Abstract
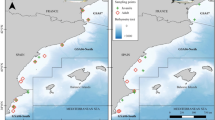
The Osumi Strait and its neighboring waters are major spawning grounds for commercially important small pelagic and coastal fish in the Pacific coastal area of Japan. Mesopelagic fish larvae are also abundant here, co-occurring with the larvae of commercial species. However, it is unknown whether these fish groups compete for prey. We examined the diet of four commercial fish groups (Engraulis japonicus, Trachurus japonicus, Triglidae spp., and Sebastiscus spp.) and two mesopelagic species (Sigmops gracilis, Myctophum asperum) during their larval stages to determine whether they had the same prey source. Morphological analysis and DNA metabarcoding of gut contents showed that the main prey species of all six fish groups were calanoid copepods. However, results from metabarcoding differed greatly from morphological analysis, showing that appendicularians were abundant in the guts of S. gracilis, M. asperum, and Sebastiscus spp. This may be because the methods differ in how they determine prey composition, with morphological analysis relying on counts of identifiable prey parts and metabarcoding being only a semi-quantitative method. In addition, appendicularians might be underestimated by metabarcoding analysis due to variation in genome size, copy number of target genes, and primer mismatches. Stable isotope analysis supported the importance of appendicularians as prey for S. gracilis, M. asperum, and Sebastiscus spp. Our results indicate that the two mesopelagic species and Sebastiscus spp. may compete for prey, but E. japonicus, T. japonicus, and Triglidae spp. occupy niches different from those of the two mesopelagic species. Our results imply that the trophic pathway via appendicularians may support the feeding of dominant mesopelagic species and enable coexistence with commercial species in the study area.





Similar content being viewed by others
References
Albaina A, Aguirre M, Abad D, Santos M, Estonba A (2016) 18S rRNA V9 metabarcoding for diet characterization: a critical evaluation with two sympatric zooplanktivorous fish species. Ecol Evol 6:1809–1824. https://doi.org/10.1002/ece3.1986
Amaral-Zettler LA, McCliment EA, Ducklow HW, Huse SM (2009) A method for studying protistan diversity using massively parallel sequencing of V9 hypervariable regions of small-subunit ribosomal RNA genes. PLoS ONE 4:e6372. https://doi.org/10.1371/journal.pone.0006372
Aritomi A, Andou E, Yonezawa T, Kume G (2017) Monthly occurrence and feeding habits of larval and juvenile Ryukyu-ayu Plecoglossus altivelis ryukyuensis in an estuarine lake and coastal area of the Kawauchi River, Amami-oshima Island, southern Japan. Ichthyol Res 64:159–168. https://doi.org/10.1007/s10228-016-0550-7
Bolger AM, Lohse M, Usadel B (2014) Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30:2114–2120. https://doi.org/10.1093/bioinformatics/btu170
Bucklin A, DiVito KR, Smolina IV, Choquet M, Questel JM, Hoarau G, O’Neill RJ (2018) Population genomics of marine zooplankton. In: Oleksiak M, Rajora O (eds) Population genomics: marine organisms, population genomics, pp 61−102. doi: https://doi.org/10.1007/13836_2017_9
Chesson J (1978) Measuring preference in selective predation. Ecology 59:211–215. https://doi.org/10.2307/1936364
Contreras T, Olivar MP, Bernal A, Sabatés A (2015) Comparative feeding patterns of early stages of mesopelagic fishes with vertical habitat partitioning. Mar Biol 162:2265–2277. https://doi.org/10.1007/s00227-015-2749-y
