Abstract
The major histocompatibility complex (MHC) includes highly polymorphic gene families encoding proteins crucial to the vertebrate acquired immune system. Classical MHC class I (MHCI) genes code for molecules expressed on the surfaces of most nucleated cells and are associated with defense against intracellular pathogens, such as viruses. These genes have been studied in a few wild bird species, but have not been studied in long-distance migrating shorebirds. Red Knots Calidris canutus are medium-sized, monogamous sandpipers with migratory routes that span the globe. Understanding how such long-distance migrants protect themselves from disease has gained new relevance since the emergence of avian-borne diseases, including intracellular pathogens recognized by MHCI molecules, such as avian influenza. In this study, we characterized MHCI genes in knots and found 36 alleles in eight individuals and evidence for six putatively functional and expressed MHCI genes in a single bird. We also found evidence for recombination and for positive selection at putative peptide binding sites in exons 2 and 3. These results suggest surprisingly high MHC diversity in knots, given their demographic history. This may be a result of selection from diverse pathogens encountered by shorebirds throughout their annual migrations.



Similar content being viewed by others
References
Afanassieff M, Goto RM, Ha J, Sherman MA, Zhong L, Auffray C, Coudert F, Zoorob R, Miller MM (2001) At least one class I gene in restriction fragment pattern-Y (Rfp-Y), the second MHC gene cluster in the chicken, is transcribed, polymorphic, and shows divergent specialization in antigen binding region. J Immunol 166:3324–3333
Alcaide M, Edwards SV, Cadahía L, Negro JJ (2009) MHC class I genes of birds of prey: isolation, polymorphism and diversifying selection. Cons Gen 10:1349–1355
Altizer S, Bartel R, Han BA (2011) Animal migration and infectious disease risk. Science 331:296–302
Baker AJ, Pereira SL, Paton TA (2007) Phylogenetic relationships and divergence times of Charadriiformes genera: multigene evidence for the Cretaceous origin of at least 14 clades of shorebirds. Biol Lett 3:205–209
Balakrishnan CN, Ekblom R, Völker M, Westerdahl H, Godinez R, Kotkiewicz H, Burt DW, Graves T, Griffin DK, Warren WC, Edwards SV (2010) Gene duplication and fragmentation in the zebra finch major histocompatibility complex. BMC Biol 8:29. doi:10.1186/1741-7007-8-29
Betts M, Russell R (2003) Amino acid properties and consequences of substitutions. In: Barnes M, Gray I (eds) Bioinformatics for geneticists. Wiley, Chichester, pp 289–316
Bjorkman P, Saper M, Samraoui B, Bennett W, Strominger J, Wiley D (1987) The foreign antigen binding site and T cell recognition regions of class I histocompatibility antigens. Nature 329:512–518
Bjorkman PJ, Parham P (1990) Structure, function, and diversity of class I major histocompatibility complex molecules. Ann Rev Biochem 59:253–388
Bonneaud C, Sorci G, Morin V, Westerdahl H, Zoorob R, Wittzell H (2004) Diversity of Mhc class I and IIB genes in house sparrows (Passer domesticus). Immunogenetics 55:855–865
Brown JD, Luttrell MP, Berghaus RD, Kistler W, Keeler SP, Howey A, Wilcox B, Hall J, Niles L, Dey A, Knutsen G, Fritz K, Stallknecht DE (2010) Prevalence of antibodies to type A influenza virus in wild avian species using two serologic assays. J Wildlife Dis 46:896–911
Buehler, D. M. (2003) Phylogeography and genetic diversity in Red Knots (Calidris canutus). MSc Thesis. Department of Zoology. Department of Zoology. University of Toronto, Toronto.
