Abstract
The susceptible-infected-removed (SIR) and the susceptible-exposed-infected-removed (SEIR) epidemic models with constant parameters are adequate for describing the time evolution of seasonal diseases for which available data usually consist of fatality reports. The problems associated with the determination of system parameters starts with the inference of the number of removed individuals from fatality data, because the infection to death period may depend on health care factors. Then, one encounters numerical sensitivity problems for the determination of the system parameters from a correct but noisy representative of the number of removed individuals. Finally as the available data is necessarily a normalized one, the models fitting this data may not be unique. We prove that the parameters of the (SEIR) model cannot be determined from the knowledge of a normalized curve of “Removed” individuals and we show that the proportion of removed individuals, \(R(t)\), is invariant under the interchange of the incubation and infection periods and corresponding scalings of the contact rate. On the other hand we prove that the SIR model fitting a normalized curve of removed individuals is unique and we give an implicit relation for the system parameters in terms of the values of \(R_m/R_f\) and \(R^{\prime }_m/R_f\), where \(R_f\) is the steady state value of \(R(t)\) and \(R_m\) and \(R^{\prime }_m\) are the values of \(R(t)\) and its derivative at the inflection point \(t_m\) of \(R(t)\). We use these implicit relations to provide a robust method for the estimation of the system parameters and we apply this procedure to the fatality data for the H1N1 epidemic in the Czech Republic during 2009. We finally discuss the inference of the number of removed individuals from observational data, using a clinical survey conducted at major hospitals in Istanbul, Turkey, during 2009 H1N1 epidemic.











Similar content being viewed by others
References
Anderson RM, May RM (1979) Population biology of infectious diseases: part I. Nature 280:361–367
Balcan D, Colizza V, Goncalves B, Hu H, Ramasco JJ, Vespignani A (2009) Multiscale mobility networks and the spatial spreading of infectious diseases. Proc Natl Acad Sci USA 106(51):21484–21489
Barakat A, Ihazmad H, El Falaki F, Tempia S, Cherkaoui I, El Aouad R (2012) 2009 Pandemic influenza A virus subtype H1N1 in Morocco, 2009–2010: epidemiology, transmissibility, and factors associated with fatal cases. J Infect Dis 206(1):S94–S100
Bilge AH, Samanlioglu F, Ergonul O (2012) A susceptible-exposed-infected-removed (SEIR) model for the 2009–2010 A/H1N1 epidemic in Istanbul. arXiv:1205.2497v1 [q-bio.QM]
Boni MF, Manh BH, Thai PQ, Farrar J, Hien TT, Kinh NV, Horby P (2009) Modelling the progression of pandemic influenza A (H1N1) in Vietnam and the opportunities for reassortment with other influenza viruses. BMC Med 7:43
Chowell G, Nishiura H, Bettencourt LMA (2007) Comparative estimation of the reproduction number for pandemic influenza from daily case notification data. J R Soc Interf 4:155–166
Chowell G, Viboud C, Munayco CV, Gomez J, Simonsen L, Miller MA, Tamerius J, Fiestas V, Halsey ES, Laguna-Torres VA (2011) Spatial and temporal characteristics of the 2009 A/H1N1 influenza pandemic in Peru. Plos One 6(6):Article number e21287
Coelho FC, Codeco CT, Gomes MGM (2011) A Bayesian framework for parameter estimation in dynamical models. Plos One 6(5):e19616
Crow EL, Shimizu K (1988) Lognormal distributions:theory and applications. Marcel Dekker, New York
Diekmann O, Heesterbeek H, Britton T (2012) Mathamatical tools for understanding infectious disease dynamics. Princeton series in theoretical and computational biology
Ergönül Ö, Alan S, Ak Ö, Sargın F, Kantürk A, Gündüz A, Engin D, Öncül O, Balkan II, Ceylan B, Benzonana N, Yazıcı S, Şimşek F, Uzun N, Inan A, Gulhan E, Ciblak M, Midilli K, Ozyurt M, Badur S, Gencer S, Nazlıcan O, Özer S, Özgüneş N, Yıldırmak T, Aslan T, Göktaş P, Saltoǧlu N, Fincancı M, Dokucu AI, Eraksoy H, Turkish Society of Clinical Microbiology and Infectious Diseases (KLIMIK) Pandemic Influenza Study Group (2014) Predictors of fatality in pandemic influenza A (H1N1) virus infection among adults. BMC Infect Dis. 14(1):317. doi:10.1186/1471-2334-14-317
Goldstein E, Dushoff J, Ma J, Plotkin JB, Earn DJ, Lipsitch M (2009) Reconstructing influenza incidence by deconvolution of daily mortality time series. PNAS 106:21825–21829
Greenhalgh D (1992) Some results for a SEIR epidemic model with density dependence in the death rate. IMA J Math Appl Med Biol 9:67
Haghdoost AA, Gooya MM, Baneshi MR (2009) Modelling of H1N1 flu in Iran. Arch Iran Med 12(6):533–541
Hethcote HW (2000) The mathematics of infectious diseases. SIAM Rev. 42:599–653
Hsieh YH (2010) Pandemic influenza A (H1N1) during winter influenza season in the southern hemisphere. Influ Respir Virus 4(4):187–197
