Abstract
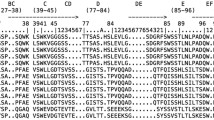
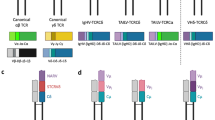
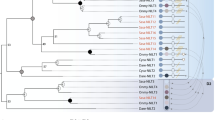
In testing the hypothesis that all jawed vertebrate classes employ immunoglobulin heavy chain V (IgHV) gene segments in their T cell receptor (TCR)δ encoding loci, we found that some basic characterization was required of zebrafish TCRδ. We began by annotating and characterizing the TCRα/δ locus of Danio rerio based on the most recent genome assembly, GRCz10. We identified a total of 141 theoretically functional V segments which we grouped into 41 families based upon 70 % nucleotide identity. This number represents the second greatest count of apparently functional V genes thus far described in an antigen receptor locus with the exception of cattle TCRα/δ. Cloning, relative quantitative PCR, and deep sequencing results corroborate that zebrafish do express TCRδ, but these data suggest only at extremely low levels and in limited diversity in the spleens of the adult fish. While we found no evidence for IgH-TCRδ rearrangements in this fish, by determining the locus organization we were able to suggest how the evolution of the teleost α/δ locus could have lost IgHVs that exist in sharks and frogs. We also found evidence of surprisingly low TCRδ expression and repertoire diversity in this species.







Similar content being viewed by others
References
Agard EA, Lewis SM (2000) Postcleavage sequence specificity in V(D)J recombination. Mol Cell Biol 20:5032–5040
Amemiya CT, Alfoldi J, Lee AP, Fan S, Philippe H, Maccallum I, Braasch I, Manousaki T, Schneider I, Rohner N, Organ C, Chalopin D, Smith JJ, Robinson M, Dorrington RA, Gerdol M, Aken B, Biscotti MA, Barucca M, Baurain D, Berlin AM, Blatch GL, Buonocore F, Burmester T, Campbell MS, Canapa A, Cannon JP, Christoffels A, De Moro G, Edkins AL, Fan L, Fausto AM, Feiner N, Forconi M, Gamieldien J, Gnerre S, Gnirke A, Goldstone JV, Haerty W, Hahn ME, Hesse U, Hoffmann S, Johnson J, Karchner SI, Kuraku S, Lara M, Levin JZ, Litman GW, Mauceli E, Miyake T, Mueller MG, Nelson DR, Nitsche A, Olmo E, Ota T, Pallavicini A, Panji S, Picone B, Ponting CP, Prohaska SJ, Przybylski D, Saha NR, Ravi V, Ribeiro FJ, Sauka-Spengler T, Scapigliati G, Searle SM, Sharpe T, Simakov O, Stadler PF, Stegeman JJ, Sumiyama K, Tabbaa D, Tafer H, Turner-Maier J, van Heusden P, White S, Williams L, Yandell M, Brinkmann H, Volff JN, Tabin CJ, Shubin N, Schartl M, Jaffe DB, Postlethwait JH, Venkatesh B, Di Palma F, Lander ES, Meyer A, Lindblad-Toh K (2013) The African coelacanth genome provides insights into tetrapod evolution. Nature 496:311–316
André S, Kerfourn F, Affaticati P, Guerci A, Ravassard P, Fellah JS (2007) Highly restricted diversity of TCR delta chains of the amphibian Mexican axolotl (Ambystomamexicanum) in peripheral tissues. Eur J Immunol 37:1621–1633
Bonneville M, O’Brien RL, Born WK (2010) γδ T cell effector functions: a blend of innate programming and acquired plasticity. Nat Rev Immunol 10:467–478
Buonocore F, Castro R, Randelli E, Lefranc MP, Six A, Kuhl H, Reinhardt R, Facchiano A, Boudinot P, Scapigliati G (2012) Diversity, molecular characterization and expression of T cell receptor gamma in a teleost fish, the sea bass (Dicentrarchus labrax, L). PLoS One 7, e47957
Chakrabarti S, Streisinger G, Singer F, Walker C (1983) Frequency of gamma-Ray induced specific locus and recessive lethal mutations in mature germ cells of the zebrafish, Brachydanio rerio. Genetics 103:109–123
Chambers JM, Freeny A, Heiberger RM (1992) Analysis of variance; designed experiments. In Hastie JMCaTJ (ed.) Statistical models in S. Wadsworth & Brooks/Cole
Connelley TK, Degnan K, Longhi CW, Morrison WI (2014) Genomic analysis offers insights into the evolution of the bovine TRA/TRD locus. BMC Genomics 15:994
Criscitiello MF (2014) What the shark immune system can and cannot provide for the expanding design landscape of immunotherapy. Expert Opin Drug Disc 9:725–739
