Abstract
As a result of two-round whole genome duplications, four or more paralogous Hox clusters exist in vertebrate genomes. The paralogous genes in the Hox clusters show similar expression patterns, implying shared regulatory mechanisms for expression of these genes. Previous studies partly revealed the expression mechanisms of Hox genes. However, cis-regulatory elements that control these paralogous gene expression are still poorly understood. Toward solving this problem, the authors searched conserved non-coding sequences (CNSs), which are candidates of cis-regulatory elements. When comparing orthologous Hox clusters of 19 vertebrate species, 208 intergenic conserved regions were found. The authors then searched for CNSs that were conserved not only between orthologous clusters but also among the four paralogous Hox clusters. The authors found three regions that are conserved among all the four clusters and eight regions that are conserved between intergenic regions of two paralogous Hox clusters. In total, 28 CNSs were identified in the paralogous Hox clusters, and nine of them were newly found in this study. One of these novel regions bears a RARE motif. These CNSs are candidates for gene expression regulatory regions among paralogous Hox clusters. The authors also compared vertebrate CNSs with amphioxus CNSs within the Hox cluster, and found that two CNSs in the HoxA and HoxB clusters retain homology with amphioxus CNSs through the two-round whole genome duplications.





Similar content being viewed by others
References
Altschul SF, Madden TL, Schäffer AA, Zhang J, Zhang Z, Miller W, Lipman DJ (1997) Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res 25:3389–3402
Amemiya CT, Prohaska SJ, Hill-Force A, Wasserscheid J, Ferrier DEK, Pascual-Anaya J, Garcia-Fernàndez J, Dewar K, Stadler PF (2008) The amphioxus Hox cluster: characterization, comparative genomics, and evolution. J Exp Zool (Mol Dev Evol) 310B:465–477
Bejerano G, Pheasant M, Makunin I, Stephen S, Kent WJ, Mattick JS, Haussler D (2004) Ultraconserved elements in the human genome. Science 304:1321–1325
Carroll SB (2001) Chance and necessity: the evolution of morphological complexity and diversity. Nature 409:1102–1109
Chiu CH, Amemiya C, Dewar K, Kim CB, Ruddle F, Wagner GP (2002) Molecular evolution of the HoxA cluster in three major gnathostome lineages. Proc Natl Acad Sci USA 99:5492–5497
Dehal P, Boore JL (2005) Two rounds of whole genome duplication in the ancestral vertebrate. PLoS Biol 3:1700–1708
Dinger ME, Amaral PP, Mercer TR, Pang KC, Bruce SJ, Gardiner BB, Askarian-Amiri ME, Ru K, Soldà G, Simons C, Sunkin SM, Crowe ML, Grimmond SM, Perkins AC, Mattick JS (2008) Long noncoding RNAs in mouse embryonic stem cell pluripotency and differentiation. Genome Res 18:1433–1445
Dubrulle J, Pourquié O (2004) Coupling segmentation to axis formation. Development 131:5783–5793
Ferrier DE, Minguillon C, Holland PW, Garcia-Fernàndez J (2000) The amphioxus Hox cluster: deuterostome posterior flexibility and Hox14. Evol Dev 2:284–293
Fitch WM (1971) Towards defining the course of evolution: minimum change for a specific tree topology. Syst Zool 20:406–416
Garcia-Fernàndez J (2005) The genesis and evolution of homeobox gene clusters. Nat Rev Genet 6:881–892
Gibson TJ, Spring J (2000) Evidence in favour of ancient octaploidy in the vertebrate genomes. Biochem Soc Trans 28:259–264
Hancock JM, Shaw PJ, Bonneton F, Dover GA (1999) High sequence turnover in the regulatory regions of the developmental gene hunchback in insects. Mol Biol Evol 284:1083–1094
Holland PW, Garcia-Fernàndez J, Williams NA, Sidow A (1994) Gene duplications and the origins of vertebrate development. Dev Suppl 120(SUPPL.):125–133
Hughes AL, da Silva J, Friedman R (2001) Ancient genome duplications did not structure the human Hox-bearing chromosomes. Genome Res 11:771–780
