis: meddling with the metal. Cell Host Microbe 11:106–115. doi:10.1016/j.chom.2012.01.009
Sugareva V, Hartl A, Brock M, Hubner K, Rohde M, Heinekamp T, Brakhage AA (2006) Characterisation of the laccase-encoding gene abr2 of the dihydroxynaphthalene-like melanin gene cluster of Aspergillus fumigatus. Arch Microbiol 186:345–355. doi:10.1007/s00203-006-0144-2
Tao L, Yu JH (2011) AbaA and WetA govern distinct stages of Aspergillus fumigatus development. Microbiology 157:313–326. doi:10.1099/mic.0.044271-0
Tsai HF, Chang YC, Washburn RG, Wheeler MH, Kwon-Chung KJ (1998) The developmentally regulated alb1 gene of Aspergillus fumigatus: its role in modulation of conidial morphology and virulence. J Bacteriol 180:3031–3038
Upadhyay S, Torres G, Lin X (2013) Laccases involved in 1,8-dihydroxynaphthalene melanin biosynthesis in Aspergillus fumigatus are regulated by developmental factors and copper homeostasis. Eukaryot Cell 12:1641–1652. doi:10.1128/EC.00217-13
Valentine JS, Gralla EB (1997) Delivering copper inside yeast and human cells. Science 278:817–818
Waterman SR, Hacham M, Hu G, Zhu X, Park Y-D, Shin S, Panepinto J, Valyi-Nagy T, Beam C, Husain S (2007) Role of a CUF1/CTR4 copper regulatory axis in the virulence of Cryptococcus neoformans. J Clin Invest 117:794–802. doi:10.1172/JCI30006
Waterman SR, Park YD, Raja M, Qiu J, Hammoud DA, O’Halloran TV, Williamson PR (2012) Role of CTR4 in the virulence of Cryptococcus neoformans. MBio 3:pii: e00285-12. doi:10.1128/mBio.00285-12
Yamaguchi-Iwai Y, Serpe M, Haile D, Yang W, Kosman DJ, Klausner RD, Dancis A (1997) Homeostatic regulation of copper uptake in yeast via direct binding of MAC1 protein to upstream regulatory sequences of FRE1 and CTR1. J Biol Chem 272:17711–17718
Yu JH (2010) Regulation of development in Aspergillus nidulans and Aspergillus fumigatus. Mycobiology 38:229–237 doi:10.4489/NYCO.2010.38.4.229
Yu JH, Mah JH, Seo JA (2006) Growth and developmental control in the model and pathogenic aspergilli. Eukaryot Cell 5:1577–1584. doi:10.1128/EC.00193-06
Yun CW, Bauler M, Moore RE, Klebba PE, Philpott CC (2001) The role of the FRE family of plasma membrane reductases in the uptake of siderophore-iron in Saccharomyces cerevisiae. J Biol Chem 276:10218–10223. doi:10.1074/jbc.M010065200
Zhu Z, Labbe S, Pena MM, Thiele DJ (1998) Copper differentially regulates the activity and degradation of yeast Mac1 transcription factor. J Biol Chem 273:1277–1280
Acknowledgements
We would like to thank Taku Oshima (Nara Institute of Science and Technology) and Takashi Umeyama (National Institute of Infectious Diseases) for their fruitful discussions, and Ryoko Mori for experimental assistance. This work has been partly supported by the Takeda Science Foundation to H. T., MEXT KAKENHI (Nos. 16K18671, 16H06279 to H. T., and 25290079 to T. Y.), the Institute for Global Prominent Research, Chiba University to H.T., and the Tenure Tracking System Program of MEXT to Y.K. and H.T. We thank the National BioResource Project—Pathogenic Microbes in Japan (http://www.nbrp.jp/).
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by M. Kupiec.
Electronic supplementary material
Below is the link to the electronic supplementary material.
294_2017_681_MOESM1_ESM.pdf
Split-marker strategy for the ∆Afmac1 strain. (a) The preparation of two fusion PCR products. Fragments A and B were amplified from A. fumigatus chromosomes. The figure is a schematic representation of the location of primers used for recombinant PCR. The ptrA gene was amplified from the pPTRI plasmid. (b) Homologous recombination. The two fusion PCR products were used directly for transformation. Homologous recombination between the overlapping regions of ptrA, and between the flank regions and chromosomal DNA results in Afmac1 deletion (PDF 155 KB)
294_2017_681_MOESM2_ESM.pdf
Confirmation of the constructed mutants, ∆Afmac1 and Co-Afmac1. (a) Genomic architectures of Afmac1::ptrA in ∆Afmac1 and Afmac1-HygB rin Co-Afmac1. (b) Agarose gel electrophoresis of PCR products. Five primer sets were designed for confirmation of ∆Afmac1 and Co-Afmac1. (c) Quantitative real-time RT-PCR analyses of Afmac1 in WT, ∆Afmac1, and Co-Afmac1 strains. Average values and error bars were obtained from three experiments (PDF 240 KB)
294_2017_681_MOESM4_ESM.pdf
Phialides and conidia of WT and ∆Afmac1 strain. The structure of conidiogenesis was observed by performing a slide culture on the AMM medium (PDF 569 KB)
294_2017_681_MOESM5_ESM.pdf
Quantitative real-time RT-PCR analyses of wetA and pksP at a high copper concentration (AMM containing 100 µM CuSO4). Average values and error bars were obtained from three independent experiments (PDF 138 KB)
Rights and permissions
About this article
Cite this article
Kusuya, Y., Hagiwara, D., Sakai, K. et al. Transcription factor Afmac1 controls copper import machinery in Aspergillus fumigatus . Curr Genet 63, 777–789 (2017). https://doi.org/10.1007/s00294-017-0681-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00294-017-0681-z