Abstract
Although many changes have been discovered during heart maturation, the genetic mechanisms involved in the changes between immature and mature myocardium have only been partially elucidated. Here, gene expression profile changed between the human fetal and adult heart was characterized. A human microarray was applied to define the gene expression signatures of the fetal (13–17 weeks of gestation, n = 4) and adult hearts (30–40 years old, n = 4). Gene ontology analyses, pathway analyses, gene set enrichment analyses, and signal transduction network were performed to predict the function of the differentially expressed genes. Ten mRNAs were confirmed by quantificational real-time polymerase chain reaction. 5547 mRNAs were found to be significantly differentially expressed. “Cell cycle” was the most enriched pathway in the down-regulated genes. EFGR, IGF1R, and ITGB1 play a central role in the regulation of heart development. EGFR, IGF1R, and FGFR2 were the core genes regulating cardiac cell proliferation. The quantificational real-time polymerase chain reaction results were concordant with the microarray data. Our data identified the transcriptional regulation of heart development in the second trimester and the potential regulators that play a prominent role in the regulation of heart development and cardiac cells proliferation.



Similar content being viewed by others
References
Iruretagoyena JI, Davis W, Bird C, Olsen J, Radue R, Teo Broman A, Kendziorski C, Splinter BonDurant S, Golos T, Bird I, Shah D (2014) Metabolic gene profile in early human fetal heart development. Mol Hum Reprod 20:690–700
Laflamme MA, Murry CE (2011) Heart regeneration. Nature 473:326–335
Senyo SE, Steinhauser ML, Pizzimenti CL, Yang VK, Cai L, Wang M, Wu TD, Guerquin-Kern JL, Lechene CP, Lee RT (2013) Mammalian heart renewal by pre-existing cardiomyocytes. Nature 493:433–436
Sturzu AC, Rajarajan K, Passer D, Plonowska K, Riley A, Tan TC, Sharma A, Xu AF, Engels MC, Feistritzer R, Li G, Selig MK, Geissler R, Robertson KD, Scherrer-Crosbie M, Domian IJ, Wu SM (2015) Fetal mammalian heart generates a robust compensatory response to cell loss. Circulation 132:109–121
Kanehisa M, Goto S, Kawashima S, Okuno Y, Hattori M (2004) The KEGG resource for deciphering the genome. Nucleic Acids Res 32:D277–D280
Soufan AT, van den Berg G, Ruijter JM, de Boer PA, van den Hoff MJ, Moorman AF (2006) Regionalized sequence of myocardial cell growth and proliferation characterizes early chamber formation. Circ Res 99:545–552
Ikenishi A, Okayama H, Iwamoto N, Yoshitome S, Tane S, Nakamura K, Obayashi T, Hayashi T, Takeuchi T (2012) Cell cycle regulation in mouse heart during embryonic and postnatal stages. Dev Growth Differ 54:731–738
Tane S, Kubota M, Okayama H, Ikenishi A, Yoshitome S, Iwamoto N, Satoh Y, Kusakabe A, Ogawa S, Kanai A, Molkentin JD, Nakamura K, Ohbayashi T, Takeuchi T (2014) Repression of cyclin D1 expression is necessary for the maintenance of cell cycle exit in adult mammalian cardiomyocytes. J Biol Chem 289:18033–18044
Paradis AN, Gay MS, Zhang L (2014) Binucleation of cardiomyocytes: the transition from a proliferative to a terminally differentiated state. Drug Discov Today 19:602–609
Kong B, Liu YL, Lu XD (2008) Microarray-bioinformatics analysis of altered genomic expression profiles between human fetal and infant myocardium. Chin Med J 121:1257–1264
Lavine KJ, Yu K, White AC, Zhang X, Smith C, Partanen J, Ornitz DM (2005) Endocardial and epicardial derived FGF signals regulate myocardial proliferation and differentiation in vivo. Dev Cell 8:85–95
Gan J, Sonntag HJ, Tang MK, Cai D, Lee KK (2015) Integrative analysis of the developing postnatal mouse heart transcriptome. PloS ONE 10:e0133288
Munshi NV (2012) Gene regulatory networks in cardiac conduction system development. Circ Res 110:1525–1537
Lee KF, Simon H, Chen H, Bates B, Hung MC, Hauser C (1995) Requirement for neuregulin receptor erbB2 in neural and cardiac development. Nature 378:394–398
Gassmann M, Casagranda F, Orioli D, Simon H, Lai C, Klein R, Lemke G (1995) Aberrant neural and cardiac development in mice lacking the ErbB4 neuregulin receptor. Nature 378:390–394
Iwamoto R, Yamazaki S, Asakura M, Takashima S, Hasuwa H, Miyado K, Adachi S, Kitakaze M, Hashimoto K, Raab G, Nanba D, Higashiyama S, Hori M, Klagsbrun M, Mekada E (2003) Heparin-binding EGF-like growth factor and ErbB signaling is essential for heart function. Proc Natl Acad Sci USA 100:3221–3226
Meyer D, Birchmeier C (1995) Multiple essential functions of neuregulin in development. Nature 378:386–390
Li P, Cavallero S, Gu Y, Chen TH, Hughes J, Hassan AB, Bruning JC, Pashmforoush M, Sucov HM (2011) IGF signaling directs ventricular cardiomyocyte proliferation during embryonic heart development. Development 138:1795–1805
Funding
This work was supported by National Natural Science Foundation of China (Grant Number 81670255), Zhejiang Provincial Natural Science Foundation of China (Grant Number LY14H020008), Science and Technology Department of Zhejiang Province (Grant Number 2015C33163), Zhejiang Provincial Natural Science Foundation of China (Grant Number LY13H020005), Scientific Research Foundation of Wenzhou (Grant Number Y20150015), Zhejiang Provincial Medical and Health Science and Technology plan (Grant Number 2016KYB197).
