Abstract
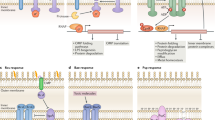
The cell envelope stress response (CESR) encompasses all regulatory events that enable a cell to protect the integrity of its envelope, an essential structure of any bacterial cell. The underlying signaling network is particularly well understood in the Gram-positive model organism Bacillus subtilis. It consists of a number of two-component systems (2CS) and extracytoplasmic function σ factors that together regulate the production of both specific resistance determinants and general mechanisms to protect the envelope against antimicrobial peptides targeting the biogenesis of the cell wall. Here, we summarize the current picture of the B. subtilis CESR network, from the initial identification of the corresponding signaling devices to unraveling their interdependence and the underlying regulatory hierarchy within the network. In the course of detailed mechanistic studies, a number of novel signaling features could be described for the 2CSs involved in mediating CESR. This includes a novel class of so-called intramembrane-sensing histidine kinases (IM-HKs), which—instead of acting as stress sensors themselves—are activated via interprotein signal transfer. Some of these IM-HKs are involved in sensing the flux of antibiotic resistance transporters, a unique mechanism of responding to extracellular antibiotic challenge.
Similar content being viewed by others
References
Bernard R, Guiseppi A, Chippaux M, Foglino M, Denizot F (2007) Resistance to bacitracin in Bacillus subtilis: unexpected requirement of the BceAB ABC transporter in the control of expression of its own structural genes. J Bacteriol 189:8636–8642. doi:10.1128/jb.01132-07
Bugg TD, Braddick D, Dowson CG, Roper DI (2011) Bacterial cell wall assembly: still an attractive antibacterial target. Trends Biotechnol 29:167–173. doi:10.1016/j.tibtech.2010.12.006
Butcher BG, Lin Y-P, Helmann JD (2007) The yydFGHIJ operon of Bacillus subtilis encodes a peptide that induces the LiaRS two-component system. J Bacteriol 189:8616–8625. doi:10.1128/JB.01181-07
Cao M, Helmann JD (2002) Regulation of the Bacillus subtilis bcrC bacitracin resistance gene by two extracytoplasmic function sigma factors. J Bacteriol 184:6123–6129. doi:10.1128/JB.184.22.6123-6129.2002
Cao M, Kobel PA, Morshedi MM, Wu MF, Paddon C, Helmann JD (2002) Defining the Bacillus subtilis σW regulon: a comparative analysis of promoter consensus search, run-off transcription/macroarray analysis (ROMA), and transcriptional profiling approaches. J Mol Biol 316:443–457. doi:10.1006/jmbi.2001.5372
Dintner S, Staron A, Berchtold E, Petri T, Mascher T, Gebhard S (2011) Co-evolution of ABC-transporters and two-component regulatory systems as resistance modules against antimicrobial peptides in Firmicutes bacteria. J Bacteriol 193:3851–3862. doi:10.1128/JB.05175-11
Dintner S, Heermann R, Fang C, Jung K, Gebhard S (2014) A sensory complex consisting of an ATP-binding cassette transporter and a two-component regulatory system controls bacitracin resistance in Bacillus subtilis. J Biol Chem 289:27899–27910. doi:10.1074/jbc.M114.596221
Dufresne K, Paradis-Bleau C (2015) Biology and assembly of the bacterial envelope. In: Krogan PJN, Babu PM (eds) Prokaryotic systems biology. Springer, Cham, pp 41–76
Economou NJ, Cocklin S, Loll PJ (2013) High-resolution crystal structure reveals molecular details of target recognition by bacitracin. Proc Natl Acad Sci USA 110:14207–14212. doi:10.1073/pnas.1308268110
Eiamphungporn W, Helmann JD (2008) The Bacillus subtilis σM regulon and its contribution to cell envelope stress responses. Mol Microbiol 67:830–848. doi:10.1111/j.1365-2958.2007.06090.x