Jesan T, Menon GI, Sinha S (2011) Epidemiological dynamics of the 2009 influenza A(H1N1) outbreak in India. Curr Sci 100(7):1051–1054
Katriel G, Yaari R, Huppert A, Roll U, Stone L (2011) Modelling the initial phase of an epidemic using incidence and infection network data: 2009 H1N1 pandemic in Israel as a case study. J R Soc Interf 8(59):856–867
Kermack WO, McKendrick AG (1927) Contributions to the theory of epidemics-I. Proc R Soc 115A:700–721. (Reprinted in Bulletin of Mathematical Biology 1991; 53(1–2):33–55)
Lekone PE, Finkenstadt BF (2006) Statistical inference in a stochastic epidemic SEIR model with control intervention: Ebola as a case study. Biometrics 62:1170–1177
Li MY, Muldowney JS (1995) Global stability for the SEIR model in epidemiology. Math Biosci 125(2):155–164
Li MY, Muldowney JS, Van Den Driessche P (1999a) Global stability of SEIRS models in epidemiology. Can Appl Math Q 7(4):409–425
Li MY, Graef JR, Wang L, Karsai J (1999b) Global dynamics of a SEIR model with varying total population size. Math Biosci 160:191–213
Li MY, Smith HL, Wang L (2001) Global dynamics of an SEIR epidemic model with vertical transmission. SIAM J Appl Math 62:58
Li G, Jin Z (2005) Global stability of a SEIR epidemic model with infectious force in latent, infected and immune period. Chaos Solitons Fractals 25:1177–1184
Merler S, Ajelli M, Pugliese A, Ferguson NM (2011) Determinants of the spatiotemporal dynamics of the 2009 H1N1 pandemic in Europe: implications for real-time modelling. Plos Comput Biol 7(9):e1002205
Navarro-Robles E, Martinez-Matsushita L, Lopez-Molina R, Fritz-Hernandez J, Flores-Aldana BA, Mendoza-Perez JC (2012) Model to estimate epidemic patterns of influenza A (H1N1) in Mexico. Revista PanAmericana de Salud Publica-Pan. Am J Public Health 31(4):269–274
Paine S, Mercer GN, Kelly PM, Bandaranayake D, Baker MG, Huang QS, Mackereth G, Bissielo A, Glass K, Hope V (2010) Transmissibility of 2009 pandemic influenza A(H1N1) in New Zealand: effective reproduction number and influence of age, ethnicity and importations. Eurosurveillance 15(24):9–17
Presanis AM, De Angelis D, The New York City Swine Flu Investigation Team, Hagy A, Reed C, Riley S, Cooper BS, Finelli L, Biedrzycki P, Lipsitch M (2009) The severity of pandemic H1N1 influenza in the United States, from April to July 2009: a Bayesian analysis. PLoS Medicine 6(12):1–12. www.plosmedicine.org
Reed C, Angulo FJ, Swerdlow DL, Lipsitch M, Meltzer MI, Jernigan D, Finelli L (2009) Estimates of the prevalence of pandemic (H1N1) 2009, United States, April–July 2009. Emerg Infect Dis 15(12):2004–2007
Roberts MG, Nishiura H (2011) Early estimation of the reproduction number in the presence of imported cases: pandemic influenza H1N1-2009 in New Zealand. Plos One 6(5):Article number e17835
Shu H, Fan D, Wei J (2012) Global stability of multi-group SEIR epidemic models with distributed delays and nonlinear transmission. Nonlinear Anal Real World Appl 13(4):1581–1592
Trottier H, Philippe P (2002) Deterministic modeling of infectious diseases: applications to measles and other similar infections. Intern J Infect Dis 2(1)
Tuite AR, Greer AL, Whelan M, Winter AL, Lee B, Yan P, Wu JH, Moghadas S, Buckeridge D, Pourbohloul B, Fisman DN (2010) Estimated epidemiologic parameters and morbidity associated with pandemic H1N1 influenza. Can Med Assoc J 182(2):128–133
Van Kerkhove MD, Asikainen T, Becker NG, Bjorge S, Desenclos J, dos Santos T, Fraser C, Leung GM, Lipsitch M, Longini Jr IM, McBryde ES, Roth CE, Shay DK, Smith DJ, Wallinga J, White PJ, Ferguson NM, Riley S for the WHO Informal Network for Mathematical Modelling for Pandemic Influenza H1N1 2009 (Working Group on Data Needs) (2010) Studies needed to address public health challenges of the 2009 H1N1 influenza pandemic: insights from modeling. PLoS Medicine 7(6):1–6. www.plosmedicine.org
Wearing HJ, Rohani P, Keeling MJ (2005) Appropriate models for the management of infectious diseases. PLoS Medicine 2(7):e621–e627. www.plosmedicine.org
White LF, Wallinga J, Finelli L, Reed C, Riley S, Lipsitch M, Pagano M (2009) Estimation of the reproductive number and the serial interval in early phase of the 2009 influenza A/H1N1 pandemic in the USA. Influ Respir Virus 3(6):267–276
Zhang J, Ma Z (2003) Global dynamics of an SEIR epidemic model with saturating contact rate. Math Biosci 185:15–32
Author information
Authors and Affiliations
Corresponding author
Appendix A
Appendix A
