Abstract
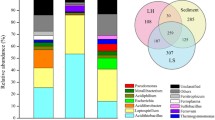
The microbial communities are important for minerals decomposition in biological heap leaching system. However, the differentiation and relationship of composition and function of microbial communities between leaching heap (LH) and leaching solution (LS) are still unclear. In this study, 16S rRNA gene sequencing was used to assess the microbial communities from the two subsystems in ZiJinShan copper mine (Fujian province, China). Results of PCoA and dissimilarity test showed that microbial communities in LH samples were significantly different from those in LS samples. The dominant genera of LH was Acidithiobacillus (57.2 ∼ 87.9 %), while Leptospirillum (48.6 ∼ 73.7 %) was predominant in LS. Environmental parameters (especially pH) were the major factors to influence the composition and structure of microbial community by analysis of Mantel tests. Results of functional test showed that microbial communities in LH utilized sodium thiosulfate more quickly and utilized ferrous sulfate more slowly than those in LS, which further indicated that the most sulfur-oxidizing processes of bioleaching took place in LH and the most iron-oxidizing processes were in LS. Further study found that microbial communities in LH had stronger pyrite leaching ability, and iron extraction efficiency was significantly positively correlated with Acidithiobacillus (dominated in LH), which suggested that higher abundance ratio of sulfur-oxidizing microbes might in favor of minerals decomposition. Finally, a conceptual model was designed through the above results to better exhibit the sulfur and iron metabolism in bioleaching systems.






Similar content being viewed by others
References
Anderson I, Chertkov O, Chen A, Saunders E, Lapidus A (2012) Complete genome sequence of the moderately thermophilic mineral-sulfide-oxidizing firmicute Sulfobacillus acidophilus type strain (NAL (T)). Stand Genomic Sci 6(3):293–303. doi:10.4056/sigs.2736042
Dixon P (2003) VEGAN, a package of R functions for community ecology. J Vegetation Sci 14(6):927–930. doi:10.1111/j.1654-1103.2003.tb02228.x
Ehrlich HL (2001) Past, present and future of biohydrometallurgy. Hydrometallurgy 59:127–134. doi:10.1016/S0304-386X(00)00165-1
García-Moyano A, González-Toril E, Moreno-Paz M, Parro V, Amils R (2007) Microbial ecology of Leptospirillum spp. in Río Tinto, a model of interest to biohydrometallurgy. Adv Mater Res 20-21:409–412. doi:10.4028/www.scientific.net/AMR.20-21.409
Goltsman DSA, Denef VJ, Singer SW, Verberkmoes NC, Mark L (2009) Community genomic and proteomic analyses of chemoautotrophic iron-oxidizing "Leptospirillum rubarum" (Group II) and "Leptospirillum ferrodiazotrophum" (Group III) bacteria in acid mine drainage biofilms. Appl Environ Microb 75(13):4599–4615. doi:10.1128/AEM.02943-08
Griffiths BS, Ritz K, Bardgett RD, Cook R, Christensen S (2000) Ecosystem response of pasture soil communities to fumigation-induced microbial diversity reductions: an examination of the biodiversity–ecosystem function relationship. Oikos 90(2):279–294. doi:10.1007/s00248-002-2043-7
Hu Q, Guo X, Liang Y, Hao X, Ma L, Yin H, Liu X (2015) Comparative metagenomics reveals microbial community differentiation in a biological heap leaching system. Res Microbiol 166(6):525–534. doi:10.1016/j.resmic.2015.06.005
Huttenhower C, Gevers D, Knight R, Abubucker S, Badger JH (2012) Structure, function and diversity of the healthy human microbiome. Nature 486(7402):207–214. doi:10.1038/nature11234
Johnson DB (2008) Biodiversity and interactions of acidophiles: key to understanding and optimizing microbial processing of ores and concentrates. T Nonferr Metal Soc 18(6):1367–1373. doi:10.1016/S1003-6326(09)60010-8