Buehler DM, Baker AJ (2005) Population divergence times and historical demography in Red Knots and Dunlins. Condor 107:497–513
Buehler DM, Baker AJ, Piersma T (2006) Reconstructing palaeoflyways of the late Pleistocene and early Holocene Red Knot (Calidris canutus). Ardea 94:485–498
Buehler DM, Piersma T (2008) Travelling on a budget: predictions and ecological evidence for bottlenecks in the annual cycle of long-distance migrants. Phil Trans R Soc Lond B 363:247–266
Buehler DM, Tieleman BI, Piersma T (2010) How do migrant species stay healthy over the annual cycle? A conceptual model for immune function and disease resistance. Int Comp Biol 50:346–357
Cheng Y, Stuart A, Morris K, Taylor R, Siddle HV, Deakin JE, Jones M, Amemiya CT, Belov K (2012) Antigen-presenting genes and genomic copy number variations in the Tasmanian devil MHC. BMC Genomics 13:87
Cloutier A, Mills J, Baker A (2011) Characterization and locus-specific typing of MHC class I genes in the red-billed gull (Larus scopulinus) provides evidence for major, minor, and nonclassical loci. Immunogenetics. doi:10.1007/s00251-011-0516-x
Cresswell P, Ackerman AL, Giodini A, Peaper DR, Wearsch PA (2005) Mechanisms of MHC class I-restricted antigen processing and cross-presentation. Immunol Rev 207:145–157
Dean A, Greenwald J (1995) Use of filtered pipet tips to elute DNA from agarose gels. Biotechniques 18:980
Drummond, A. J., Ashton, B., Buxton, S., Cheung, M., Cooper, A., Duran, C., Field, M., Heled, J., Kearse, M., Markowitz, S., Moir, R., Stones-Havas, S., Sturrock, S., Thierer, T., and Wilson, A. (2011) Geneious v5.4. http://www.geneious.com, pp. Available from www.geneious.com.
Eimes J, Bollmer J, Whittingham L, Johnson J, Van Oosterhout C, Dunn P (2011) Rapid loss of MHC class II variation in a bottlenecked population is explained by drift and loss of copy number variation. J Evol Biol. doi:10.1111/j.1420-9101.2011.02311.x
Grossberger D, Parham P (1992) Reptilian class I major histocompatibility complex genes reveal conserved elements in class I structure. Immunogenetics 36:166–174
Hanson BA, Luttrell MP, Goekjian VH, Niles L, Swayne DE, Senne DA, Stallknecht DE (2008) Is the occurrence of avian influenza virus in Charadriiformes species and location dependent? J Wildlife Dis 44:351–361
Hunt HD, Goto RM, Foster DN, Bacon LD, Miller MM (2006) At least one YMHCI molecule in the chicken is alloimmunogenic and dynamically expressed on spleen cells during development. Immunogenetics 58:297–307
Huson DH, Bryant D (2006) Application of phylogenetic networks in evolutionary studies. Mol Biol Evol 23:254–267
Jarvi S, Goto R, Gee G, Briles W, Miller M (1999) Identification, inheritance, and linkage of B-G-like and MHC class I genes in cranes. J Heredity 90:152–159
Kaufman J (2008) The Avian MHC. In: Davison F, Kaspars B, Schat KA (eds) Avian immunology. Elsevier, London, San Diego, pp 323–338
Kaufman J, Milne S, Göbel TW, Walker BA, Jacob JP, Auffray C, Zoorob R, Beck S (1999) The chicken B locus is a minimal essential major histocompatibility complex. Nature 401:923–925
Kaufman J, Salomonsen J, Flajnik M (1994) Evolutionary conservation of MHC class I and class II molecules—different yet the same. Seminars Immunol 6:411–424
Kelley J, Walter L, Trowsdale J (2005) Comparative genomics of major histocompatibility complexes. Immunogenetics 56:683–695
Klein J (1986) Natural history of the major histocompatibility complex. Wiley, New York
Klein J, Bontrop RE, Dawkins RL, Erlich HA, Gyllensten UB, Heise ER, Jones PP, Parham P, Wakeland EK, Watkins DI (1990) Nomenclature for the major histocompatibility complexes of different species: a proposal. Immunogenetics 31:217–219
Kosakovsky Pond SL, Posada D, Gravenor MB, Woelk CH, Frost SDW (2006) GARD: a genetic algorithm for recombination detection. Bioinformatics 22:3096–3098
Krauss S, Obert CA, Franks J, Walker D, Jones K, Seiler P, Niles L, Pryor SP, Obenauer JC, Naeve CW, Widjaja L, Webby RJ, Webster RG (2007) Influenza in migratory birds and evidence of limited intercontinental virus exchange. PLoS Pathogens 3:e167
Kryazhimskiy S, Plotkin JB (2008) The population genetics of d N /d S . PLoS Genetics 4:e1000304