de Vargas C, Audic S, Henry N, Decelle J et al (2015) Eukaryotic plankton diversity in the sunlit ocean. Science 348:1261605. https://doi.org/10.1126/science.1261605
Deibel D (1998) Feeding and metabolism of Appendicularia. In: Bone Q (ed) The biology of pelagic tunicates. Oxford University Press, Oxford, pp 139–149
DeNiro MJ, Epstein S (1977) Mechanism of carbon isotope fractionation associated with lipid synthesis. Science 197:261–263. https://doi.org/10.1126/science.327543
Devloo-Delva F, Huerlimann R, Chua G, Matley JK, Heupel MR, Simpfendorfer CA, Maes GE (2018) How does marker choice affect your diet analysis: comparing genetic markers and digestion levels for diet metabarcoding of tropical-reef piscivores. Mar Freshw Res 70:8–18. https://doi.org/10.1071/MF17209
Edgar RC, Haas BJ, Clemente JC, Quince C, Knight R (2011) UCHIME improves sensitivity and speed of chimera detection. Bioinformatics 27:2194–2200. https://doi.org/10.1093/bioinformatics/btr381
Flood PR, Deibel D (1998) The appendicularian house. In: Bone Q (ed) The biology of pelagic tunicates. Oxford University Press, Oxford, pp 105–124
Fry B (2008) Stable isotope ecology, 3rd edn. Springer, New York. doi: https://doi.org/10.1007/0-387-33745-8
Gorsky G, Fenaux R (1998) The role of Appendicularia in marine food webs. In: Bone Q (ed) The biology of pelagic tunicates. Oxford University Press, Oxford, pp 161–169
Hamada R, Kinoshita I (1988) Feeding habit of larval and juvenile ayu, Plecoglossus altivelis in the surf zone of Tosa Bay, Japan. Jpn J Ichthyol 35:382–388. https://doi.org/10.11369/jji1950.35.382
Hattori S (1964) Studies on fish larvae in the Kuroshio and adjacent waters. Bull Tokai Reg Fish Res Lab 40:1–158
Hirai J, Hidaka K, Nagai S, Ichikawa T (2017) Molecular-based diet analysis of the early post-larvae of Japanese sardine Sardinops melanostictus and Pacific round herring Etrumeus teres. Mar Ecol Prog Ser 564:99–113. https://doi.org/10.3354/meps12008
Hirai J, Kuriyama M, Ichikawa T, Hidaka K, Tsuda A (2015) A metagenetic approach for revealing community structure of marine planktonic copepods. Mol Ecol Resour 15:68–80. https://doi.org/10.1111/1755-0998.12294
Hirota Y, Honda H, Sakaji H, Uehara S (2009) Ontogenetic and diel variation in the vertical distribution of larvae of jack mackerel Trachurus japonicus in the East China sea. Fish Sci 75:577–584. https://doi.org/10.1007/s12562-009-0075-3
Hirota Y, Honda H, Sakaji H, Uehara S, Ichikawa T (2016) Ontogenetic changes of feeding habits of larval jack mackerel Trachurus japonicus in the Kuroshio frontal area of the southern East China Sea. Fish Sci 82:573–583. https://doi.org/10.1007/s12562-016-0989-5
Hjort J (1914) Fluctuations in the great fisheries of northern Europe viewed in the light of biological research. Int Council Explor Sea 20:1–228
Huse SM, Welch DM, Morrison HG, Sogin ML (2010) Ironing out the wrinkles in the rare biosphere through improved OTU clustering. Environ Microbial 12:1889–1898. https://doi.org/10.1111/1462-2920.2010.02193.x
Kobari T, Makihara W, Kawafuchi T, Sato K, Kume G (2018) Geographic variability in taxonomic composition, standing stock, and productivity of the mesozooplankton community around the Kuroshio Current in the East China Sea. Fish Oceanogr 27:336–350. https://doi.org/10.1111/fog.12256