Authors’ Contributions
MPC designed the study; JW and ZMG analyzed the data and wrote the manuscript; LLP, XLC, and JTZ contributed to the collection of samples and analyzed the data; ML revised the paper. All authors read and approved the final version.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare that they have no conflict of interests.
Additional information
Zhimin Geng and Jue Wang have contributed equally to this study.
Electronic supplementary material
Below is the link to the electronic supplementary material.
246_2017_1569_MOESM1_ESM.tif
Extent of cardiomyocyte proliferation in fetal and adult hearts. Immunofluorescent staining shows the proliferating Ki-67+ cardiomyocytes (green arrow heads) in the fetal heart and no proliferating Ki-67+ cardiomyocytes in the adult heart. Scale bar: 50 μm. Supplementary material 1 (TIF 4921 KB)
246_2017_1569_MOESM2_ESM.tif
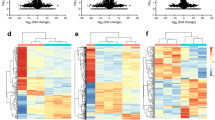
Differentially expressed genes between the fetal and adult heart. The scatterplot (A) is a visualization used to assess the variation between the chips. The X and Y axes in the scatterplot represent the normalized signal values of each group (log2 scaled). The green lines represent FC (FC = 2.0) lines. Hierarchical clustering (B) shows the expression values of differentially expressed genes. ‘Red’ indicates higher expression, whereas ‘green’ indicates lower expression. The bar code represents the color scale of log2 transformed values. Supplementary material 2 (TIF 2771 KB)
246_2017_1569_MOESM3_ESM.tif
GO analysis. All the gene ontology terms that exhibited significant differences between the fetal hearts and adult hearts are listed. A, Gene ontology terms involved in the down-regulated genes in the adult group. B, Gene ontology terms involved in the up-regulated genes. Higher -lg P value indicates higher significance level. Supplementary material 3 (TIF 1449 KB)
246_2017_1569_MOESM4_ESM.tif
Cell cycle pathway. Red frames represent the down-regulated genes and purple frames represent the up-regulated genes in the adult group. Supplementary material 4 (TIF 3946 KB)
246_2017_1569_MOESM5_ESM.tif
GSEA of the cell cycle pathway gene set. GSEA analysis indicated that the cell cycle pathway gene set was significantly enriched (A). The x-axis represents the genes ranked from the most down-regulated to the most up-regulated in the adult group. The y-axis represents enrichment score (ES), which increases when a gene is in the gene set and decreases when it is not. FDR q-value = 0.043, P < 0.001. B represents the genes included in the cell cycle pathway gene sets, with “red” representing the up-regulated genes and blue representing the down-regulated genes. Supplementary material 5 (TIF 3234 KB)
246_2017_1569_MOESM6_ESM.tif
Signal transduction network of the focused genes. Red circles represent down-regulated genes and blue ones represent up-regulated genes in the adult group. The larger the circle is, the higher the degree would be. The interactions between the genes include binding/association (b), phosphorylation (pho), ubiquination (ubi), expression (ex), dephosphorylation (de), activation (a) and indirect (indi, dotted line). Supplementary material 6 (TIF 1149 KB)
Rights and permissions
About this article
Cite this article
Geng, Z., Wang, J., Pan, L. et al. Microarray Analysis of Differential Gene Expression Profile Between Human Fetal and Adult Heart. Pediatr Cardiol 38, 700–706 (2017). https://doi.org/10.1007/s00246-017-1569-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00246-017-1569-x