Johnson DB (1998) Biodiversity and ecology of acidophilic microorganisms. FEMS Microbiol Ecol 27(4):307–317. doi:10.1016/S0168-6496(98)00079-8
Kuang JL, Huang LN, Chen LX, Hua ZS, Li SJ, Hu M, Li JT, Shu WS (2013) Contemporary environmental variation determines microbial diversity patterns in acid mine drainage. Isme J 7(5):1038–1050. doi:10.1038/ismej.2012.139
Li T, Meng L, Gatesoupe FJ, Zhang Q, Li A, Gong X (2015) Comparative analysis of the intestinal bacterial communities in different species of carp by pyrosequencing. Microb Ecol 69(1):25–36. doi:10.1007/s00248-014-0480-8
Liu X, Chen B, Wen J, Renman R (2010) Leptospirillum forms a minor portion of the population in Zijinshan commercial non-aeration copper bioleaching heap identified by 16S rRNA clone libraries and real-time PCR. Hydrometallurgy 104(3):399–403. doi:10.1016/j.hydromet.2010.03.024
Liu Y, Yin H, Liang Y, Shen L, Liu Y (2011) Changes in the composition of an acid mine drainage microbial community upon successive transfers in medium containing low-grade copper sulfide. Bioresource Technol 102(20):9388–9394. doi:10.1016/j.biortech.2011.05.095
Loranger H, Weisser WW, Ebeling A, Eggers T, Luca ED, Loranger J, Roscher C, Meyer ST (2013) Invertebrate herbivory increases along an experimental gradient of grassland plant diversity. Oecologia 174(1):1–11. doi:10.1007/s00442-013-2741-5
Mendez-Garcia C, Pelaez AI, Mesa V, Sanchez J, Golyshina OV, Ferrer M (2015) Microbial diversity and metabolic networks in acid mine drainage habitats. Front Microbiol 6:475. doi:10.3389/fmicb.2015.00475
Ouml BZ, Erkan S, Pauliina N, Kaksonen AH, Puhakka JA (2007) Kinetics of iron oxidation by Leptospirillum ferriphilum dominated culture at pH below one. Biotechnol Bioeng 97(5):1121–1127. doi:10.1002/bit.21313
Peres-Neto PR, Jackson DA (2001) How well do multivariate data sets match? The advantages of a Procrustean superimposition approach over the Mantel test. Oecologia 129(2):169–178. doi:10.1007/s004420100720
Pielou EC (1977) Mathematical ecology. J Anim Ecol 47
Qiu GZ, Wan MX, Qian L, Huang ZY, Liu K, Liu XD, Shi WY, Yang Y (2008) Archaeal diversity in acid mine drainage from Dabaoshan Mine, China. J Basic Microbiol 48(5):401–409. doi:10.1002/jobm.200800002
Rodríguez Y, Ballester A, Blázquez ML, González F, Muñoz JA (2003b) New information on the pyrite bioleaching mechanism at low and high temperature. Hydrometallurgy 71(1–2):37–46. doi:10.1016/S0304-386X(03)00172-5
Rodríguez Y, Ballester A, Blázquez ML, González F, Muñoz JA (2003a) New information on the chalcopyrite bioleaching mechanism at low and high temperature. Hydrometallurgy 71(1–2):47–56. doi:10.1016/S0304-386X(03)00173-7
Shannon CE (1948) A mathematical theory of communication. Bell System Technical Journal 27(3):379–423. doi:10.1145/584091.584093
Smith TB, Wayne RK, Girman DJ, Bruford MW (1997) A role for ecotones in generating rainforest biodiversity. Science 276(5320):1855–1857. doi:10.1126/science.276.5320.1855
Sophie W, Valérie D, Prosser JI, Franck P, Claire C, Nadine G, Le RX (2007) Decline of soil microbial diversity does not influence the resistance and resilience of key soil microbial functional groups following a model disturbance. Environ Microbiol 9(9):2211–2219. doi:10.1111/j.1462-2920.2007.01335.x
Tamaki H, Tanaka Y, Matsuzawa H, Muramatsu M, Meng XY, Hanada S, Mori K, Kamagata Y (2011) Armatimonas rosea gen. nov., sp. nov., of a novel bacterial phylum, Armatimonadetes phyl. nov., formally called the candidate phylum OP10. Int J Syst Evol Micr 61(Pt6):1442–1447. doi:10.1099/ijs.0.025643-0