Møller AP, Erritzøe J (1998) Host immune defence and migration in birds. Evol Ecol 12:945–953
Moon DA, Veniamin SM, Parks-Dely JA, Magor KE (2005) The MHC of the duck (Anas platyrhynchos) contains five differentially expressed class I genes. J Immunol 175:6702–6712
Mortazavi A, Williams BA, McCue K, Schaeffer L, Wold B (2008) Mapping and quantifying mammalian transcriptomes by RNA-Seq. Nature Methods 5:621–628
Murphy K, Travers P, Walport M (2008) Janeway’s immunobiology, 7th edn. Garland Science, New York
Nei M, Gu X, Sitnikova T (1997) Evolution by the birth-and-death process in multigene families of the vertebrate immune system. Proc Natl Acad Sci USA 94:7799–7806
Piersma T (1997) Do global patterns of habitat use and migration strategies co-evolve with relative investments in immunocompetence due to spatial variation in parasite pressure? Oikos 80:623–631
Piersma T (2006) Understanding the numbers and distributions of waders and other animals in a changing world: habitat choice as the lock and the key. Stilt 50:3–14
Piersma T (2007) Using the power of comparison to explain habitat use and migration strategies of shorebirds worldwide. J Ornithol 148(Suppl 1):S45–S59
Piersma T, Baker AJ (2000) Life history characteristics and the conservation of migratory shorebirds. In: Gosling LM, Sutherland WJ (eds) Behaviour and Conservation. Cambridge University Press, Cambridge, UK, pp 105–124
Piersma T, Davidson NC (1992) The migrations and annual cycles of five subspecies of knots in perspective. Wader Study Group Bull 64(Suppl):187–197
Piertney SB, Oliver MK (2006) The evolutionary ecology of the major histocompatibility complex. Heredity 96:7–21
Piontkivska H, Nei M (2003) Birth-and-death evolution in primate MHC class I genes: divergence time estimates. Mol Biol Evol 20:601–609
Posada D (2008) jModelTest: phylogenetic model averaging. Mol Biol Evol 25:1253–1256
Potter CJ, Luo L (2010) Splinkerette PCR for mapping transposable elements in Drosophila. PLoS One 5(10168). doi:10.1371/journal.pone.0010168
Promerová M, Albrecht T, Bryja J (2009) Extremely high MHC class I variation in a population of a long-distance migrant, the scarlet rosefinch (Carpodacus erythrinus). Immunogenetics 61:451–461
Reed KM, Bauer MM, Monson MS, Benoit B, Chaves LD, O’Hare TH, Delany ME (2012) Defining the Turkey MHC: identification of expressed class I- and class IIB-like genes independent of the MHC-B. Immunogenetics 63:753–771
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Sambrook J, Russell DW (2001) Molecular cloning: a laboratory manual. Cold Spring Harbor Laboratory, Cold Spring Harbor, NY
Saper M, Bjorkman P, Wiley D (1991) Refined structure of the human histocompatibility antigen HLA-A2 at 2.6Å resolution. J Mol Biol 219:277–319
Schut E, Aguilar J, Merino S, Magrath M, Komdeur J, Westerdahl H (2011) Characterization of MHC-I in the blue tit (Cyanistes caeruleus) reveals low levels of genetic diversity and trans-population evolution across European populations. Immunogenetics 63:531–542
Sepil I, Moghadam HK, Huchard E, Sheldon BC (2012) Characterization and 454 pyrosequencing of major histocompatibility complex class I genes in the great tit reveal complexity in a passerine system. BMC Evol Biol 12:68
Shiina T, Hosomichi K, Hanzawa K (2006) Comparative genomics of the poultry major histocompatibility complex. Anim Sci Journal 77:151–162
Sommer S (2005) The importance of immune gene variability (MHC) in evolutionary ecology and conservation. Frontiers Zool 2:16. doi:10.1186/1742-9994-2-16
Spurgin LG, Oosterhout CV, Illera JC, Bridgett S, Gharbi K, Emerson BC, Richardson DS (2011) Gene conversion rapidly generates major histocompatibility complex diversity in recently founded bird populations. Mol Ecol 20:5213–5225
Spurgin LG, Richardson DS (2010) How pathogens drive genetic diversity: MHC, mechanisms and misunderstandings. Proc R Soc Lond B 277:979–988
Strand T, Westerdahl H, Höglund J, Alatalo RV, Siitari H (2007) The Mhc class II of the Black grouse (Tetrao tetrix) consists of low numbers of B and Y genes with variable diversity and expression. Immunogenetics 59:725–734