Fritz G, Dintner S, Treichel NS, Radeck J, Gerland U, Mascher T, Gebhard S (2015) A new way of sensing: need-based activation of antibiotic resistance by a flux-sensing mechanism. MBio 6:e00975. doi:10.1128/mBio.00975-15
Gauntlett JC, Gebhard S, Keis S, Manson JM, Pos KM, Cook GM (2008) Molecular analysis of BcrR, a membrane-bound bacitracin sensor and DNA-binding protein from Enterococcus faecalis. J Biol Chem 283:8591–8600. doi:10.1074/jbc.M709503200
Gebhard S, Gaballa A, Helmann JD, Cook GM (2009) Direct stimulus perception and transcription activation by a membrane-bound DNA binding protein. Mol Microbiol 73:482–491. doi:10.1111/j.1365-2958.2009.06787.x
Gonzalez-Pastor JE (2011) Cannibalism: a social behavior in sporulating Bacillus subtilis. FEMS Microbiol Rev 35:415–424. doi:10.1111/j.1574-6976.2010.00253.x
Helmann JD (2016) Bacillus subtilis extracytoplasmic function (ECF) sigma factors and defense of the cell envelope. Curr Opin Microbiol 30:122–132. doi:10.1016/j.mib.2016.02.002
Hiron A, Falord M, Valle J, Debarbouille M, Msadek T (2011) Bacitracin and nisin resistance in Staphylococcus aureus: a novel pathway involving the BraS/BraR two-component system (SA2417/SA2418) and both the BraD/BraE and VraD/VraE ABC transporters. Mol Microbiol 81:602–622. doi:10.1111/j.1365-2958.2011.07735.x
Höfler C, Heckmann J, Fritsch A, Popp P, Gebhard S, Fritz G, Mascher T (2016) Cannibalism stress response in Bacillus subtilis. Microbiology 162:164–176. doi:10.1099/mic.0.000176
Horsburgh MJ, Moir A (1999) σM, an ECF RNA polymerase sigma factor of Bacillus subtilis 168, is essential for growth and survival in high concentrations of salt. Mol Microbiol 32:41–50. doi:10.1046/j.1365-2958.1999.01323.x
Inoue H, Suzuki D, Asai K (2013) A putative bactoprenol glycosyltransferase, CsbB, in Bacillus subtilis activates SigM in the absence of co-transcribed YfhO. Biochem Biophys Res Commun 436:6–11. doi:10.1016/j.bbrc.2013.04.064
Jeong DW, Cho H, Jones MB, Shatzkes K, Sun F, Ji Q, Liu Q, Peterson SN, He C, Bae T (2012) The auxiliary protein complex SaePQ activates the phosphatase activity of sensor kinase SaeS in the SaeRS two-component system of Staphylococcus aureus. Mol Microbiol 86:331–348. doi:10.1111/j.1365-2958.2012.08198.x
Jordan S, Junker A, Helmann JD, Mascher T (2006) Regulation of LiaRS-dependent gene expression in Bacillus subtilis: identification of inhibitor proteins, regulator binding sites and target genes of a conserved cell envelope stress-sensing two-component system. J Bacteriol 188:5153–5166. doi:10.1128/JB.00310-06
Jordan S, Hutchings MI, Mascher T (2008) Cell envelope stress response in Gram-positive bacteria. FEMS Microbiol Rev 32:107–146. doi:10.1111/j.1574-6976.2007.00091.x
Joseph P, Fichant G, Quentin Y, Denizot F (2002) Regulatory relationship of two-component and ABC transport systems and clustering of their genes in the Bacillus/Clostridium group, suggest a functional link between them. J Mol Microbiol Biotechnol 4:503–513
Kallenberg F, Dintner S, Schmitz R, Gebhard S (2013) Identification of regions important for resistance and signalling within the antimicrobial peptide transporter BceAB of Bacillus subtilis. J Bacteriol 195:3287–3297. doi:10.1128/JB.00419-13
Kingston AW, Liao X, Helmann JD (2013) Contributions of the σW, σM and σX regulons to the lantibiotic resistome of Bacillus subtilis. Mol Microbiol 90:502–518. doi:10.1111/mmi.12380
Kingston AW, Zhao H, Cook GM, Helmann JD (2014) Accumulation of heptaprenyl diphosphate sensitizes Bacillus subtilis to bacitracin: implications for the mechanism of resistance mediated by the BceAB transporter. Mol Microbiol 93:37–49. doi:10.1111/mmi.12637