Tom C, Elke V, Emly S, Enevold F, Peter L, Peter V (2005) Advenella incenata gen. nov., sp nov., a novel member of the Alcaligenaceae, isolated from various clinical samples. Int J Syst Evol Micr 55(Part 1):251–256. doi:10.1099/ijs.0.63267-0
Troesch A, Duperray A, Polack B, Marguerie G (1990) Comparative study of the glycosylation of platelet glycoprotein GPIIb/GPIIa and the vitronectin receptor. Biochem J 268. doi:10.1042/bj2680129
Vardanyan NS, Vardanyan AK (2014) New sulphur oxidizing bacteria isolated from bioleaching pulp of zinc and copper concentrates. Univ J Microbiol Res 2(2):27–31. doi:10.13189/ujmr.2014.020201
Watling HR, Collinson DM, Shiers DW, Bryan CG, Watkin ELJ (2013) Effects of pH, temperature and solids loading on microbial community structure during batch culture on a polymetallic ore. Miner Eng 48:68–76. doi:10.1016/j.mineng.2012.10.014
Wikström P, Andersson AC, Forsman M (1999) Biomonitoring complex microbial communities using random amplified polymorphic DNA and principal component analysis. FEMS Microbiol Ecol 28(2):131–139. doi:10.1111/j.1574-6941.1999.tb00568.x
Williams KP, Kelly DP (2013) Proposal for a new class within the phylum Proteobacteria, Acidithiobacillia classis nov., with the type order Acidithiobacillales, and emended description of the class Gammaproteobacteria. Int J Syst Evol Micr 63(4):2901–2906. doi:10.1099/ijs.0.049270-0
Xiao Y, Xu Y, Dong W, Liang Y, Fan F (2015) The complicated substrates enhance the microbial diversity and zinc leaching efficiency in sphalerite bioleaching system. Appl Microbiol Biotechnol 99(23):10311–10322. doi:10.1007/s00253-015-6881-x
Yang Y, Yang LI, Sun QY (2014) Archaeal and bacterial communities in acid mine drainage from metal-rich abandoned tailing ponds, Tongling, China. Tran Nonferr Metal Soc 24(10):3332–3342. doi:10.1016/S1003-6326(14)63474-9
Yin H, Cao L, Qiu G, Wang D, Kellogg L (2008) Molecular diversity of 16S rRNA and gyrB genes in copper mines. Arch Microbiol 189(2):101–110. doi:10.1007/s00203-007-0298-6
Yin H, Niu J, Ren Y, Cong J, Zhang X, Fan F, Xiao Y, Zhang X, Deng J, Xie M (2015) An integrated insight into the response of sedimentary microbial communities to heavy metal contamination. Sci Rep 22:93–102. doi:10.1038/srep14266
Zhang L, Mao F, Li K, Wang Y, Chen X, Zhou H (2015) Enhancement in copper extraction from chalcopyrite by re-inoculation of different acidophilic, moderately thermophilic microorganisms. Hydrometallurgy 156:142–151. doi:10.1016/j.hydromet.2015.06.009
Zhang X, Niu J, Liang Y, Liu X, Yin H (2016) Metagenome-scale analysis yields insights into the structure and function of microbial communities in a copper bioleaching heap. BMC Genet 17:1–12. doi:10.1186/s12863-016-0330-4
Acknowledgments
The study was supported by the National Nature Science Foundation of China (No. 31570113 and No. 41573072). Thanks to prof. Huaqun Yin and Xueduan Liu who helped design this study and contributed material essential for the study, to Weiling Dong, Liyuan Ma, Xiaodong Hao Yili Liang, Yabin Gu, and Siyuan She for their help finish this experiment, and to Jiaojiao Niu and Xian Zhang for data analysis.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
This article does not contain any studies with human participants or animals performed by any of the authors.
Conflict of interest
The authors declare that they have no competing interests.
Electronic supplementary material
ESM 1
(PDF 649 kb)
Rights and permissions
About this article
Cite this article
Xiao, Y., Liu, X., Ma, L. et al. Microbial communities from different subsystems in biological heap leaching system play different roles in iron and sulfur metabolisms. Appl Microbiol Biotechnol 100, 6871–6880 (2016). https://doi.org/10.1007/s00253-016-7537-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-016-7537-1