Criscitiello MF, Kamper SM, McKinney EC (2004a) Allelic polymorphism of TCRalpha chain constant domain genes in the bicolor damselfish. Dev Comp Immunol 28:781–792
Criscitiello MF, Wermenstam NE, Pilstrom L, McKinney EC (2004b) Allelic polymorphism of T-cell receptor constant domains is widespread in fishes. Immunogenetics 55(12):818–824
Criscitiello MF, Ohta Y, Saltis M, McKinney EC, Flajnik MF (2010) Evolutionarily conserved TCR binding sites, identification of T cells in primary lymphoid tissues, and surprising trans-rearrangements in nurse shark. J Immunol 184:6950–6960
Danilova N, Hohman VS, Sacher F, Ota T, Willett CE, Steiner LA (2004) T cells and the thymus in developing zebrafish. Dev Comp Immunol 28:755–767
Fischer C, Bouneau L, Ozouf-Costaz C, Crnogorac-Jurcevic T, Weissenbach J, Bernot A (2002) Conservation of the T-cell receptor α/δ linkage in the teleost fish Tetraodon nigroviridis. Genomics 79:241–248
Haire RN, Rast JP, Litman RT, Litman GW (2000) Characterization of three isotypes of immunoglobulin light chains and T-cell antigen receptor α in zebrafish. Immunogenetics 51:915–923
Harper CCL (2011) The laboratory zebrafish. CRC, New York
Hein WR, Mackay CR (1991) Prominence of gamma delta T cells in the ruminant immune system. Immunol Today 12:30–34
Holderness J, Hedges JF, Ramstead A, Jutila MA (2013) Comparative biology of gammadelta T cell function in humans, mice, and domestic animals. Annu Rev Anim Biosci 1:99–124
Hope RM (2013) Rmisc: Rmisc: Ryan Miscellaneous
Itohara S, Farr AG, Lafaille JJ, Bonneville M, Takagaki Y, Haas W, Tonegawa S (1990) Homing of a y8 thymocyte subset with homogeneous T-cell receptors to mucosal epithelia. Nature 343:754–757
Iwanami N (2014) Zebrafish as a model for understanding the evolution of the vertebrate immune system and human primary immunodeficiency. Exp Hematol 42:697–706
Kamper SM, McKinney CE (2002) Polymorphism and evolution in the constant region of the T-cell receptor beta chain in an advanced teleost fish. Immunogenetics 53:1047–1054
Kearse M, Moir R, Wilson A, Stones-Havas S, Cheung M, Sturrock S, Buxton S, Cooper A, Markowits S, Duran C, Thierer T, Ashton B, Mentijies P, Drummond AA (2012) Geneious basic: an integrated and extendable desktop software platform for the organization and analysis of sequence data. Bioinformatics 28:1647–1649
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT method. Methods 25:402–408
Moulana M, Taylor EB, Edholm ES, Quiniou SM, Wilson M, Bengten E (2014) Identification and characterization of TCRgamma and TCRdelta chains in channel catfish, Ictalurus punctatus. Immunogenetics 66:545–561
Murphy K (2012) Janeway’s immunobiology. Garland Science, New York
Nam B-H, Hirono I, Aoki T (2003) The four TCR genes of teleost fish: the cDNA and genomic DNA analysis of Japanese flounder (Paralichthys olivaceus) TCR α-, β-, γ-, and δ-chains. J Immunol 170:3081–3090
Parra ZE, Miller RD (2012) Comparative analysis of the chicken TCR alpha/delta locus. Immunogenetics 64:641–645
Parra ZE, Baker ML, Schwarz RS, Deakin JE, Lindblad-Toh K, Miller RD (2007) A unique T cell receptor discovered in marsupials. Proc Natl Acad Sci U S A 104:9776–9781
Parra ZE, Baker ML, Hathaway J, Lopez AM, Trujillo J, Sharp A, Miller RD (2008) Comparative genomic analysis and evolution of the T cell receptor loci in the opossum Monodelphis domestica. BMC Genomics 9:111
Parra ZE, Ohta Y, Criscitiello MF, Flajnik MF, Miller RD (2010) The dynamic TCRdelta: TCRdelta chains in the amphibian Xenopus tropicalis utilize antibody-like V genes. Eur J Immunol 40(8):2319–2329
Parra ZE, Lillie M, Miller RD (2012a) A model for the evolution of the mammalian T-cell Receptor alpha/delta and mu Loci based on evidence from the duckbill platypus. Mol Biol Evol 29:3205–3214
Parra ZE, Mitchell K, Dalloul RA, Miller RD (2012b) A second TCRdelta locus in Galliformes uses antibody-like V domains: insight into the evolution of TCRdelta and TCRmu genes in tetrapods. J Immunol 188:3912–3919