Kovács ÁT (2016) Bacterial differentiation via gradual activation of global regulators. Curr Genet 62:125–128. doi:10.1007/s00294-015-0524-8
Lee YH, Helmann JD (2013) Reducing the level of undecaprenyl pyrophosphate synthase has complex effects on susceptibility to cell wall antibiotics. Antimicrob Agents Chemother 57:4267–4275. doi:10.1128/aac.00794-13
Luttmann D, Gopel Y, Gorke B (2012) The phosphotransferase protein EIIA(Ntr) modulates the phosphate starvation response through interaction with histidine kinase PhoR in Escherichia coli. Mol Microbiol 86:96–110. doi:10.1111/j.1365-2958.2012.08176.x
Mascher T (2006) Intramembrane-sensing histidine kinases: a new family of cell envelope stress sensors in Firmicutes bacteria. FEMS Microbiol Lett 264:133–144. doi:10.1111/j.1574-6968.2006.00444.x
Mascher T (2014) Bacterial (intramembrane-sensing) histidine kinases: signal transfer rather than stimulus perception. Trends Microbiol 22:559–565. doi:10.1016/j.tim.2014.05.006
Mascher T, Margulis NG, Wang T, Ye RW, Helmann JD (2003) Cell wall stress responses in Bacillus subtilis: the regulatory network of the bacitracin stimulon. Mol Microbiol 50:1591–1604. doi:10.1046/j.1365-2958.2003.03786.x
Mascher T, Zimmer SL, Smith TA, Helmann JD (2004) Antibiotic-inducible promoter regulated by the cell envelope stress-sensing two-component system LiaRS of Bacillus subtilis. Antimicrob Agents Chemother 48:2888–2896. doi:10.1128/AAC.48.8.2888-2896.2004
Mascher T, Helmann JD, Unden G (2006) Stimulus perception in bacterial signal-transducing histidine kinases. Microbiol Mol Biol Rev 90:910–938. doi:10.1128/MMBR.00020-06
Mascher T, Hachmann AB, Helmann JD (2007) Regulatory overlap and functional redundancy among Bacillus subtilis extracytoplasmic function σ factors. J Bacteriol 189:6919–6927. doi:10.1128/JB.00904-07
Meeske AJ, Sham LT, Kimsey H, Koo BM, Gross CA, Bernhardt TG, Rudner DZ (2015) MurJ and a novel lipid II flippase are required for cell wall biogenesis in Bacillus subtilis. Proc Natl Acad Sci USA 112:6437–6442. doi:10.1073/pnas.1504967112
Ming L-J, Epperson JD (2002) Metal binding and structure-activity relationship of the metalloantibiotic peptide bacitracin. J Inorg Biochem 91:46–58. doi:10.1016/S0162-0134(02)00464-6
Ohki R, Giyanto Tateno K, Masuyama W, Moriya S, Kobayashi K, Ogasawara N (2003) The BceRS two-component regulatory system induces expression of the bacitracin transporter, BceAB, in Bacillus subtilis. Mol Microbiol 49:1135–1144. doi:10.1046/j.1365-2958.2003.03653.x
Petersohn A, Brigulla M, Haas S, Hoheisel JD, Völker U, Hecker M (2001) Global analysis of the general stress response of Bacillus subtilis. J Bacteriol 183:5617–5631. doi:10.1128/JB.183.19.5617-5631.2001
Pietiäinen M, Gardemeister M, Mecklin M, Leskela S, Sarvas M, Kontinen VP (2005) Cationic antimicrobial peptides elicit a complex stress response in Bacillus subtilis that involves ECF-type sigma factors and two-component signal transduction systems. Microbiology 151:1577–1592. doi:10.1099/mic.0.27761-0
Pollak S, Omer Bendori S, Eldar A (2015) A complex path for domestication of B. subtilis sociality. Curr Genet 61:493–496. doi:10.1007/s00294-015-0479-9
Radeck J, Gebhard S, Orchard PS, Kirchner M, Bauer S, Mascher T, Fritz G (2016) Anatomy of the bacitracin resistance network in Bacillus subtilis. Mol Microbiol 100:607–620. doi:10.1111/mmi.13336
Rietkötter E, Hoyer D, Mascher T (2008) Bacitracin sensing in Bacillus subtilis. Mol Microbiol 68:768–785. doi:10.1111/j.1365-2958.2008.06194.x