We use Istanbul data to develop a statistical method for shifting the observed date of fatality back or forth in order to infer a correct representative of \(R(t)\). The first step is to show that the time-to-death distribution is nearly log-normal (Crow and Shimizu 1988). To proceed further, one has to start with an a priori estimate of the mean and the standard deviation of the logarithm of the time-to-death intervals. The next step is to partition the time axis into a reasonable number of bins bounded by \(0<ln(t_1)<\cdots <ln(t_n)<\infty \). The proportion of the cases with a duration belonging to a bin \([t_{i-1},t_i]\), denoted by \(p_i\) can be computed from the log-normal distribution. We then translate the time axis for the occurrence of fatalities to the day of the first fatality and compute the proportion \(q_i\) of the fatalities that occur in the bin \([t_{i-1},t_i]\). By a simple pigeon hole principle, all fatality cases with duration belonging to the last bin, should have been occurred at the last bin. By similar arguments that will be illustrated below, we obtain a system of under-determined equations with inequality constraints, that we solve as an optimization problem, to determine the proportion \(\alpha _{i,j}\) of fatalities that occur in bin \(i\) to have duration belonging to the bin \(j\). The final step is to move the cases in the bin \(i\) back or forth according to the parameters \(\alpha _{i,j}\). We note that this process aims only to correct for excessively long hospitalization periods.
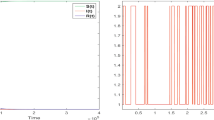
Working with Istanbul data, in Fig. 12a, b, we present the histograms for the time-to-death durations with respect to time and with respect to the logarithm of time, measured in days. The first figure seems to have an exponential decay, while the second one is centered in the 7–12 day interval. The mean, standard deviation and higher central moments are computed as
The Jarque–Bera test statistic with 0.05 confidence interval (with value \(JB=0.093\)) allows us to consider the time-to-death duration as log-normal distributed.
The histogram of the time-to-death intervals with respect to days (a). The histogram of the time-to-death intervals with respect to the logarithm of days. The distribution is approximately normal, centered at \(\mu =2.49\) (7–12 days), with standard deviation \(\sigma =0.94\), skewness 0.026 and kurtosis 3.21 (b)
We divide the logarithmic time scale into 6 bins separated by
The probabilities \(p_i\) that the duration of time-to-death falls in these bins are computed from the log-normal distribution with parameters above as
The proportion \(q_i\) of deaths that occur in these bins is an observable quantity. In the case of Istanbul data these are observed as
As above, let \(\alpha _{i,j}\) be the contribution of cases with duration in the \(j\)th bin to fatalities occurring in the \(i\)th bin. Clearly, longer durations cannot contribute to earlier fatalities. Thus \(\alpha _{i,j}=0\) for \(i<j\). If the number of bins is \(n\), the number of parameters is \(n(n+1)/2\) subject to the constraints \(0<\alpha _{i,j}<1\) and \(\sum _j \alpha _{i,j}=1\). With the bin selection as above, we aim to approximate the \(q_i\)’s by the expressions at the right hand sides of the system below.
together with the constraints
where \(k_0>0\) may be a non-zero lower bound. We search for an optimal solution minimizing the least-square error between the right and left hand sides of the expressions above, subject to the constraints. For the example under consideration, the matrices \(\alpha _{i,j}\) for lower bounds \(k_0=0\) and \(k_0=0.1\) are given below.
We define the percentage contribution of the cases with duration in the \(j\)th bin, to fatalities occurring in the \(i\)th bin, as
For the \(\alpha _{i,j}{(1)}\) matrix this gives
Note that the entries of the \(H_{i,j}\) can be interpreted as conditional probabilities. We see that contributions fatalities that occur late have longer durations. In order to construct a curve that approximates more closely the time evolution of the number of removed individuals, one can shift back the proportion \(H_{i,j}\) of the date of fatalities in the \(i\)th bin, by some value in the \(j\)th bin, then add the incubation period. If the incubation period is unknown, the process described above will give a time translate of the correct curve. For example, we have to move two thirds of the fatalities occurring later than the 54th day of the epidemic, by an amount between 54 and 148 days. Similarly, \(9/13\)th of the fatalities occurring between the 20th and 54th days of the epidemic have to be moved back.
We would like to note that the method proposed here is only a preprocessing for obtaining a better approximation to the number of removed individuals. The analysis of the previous sections have to be carried out in order to obtain the incidence curves from fatality data.
Rights and permissions
About this article
Cite this article
Bilge, A.H., Samanlioglu, F. & Ergonul, O. On the uniqueness of epidemic models fitting a normalized curve of removed individuals. J. Math. Biol. 71, 767–794 (2015). https://doi.org/10.1007/s00285-014-0838-z
Received:
Revised:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00285-014-0838-z