Abstract
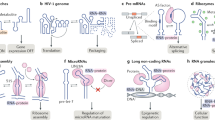
Recently a number of seminal studies have revealed that both sequence and spatio-temporal factors govern RNA decay in bacteria, which is crucial for regulation of gene expression. Ribonucleases have been described that not only exhibit sequence preferences, but also are sub-cellularly localised. Furthermore, the RNA itself is distributed in an organised manner and does not diffuse freely or randomly within the bacterial cells. Thus, even within the sub-micrometer distances of the bacterial intra-cellular space, the positions of the enzymes and their substrates are kept in check. Adding to this complexity is the secondary structure and sequence specificity that many, perhaps all, ribonucleases exhibit, including those that are responsible for “general” RNA degradation. In this review, the implications of these novel findings are discussed and specific examples from Staphylococcus aureus are analysed.


Similar content being viewed by others
References
Aït-Bara S, Carpousis AJ (2015) RNA degradosomes in bacteria and chloroplasts: classification, distribution and evolution of RNase E homologs. Mol Microbiol 97:1021–1135. doi:10.1111/mmi.13095
Bakshi S, Siryaporn A, Goulian M, Weisshaar JC (2012) Superresolution imaging of ribosomes and RNA polymerase in live Escherichia coli cells. Mol Microbiol 85:21–38. doi:10.1111/j.1365-2958.2012.08081.x
Britton RA, Wen T, Schaefer L et al (2007) Maturation of the 5′ end of Bacillus subtilis 16S rRNA by the essential ribonuclease YkqC/RNase J1. Mol Microbiol 63:127–138. doi:10.1111/j.1365-2958.2006.05499.x
Christen B, Abeliuk E, Collier JM et al (2011) The essential genome of a bacterium. Mol Syst Biol 7:528. doi:10.1038/msb.2011.58
Clarke JE, Kime L, Romero AD, McDowall KJ (2014) Direct entry by RNase E is a major pathway for the degradation and processing of RNA in Escherichia coli. Nucleic Acids Res 42:11733–11751. doi:10.1093/nar/gku808
Del Campo C, Ignatova Z (2015) Probing dimensionality beyond the linear sequence of mRNA. Curr Genet. doi:10.1007/s00294-015-0551-5
Elvekrog MM, Walter P (2015) Dynamics of co-translational protein targeting. Curr Opin Chem Biol 29:79–86. doi:10.1016/j.cbpa.2015.09.016
Figaro S, Durand S, Gilet L et al (2013) Bacillus subtilis mutants with knockouts of the genes encoding ribonucleases RNase Y and RNase J1 are viable, with major defects in cell morphology, sporulation, and competence. J Bacteriol 195:2340–2348. doi:10.1128/JB.00164-13
Giraud C, Hausmann S, Lemeille S et al (2015) The C-terminal region of the RNA helicase CshA is required for the interaction with the degradosome and turnover of bulk RNA in the opportunistic pathogen Staphylococcus aureus. RNA Biol 12:658–674. doi:10.1080/15476286.2015.1035505
Hunt A, Rawlins JP, Thomaides HB, Errington J (2006) Functional analysis of 11 putative essential genes in Bacillus subtilis. Microbiology 152:2895–2907. doi:10.1099/mic.0.29152-0
Khemici V, Poljak L, Luisi BF, Carpousis AJ (2008) The RNase E of Escherichia coli is a membrane-binding protein. Mol Microbiol 70:799–813. doi:10.1111/j.1365-2958.2008.06454.x
Khemici V, Prados J, Linder P, Redder P (2015) Decay-Initiating Endoribonucleolytic Cleavage by RNase Y Is Kept under tight control via Sequence Preference and Sub-cellular Localisation. PLoS Genet 11:e1005577. doi:10.1371/journal.pgen.1005577
Krogh A, Larsson B, von Heijne G, Sonnhammer EL (2001) Predicting transmembrane protein topology with a hidden Markov model: application to complete genomes. J Mol Biol 305:567–580. doi:10.1006/jmbi.2000.4315
Laalami S, Zig L, Putzer H (2014) Initiation of mRNA decay in bacteria. Cell Mol Life Sci 71:1799–1828. doi:10.1007/s00018-013-1472-4
Lehnik-Habrink M, Newman J, Rothe FM et al (2011) RNase Y in Bacillus subtilis: a Natively disordered protein that is the functional equivalent of RNase E from Escherichia coli. J Bacteriol 193:5431–5441. doi:10.1128/JB.05500-11
Linder P, Lemeille S, Redder P (2014) Transcriptome-wide analyses of 5′-ends in RNase J mutants of a gram-positive pathogen reveal a role in RNA maturation, regulation and degradation. PLoS Genet 10:e1004207. doi:10.1371/journal.pgen.1004207
Mackie GA (2013) RNase E: at the interface of bacterial RNA processing and decay. Nat Rev Microbiol 11:45–57. doi:10.1038/nrmicro2930
Montero Llopis P, Jackson AF, Sliusarenko O et al (2010) Spatial organization of the flow of genetic information in bacteria. Nature 466:77–81. doi:10.1038/nature09152
Murashko ON, Kaberdin VR, Lin-Chao S (2012) Membrane binding of Escherichia coli RNase E catalytic domain stabilizes protein structure and increases RNA substrate affinity. Proc Natl Acad Sci USA 109:7019–7024. doi:10.1073/pnas.1120181109
Ow MC, Kushner SR (2002) Initiation of tRNA maturation by RNase E is essential for cell viability in E. coli. Genes Dev 16:1102–1115. doi:10.1101/gad.983502
Redder P (2015) Using EMOTE to map the exact 5’-ends of processed RNA on a transcriptome-wide scale. Methods Mol Biol 1259:69–85. doi:10.1007/978-1-4939-2214-7_5
Redder P, Linder P (2012) New range of vectors with a stringent 5-fluoroorotic acid-based counterselection system for generating mutants by allelic replacement in Staphylococcus aureus. Appl Environ Microbiol 78:3846–3854. doi:10.1128/AEM.00202-12
Roux CM, DeMuth JP, Dunman PM (2011) Characterization of components of the Staphylococcus aureus mRNA degradosome holoenzyme-like complex. J Bacteriol 193:5520–5526. doi:10.1128/JB.05485-11
Shahbabian K, Jamalli A, Zig L, Putzer H (2009) RNase Y, a novel endoribonuclease, initiates riboswitch turnover in Bacillus subtilis. EMBO J 28:3523–3533. doi:10.1038/emboj.2009.283
Voss JE, Luisi BF, Hardwick SW (2014) Molecular recognition of RhlB and RNase D in the Caulobacter crescentus RNA degradosome. Nucleic Acids Res 42:13294–13305. doi:10.1093/nar/gku1134
Acknowledgments
I would like to thank Julien Prados for help with the bioinfomatics, and Patrick Linder for critical reading of the manuscript. Our research into RNA decay in S. aureus is funded by the Swiss Life Jubiläumsstiftung, the Novartis Consumer Health Foundation, the Ernst and Lucie Schmidheiny Foundation, the Swiss National Science Foundation and the Medical Faculty at the University of Geneva.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by M. Kupiec.
Rights and permissions
About this article
Cite this article
Redder, P. How does sub-cellular localization affect the fate of bacterial mRNA?. Curr Genet 62, 687–690 (2016). https://doi.org/10.1007/s00294-016-0587-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00294-016-0587-1