Juan AH, Ruddle FH (2003) Enhancer timing of Hox gene expression: deletion of the endogenous Hoxc8 early enhancer. Development 130:4823–4834
Kim CB, Amemiya C, Bailey W, Kawasaki K, Mezey J, Miller W, Minoshima S, Shimizu N, Wagner GP, Ruddle F (2000) Hox cluster genomics in the horn shark, Heterodontus francisci. Proc Natl Acad Sci USA 97:1655–1660
Lehoczky JA, Innis JW (2008) BAC transgenic analysis reveals enhancers sufficient for Hoxa13 and neighborhood gene expression in mouse embryonic distal limbs and genital bud. Evol Dev 10:421–423
Lehoczky JA, Williams ME, Innis JW (2004) Conserved expression domains for genes upstream and within the HoxA and HoxD clusters suggests a long-range enhancer existed before cluster duplication. Evol Dev 6:423–430
Lemons D, McGinnis W (2006) Genomic evolution of Hox gene clusters. Science 313:1918–1922
Ludwig MZ, Palsson A, Alekseeva E, Bergman CM, Nathan J, Kreitman M (2005) Functional evolution of a cis-regulatory module. PLoS Biol 3:588–598
Mainguy G, In der Rieden PM, Berezikov E, Woltering JM, Plasterk RH, Durston AJ (2003) A position-dependent organization of retinoid response elements is conserved in the vertebrate Hox clusters. Trends Genet 19:476–479
Mainguy G, Koster J, Woltering J, Jansen H, Durston A (2007) Extensive polycistronism and antisense transcription in the mammalian Hox clusters. PLoS One 2:e356
McEwen GK, Woolfe A, Goode D, Vavouri T, Callaway H, Elgar G (2006) Ancient duplicated conserved noncoding elements in vertebrates: a genomic and functional analysis. Genome Res 16:451–465
Morrison A, Ariza-McNaughton L, Gould A, Featherstone M, Krumlauf R (1997) HOXD4 and regulation of the group 4 paralog genes. Development 124:3135–3146
Murphy WJ, Pevzner PA, O’Brien SJ (2004) Mammalian phylogenomics comes of age. Trends Genet 20:631–639
Ohno S (1970) Evolution by gene duplication. Springer Verlag, New York
Pascual-Anaya J, D’Aniello S, Garcia-Fernàndez J (2008) Unexpected number of conserved noncoding regions within the ancestral chordate Hox cluster. Dev Genes Evol 218:591–597
Pearson JC, Lemons D, McGinnis W (2005) Modulating Hox gene functions during animal body patterning. Nat Rev Genet 6:893–904
Prohaska SJ, Fried C, Flamm C, Wagner GP, Stadler PF (2004) Surveying phylogenetic footprints in large gene clusters: application to Hox cluster duplications. Mol Phylogenet Evol 31:581–604
Putnam NH, Butts T, Ferrier DEK, Furlong RF, Hellsten U, Kawashima T, Robinson-Rechavi M, Shoguchi E, Terry A, Yu JK, Benito-Gutie′rrez E, Dubchak I, Garcia-Fernàndez J, Grigoriev IV, Horton AC, de Jong PJ, Jurka J, Kapitonov V, Kohara Y, Kuroki Y, Lindquist E, Lucas S, Osoegawa K, Pennacchio LA, Salamov AA, Satou Y, Sauka-Spengler T, Schmutz J, Shin-I T, Toyoda A, Gibson-Brown JJ, Bronner-Fraser M, Fujiyama A, Holland LZ, Holland PWH, Satoh N, Rokhsar DS (2008) The amphioxus genome and the evolution of the chordate karyotype. Nature 453:1064–1071
Ray P, Shringarpure S, Kolar M, Xing EP (2008) CSMET: comparative genomic motif detection via multi-resolution phylogenetic shadowing. PLoS Comput Biol 4:e1000090
Santini S, Boore JL, Meyer A (2003) Evolutionary conservation of regulatory elements in vertebrate Hox gene clusters. Genome Res 13:1111–1122
Spitz F, Gonzalez F, Peichel C, Vogt TF, Duboule D, Zákány J (2001) Large scale transgenic and cluster deletion analysis of the HoxD complex separate an ancestral regulatory module from evolutionary innovations. Genes Dev 15:2209–2214
Spitz F, Gonzalez F, Duboule D (2003) A global control region defines a chromosomal regulatory landscape containing the HoxD cluster. Cell 113:405–417
Thomopson JD, Higgins DG, Gibson TJ (1994) CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res 22:4673–4680
Wada H, Escriva H, Zhang S, Laudet V (2006) Conserved RARE localization in amphioxus Hox clusters and implications for Hox code evolution in the vertebrate neural crest. Dev Dyn 235:1522–1531