Schneider T, Sahl H-G (2010) An oldie but a goodie—cell wall biosynthesis as antibiotic target pathway. Int J Med Microbiol 300:161–169. doi:10.1016/j.ijmm.2009.10.005
Schrecke K, Staroń A, Mascher T (2012) Two-component signaling in the Gram-positive envelope stress response: intramembrane-sensing histidine kinases and accessory membrane proteins. In: Gross R, Beier D (eds) Two component systems in bacteria. Horizon Scientific Press, Hethersett, pp 199–229
Schrecke K, Jordan S, Mascher T (2013) Stoichiometry and perturbation studies of the LiaFSR system of Bacillus subtilis. Mol Microbiol 87:769–788. doi:10.1111/mmi.12130
Silhavy TJ, Kahne D, Walker S (2010) The bacterial cell envelope. Cold Spring Harb Perspect Biol 2:a000414. doi:10.1101/cshperspect.a000414
Staroń A, Finkeisen DE, Mascher T (2011) Peptide antibiotic sensing and detoxification modules of Bacillus subtilis. Antimicrob Agents Chemother 55:515–525. doi:10.1128/AAC.00352-10
Storm DR, Strominger JL (1974) Binding of bacitracin to cells and protoplasts of Micrococcus lysodeikticus. J Biol Chem 249:1823–1827
Wecke T, Mascher T (2011) Antibiotic research in the age of omics: from expression profiles to interspecies communication. Antimicrob Chemother 66:2689–2704. doi:10.1093/jac/dkr373
Wecke T, Zühlke D, Mäder U, Jordan S, Voigt B, Pelzer S, Labischinski H, Homuth G, Hecker M, Mascher T (2009) Daptomycin versus friulimicin B: in-depth profiling of Bacillus subtilis cell envelope stress responses. J Antimicrob Chemother 53:1619–1623. doi:10.1128/aac.01046-08
Wecke T, Bauer T, Harth H, Mäder U, Mascher T (2011) The rhamnolipid stress response of Bacillus subtilis. FEMS Microbiol Lett. doi:10.1111/j.1574-6968.2011.02367.x
Wiegert T, Homuth G, Versteeg S, Schumann W (2001) Alkaline shock induces the Bacillus subtilis σW regulon. Mol Microbiol 41:59–71. doi:10.1046/j.1365-2958.2001.02489.x
Wolf D, Kalamorz F, Wecke T, Juszczak A, Mäder U, Homuth G, Jordan S, Kirstein J, Hoppert M, Voigt B, Hecker M, Mascher T (2010) In-depth profiling of the LiaR response of Bacillus subtilis. J Bacteriol 192:4680–4693. doi:10.1128/JB.00543-10
Wolf D, Dominguez-Cuevas P, Daniel RA, Mascher T (2012) Cell envelope stress response in cell wall-deficient L-forms of Bacillus subtilis. Antimicrob Agents Chemother 56:5907–5915. doi:10.1128/AAC.00770-12
Yoshimura M, Asai K, Sadaie Y, Yoshikawa H (2004) Interaction of Bacillus subtilis extracytoplasmic function (ECF) sigma factors with the N-terminal regions of their potential anti-sigma factors. Microbiology 150:591–599. doi:10.1099/mic.0.26712-0
Acknowledgments
The authors would like to acknowledge the contributions of numerous co-workers of the Mascher group, who by their dedication, hard work and intellectual input shaped our picture of the cell envelope stress response of B. subtilis in the past decade.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Funding
Work on the cell envelope stress response of B. subtilis in the Mascher and Fritz groups was continuously supported by Grants from the Deutsche Forschungsgemeinschaft (DFG) (Grants MA 3269, MA2837/1-3, and MA2837/3-1 to TM as well as Grant FR 3673/1-2 to GF).
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Communicated by M. Kupiec.
Rights and permissions
About this article
Cite this article
Radeck, J., Fritz, G. & Mascher, T. The cell envelope stress response of Bacillus subtilis: from static signaling devices to dynamic regulatory network. Curr Genet 63, 79–90 (2017). https://doi.org/10.1007/s00294-016-0624-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00294-016-0624-0