Partula S, De Guerra A, Fellah JS, Charlemagne J (1995) Structure and diversity of the T cell antigen receptor beta-chain in a teleost fish. J Immunol 155:699–706
Pascual V, Capra JD (1991) Human immunoglobulin heavy-chain variable region genes: organization, polymorphism, and expression. Adv Immunol 49:1–74
R Core Team (2014) R: A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna
Rast JP, Litman GW (1994) T-cell receptor gene homologs are present in the most primitive jawed vertebrates. Proc Natl Acad Sci U S A 91:9248–9252
Rast JP, Anderson MK, Strong SJ, Luer C, Litman RT, Litman GW (1997) α, β, γ, and δ T cell antigen receptor genes arose early in vertebrate phylogeny. Immunity 6:1–11
Schorpp M, Bialecki M, Diekhoff D, Walderich B, Odenthal J, Maischein HM, Zapata AG, Boehm T (2006) Conserved functions of Ikaros in vertebrate lymphocyte development: genetic evidence for distinct larval and adult phases of T cell development and two lineages of B cells in zebrafish. J Immunol 177:2463–2476
Shang N, Sun XF, Hu W, Wang YP, Guo QL (2008) Molecular cloning and characterization of common carp (Cyprinus carpio L.) TCRgamma and CD3gamma/delta chains. Fish Shellfish Immunol 24:412–425
Tamura K, Stecher G, Peterson D, Filipski A, Kumar S (2013) MEGA6: Molecular evolutionary genetics analysis version 6.0. Mol Biol Evol 30:2725–2729
Tian JY, Qi ZT, Wu N, Chang MX, Nie P (2014) Complementary DNA sequences of the constant regions of T-cell antigen receptors alpha, beta and gamma in mandarin fish, Siniperca chuatsi Basilewsky, and their transcriptional changes after stimulation with Flavobacterium columnare. J Fish Dis 37:89–101
Venkatesh B, Lee AP, Ravi V, Maurya AK, Lian MM, Swann JB, Ohta Y, Flajnik MF, Sutoh Y, Kasahara M, Hoon S, Gangu V, Roy SW, Irimia M, Korzh V, Kondrychyn I, Lim ZW, Tay BH, Tohari S, Kong KW, Ho S, Lorente-Galdos B, Quilez J, Marques-Bonet T, Raney BJ, Ingham PW, Tay A, Hillier LW, Minx P, Boehm T, Wilson RK, Brenner S, Warren WC (2014) Elephant shark genome provides unique insights into gnathostome evolution. Nature 505:174–179
Walker C, Streisinger G (1983) Induction of mutations by gamma-Rays in pregonial germ cells of zebrafish embryos. Genetics 103:125–136
Wang K, Gan L, Kunisada T, Lee I, Yamagishi H, Hood L (2001) Characterization of the Japanese pufferfish (Takifugu rubripes) T-cell receptor alpha locus reveals a unique genomic organization. Immunogenetics 53:31–42
Wang X, Parra ZE, Miller RD (2011) Platypus TCRmu provides insight into the origins and evolution of a uniquely mammalian TCR locus. J Immunol 187:5246–5254
Wang F, Ekiert DC, Ahmad I, Yu W, Zhang Y, Bazirgan O, Torkamani A, Raudsepp T, Mwangi W, Criscitiello MF, Wilson IA, Schultz PG, Smider VV (2013) Reshaping antibody diversity. Cell 153:1379–1393
Weir H, Chen PL, Deiss TC, Jacobs N, Nabity MB, Young MH, Criscitiello MF (2015) DNP-KLH Yields changes in leukocyte populations and immunoglobulin isotype use with different immunization routes in zebrafish. Front Immunol 6, 606
Wickham H (2009) ggplot2: elegant graphics for data analysis. Springer, New York
Yazawa R, Cooper GA, Beetz-Sargent M, Robb A, McKinnel L, Davidson WS, Koop BF (2008a) Functional adaptive diversity of the Atlantic salmon T-cell receptor gamma locus. Mol Immunol 45:2150–2157
Yazawa R, Cooper GA, Hunt P, Beetz-Sargent M, Robb A, Conrad M, McKinnel L, So S, Jantzen S, Phillips RB, Davidson WS, Koop BF (2008b) Striking antigen recognition diversity in the Atlantic salmon T-cell receptor alpha/delta locus. Dev Comp Immunol 32:204–212
Author information
Authors and Affiliations
Corresponding author
Additional information
This work was supported by the National Science Foundation through a grant to MFC (IOS 1257829).
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Seelye, S.L., Chen, P.L., Deiss, T.C. et al. Genomic organization of the zebrafish (Danio rerio) T cell receptor alpha/delta locus and analysis of expressed products. Immunogenetics 68, 365–379 (2016). https://doi.org/10.1007/s00251-016-0904-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00251-016-0904-3