Werauch MT, Hughes TR (2010) Conserved expression without conserved regulatory sequence: the more things change, the more they stay the same. Trends Genet 26:66–74
Woolfe A, Goodson M, Goode DK, Snell P, McEwen GK, Vavouri T, Smith SF, North P, Callaway H, Kelly K, Walter K, Abnizova I, Gilks W, Edwards YJ, Cooke JE, Elgar G (2005) Highly conserved non-coding sequences are associated with vertebrate development. PLoS Biol 3:116–130
Yang Z (2007) PAML 4: phylogenic analysis by maximum likelihood. Mol Biol Evol 24:1586–1591
Yekta S, Shih IH, Bartel DP (2004) MicroRNA-directed cleavage of HOXB8 mRNA. Science 304:594–596
Acknowledgments
The authors thank Drs. Kiyoshi Ezawa, Hiroki Kokubo, Kazuho Ikeo, Toshihiko Shiroishi, and Hiroyuki Sasaki for their many helpful discussions, suggestions, and comments. The authors appreciate English polishing by Ms Rumiko Suzuki, Ms Mahoko Takahashi, and Mr. Tim Jinam. This study was supported partly by The Graduate University for Advanced Studies (Sokendai) to M.M. and by Grant-in-Aid for scientific research from the Ministry of Education, Culture, Sports, Science, and Technology of Japan to N.S.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
239_2010_9396_MOESM1_ESM.xls
Supplementary Table 1: Feature of each CNS. Table shows location, conservation, length and feature of each CNS. Especially HoxA cluster, we compared our CNSs with footprinting results of Prohaska et al. (2004) and available transcript data. (XLS 68 kb)
239_2010_9396_MOESM2_ESM.xls
Supplementary Table 2: Conservation level of each CNS. Table shows existence or nonexistence of each CNS among 19 species. The CNSs which conserve among only placental mammalians are highlighted by purple. (XLS 100 kb)
239_2010_9396_MOESM3_ESM.ppt
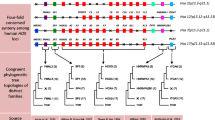
Supplementary Figure 1: The guide tree The phylogenetic tree shows vertebrate phylogeny drawn upon Murphy et al. (2004). The scale bar is one million years ago (MYA). All species in this figure have four Hox clusters. On the right of the tree, higher taxonomic groupings are shown in colors. The same coloring is used in Figure 4 and Supplementary Figure 2. (PPT 169 kb)
239_2010_9396_MOESM4_ESM.ppt
Supplementary Figure 2: The phylogenetic footprinting analysis of the TP and DP CNSs The results of phylogenetic footprinting analysis of each TP (A) - (C) and each DP (D) – (K) are shown. Graphs are the results of sliding window analysis on each multiple alignment. X-axis represents base positions. Y-axis represents substitution number of each window. In the graph, substitution numbers within placental mammalians, amniotes and vertebrates are colored in blue, red and green, respectively. Orange boxes show paralogous conserved sequences. It is noted that paralogous conserved regions are also highly conserved at the orthologous comparison. (PPT 530 kb)
239_2010_9396_MOESM5_ESM.ppt
Supplementary Figure 3: Multiple alignments of each DP CNS. (A) - (H) are results of multiple alignments of paralogous conserved regions derived from each DP CNS. Construction of each alignment is same as Figure 2. (PPT 678 kb)
239_2010_9396_MOESM6_ESM.ppt
Supplementary Figure 4: Multiple alignments of each amphiCNS. Figures show multiple alignments of CNSs which are (A) conserved among all vertebrates and (B) not conserved among all vertebrates from comparison between vertebrate CNSs and amphiCNSs. Construction of each alignment is same as Figure 2. (PPT 1064 kb)
Rights and permissions
About this article
Cite this article
Matsunami, M., Sumiyama, K. & Saitou, N. Evolution of Conserved Non-Coding Sequences Within the Vertebrate Hox Clusters Through the Two-Round Whole Genome Duplications Revealed by Phylogenetic Footprinting Analysis. J Mol Evol 71, 427–436 (2010). https://doi.org/10.1007/s00239-010-9396-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00239-010-9396-1