Strandh M, Lannefors M, Bonadonna F, Westerdahl H (2011) Characterization of MHC class I and II genes in a subantarctic seabird, the blue petrel, Halobaena caerulea (Procellariiformes). Immunogenetics 63:653–666
Stroynowski I, Lindahl KF (1994) Antigen presentation by non-classical class I molecules. Current Opin Immunol 6:38–44
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S (2011) MEGA5: Molecular Evolutionary Genetics Analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28:2731–2739
Thomas G, Reynolds JD (2007) Sexual conflict and the evolution of breeding systems in shorebirds. Advances Study Behav 37:279–342
Verkuil YI, Piersma T, Jukema J, Hooljmeijer JCEW, Zwarts L, Baker AJ (2012) The interplay between habitat availability and population differentiation: a case study on genetic and morphological structure in an inland wader (Charadriiformes). Biol J Linnean Soc 106:641–656
Westerdahl H (2007) Passerine MHC: genetic variation and disease resistance in the wild. J Ornithol 148:s469–s477
Westerdahl H, Wittzell H, von Schantz T (1999) Polymorphism and transcription of Mhc class I genes in a passerine bird, the great reed warbler. Immunogenetics 49:158–170
Westerdahl H, Wittzell H, von Schantz T (2000) Mhc diversity in two passerine birds: no evidence for a minimal essential Mhc. Immunogenetics 52:92–100
Wittzell H, Schantz T, Zoorob R, Auffray C (1995) Rfp-Y-like sequences assort independently of pheasant Mhc genes. Immunogenetics 42:68–71
Xia C, Hu T, Yang T, Wang L, Xu G, Lin C (2005) cDNA cloning, genomic structure and expression analysis of the goose (Anser cygnoides) MHC class I gene. Vet Immunol Immunopathol 107:291–302
Yang Z (2007) PAML 4: a program package for phylogenetic analysis by maximum likelihood. Mol Biol Evol 24:1586–1591
Yang Z, Wong WSW, Nielsen R (2005) Bayes empirical Bayes inference of amino acid sites under positive selection. Mol Biol Evol 22:1107–1118
Acknowledgments
We thank Alberto Castillo and Rosemary Gibson for valuable help in the lab; Oliver Haddrath for technical support in the lab and stimulating discussion; Sergio Pereira for help with RNAseq; Sue Chopra, Cathy Dutton, Kirsten Grond, Kevin Kalasz, and Mark Peck for administrative, field, and logistical support; Theunis Piersma, Anne Dekinga, and Anneke Bol for sending samples and extractions; and Alison Cloutier and anonymous reviewers for valuable comments on earlier drafts. This work was supported by funds to DMB from the Natural Science and Engineering Council of Canada (NSERC PDF-373488-2009) and the Netherlands Organisation for Scientific Research (Rubicon 825.09.0190), and to AJB from NSERC (RGPIN 200–07) and the Royal Ontario Museum Life in Crisis: Schad Gallery of Biodiversity Research Fund.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Fig. S1
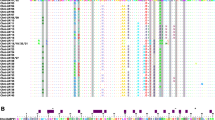
Alignment of exon 1 (signal peptide), exon 4 (α3 domain) and exons 5 to 8 (transmembrane and cytoplasmic domains) in Red Knot and Red-billed Gull MHCI amino acid sequences with chicken and goose as outgroups (see Fig. 2 for exons 2 and 3). In exon 4, CD8 binding sites are shown by yellow annotations and intra- and inter-domain contacts by purple triangles. Sequence sources are Gaga (chicken, G. gallus BF2, AL023516), Anan (domestic goose, A. anser, MHC class I antigen gene, AY387655), Lasc (Red-billed Gull, L. scopulinus, UAA, HM008713, UBA, HM008714, UCA, HM008715, UDA, HM008716). Separate alignments for different exons show only alleles with coverage across the exon in question. The first residue of the signal peptide is alignment position 1 (the start codon M) (DOC 535 kb)
Table S1
A complete list of primers used to partially characterize knot MHCI. Melting temperatures (T m ) were calculated using FastPCR v 5.4.54 (DOC 254 kb)
Rights and permissions
About this article
Cite this article
Buehler, D.M., Verkuil, Y.I., Tavares, E.S. et al. Characterization of MHC class I in a long-distance migrant shorebird suggests multiple transcribed genes and intergenic recombination. Immunogenetics 65, 211–225 (2013). https://doi.org/10.1007/s00251-012-0669-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00251-012-0669-2