Kobari T, Kobari Y, Miyamoto H, Okazaki Y, Kume G, Kondo R, Habano A (2019) Variability in taxonomic composition, standing stock and productivity of the plankton community in the Kuroshio and its neighboring waters. In: Nagai T, Saito H, Suzuki K, Takahashi, M (eds) Kuroshio current: physical, biogeochemical and ecosystem dynamics, geophysical monograph, vol 243, pp 223−243. doi: https://doi.org/10.1002/9781119428428.ch14
Kodama T, Hirai J, Tamura S, Takahashi T, Tanaka Y, Ishihara T, Tawa A, Morimoto H, Ohshimo S (2017) Diet composition, and feeding habits of larval Pacific bluefin tuna Thunnus orientalis in the Sea of Japan: integrated morphological and metagenetic analysis. Mar Ecol Prog Ser 583:211–226. https://doi.org/10.3354/meps12341
Kume G, Furumitsu K, Nakata H, Suzuki T, Handa M, Yamaguchi A (2015a) Spatiotemporal occurrence and feeding habits of tonguefish, Cynoglossus lighti Norman, 1925, larvae in Ariake Bay, Japan. J Appl Ichthyol 31:276–281. https://doi.org/10.1111/jai.12498
Kume G, Yagishita N, Furumitsu K, Nakata H, Suzuki T, Handa M, Yamaguchi A (2015b) The role of molecular methods to compare distribution and feeding habits in larvae and juveniles of two co-occurring sciaenid species Nibea albiflora and Pennahia argentata. Estuar Coast Shelf Sci 167:516–525. https://doi.org/10.1016/j.ecss.2015.10.034
Kuwahara A, Suzuki S (1982) Vertical distribution and feeding habits of a horse mackerel Trachurus japonicus and barracuda Sphyraena pinguis larvae in the western Wakasa Bay. Bull Kyoto Ocean Fish Sci 6:13–17
Le DQ, Tanaka K, Hii YS, Sano Y, Nanjo K, Shirai K (2018) Importance of seagrass-mangrove continuum as feeding grounds for juvenile pink ear emperor Lethrinus lentjan in Setiu Lagoon, Malaysia: Stable isotope approach. J Sea Res 135:1–10. https://doi.org/10.1016/j.seares.2018.02.004
Llopiz JK (2013) Latitudinal and taxonomic patterns in the feeding ecologies of fish larvae: a literature synthesis. J Mar Syst 109–110:69–77. https://doi.org/10.1016/j.jmarsys.2012.05.002
Maruyama A, Nakamura K, Yamanaka H, Kondoh M, Minamoto T (2014) The release rate of environmental DNA from juvenile and adult fish. PLoS ONE 9:e114639. https://doi.org/10.1371/journal.pone.0114639
Mei W, Umezawa U, Wan X, Sassa C, Takahashi M (2019) Factors controlling spatiotemporal variations in stable nitrogen isotopes of Trachurus japonicus larvae and juveniles in the East China Sea. Fish Sci 85:71–80. https://doi.org/10.1007/s12562-018-1267-5
Mitsuzawa A, Miyamoto H, Ueda H (2017) Feeding selectivity of early-stage fish larvae on the nauplii and eggs of different copepod species. Plankton Benthos Res 12:115–122. https://doi.org/10.3800/pbr.12.115
Monson KD, Hayes JM (1982) Carbon isotopic fractionation in the biosynthesis of bacterial fatty acids. Ozonolysis of unsaturated fatty acids as a means of determining the intramolecular distribution of carbon isotopes. Geochim Cosmochim Acta 46:139–149. https://doi.org/10.1016/0016-7037(82)90241-1
Montagnes DJS, Dower JF, Figueiredo GM (2010) The protozooplankton−ichthyoplankton trophic link: an overlooked aspect of aquatic food webs. J Eukaryot Microbiol 57:223–228. https://doi.org/10.1111/j.1550-7408.2010.00476.x
Morote E, Olivar MP, Villate F, Uriarte I (2010) A comparison of anchovy (Engraulis encrasicolus) and sardine (Sardina pilchardus) larvae feeding in the Northwest Mediterranean: influence of prey availability and ontogeny. ICES J Mar Sci 67:897–908. https://doi.org/10.1093/icesjms/fsp302
Nagai S, Yamamoto K, Hata N, Itakura S (2012) Study of DNA extraction methods for use in loop-mediated isothermal amplification detection of single resting cysts in the toxic dinoflagellates Alexandrium tamarense and A. catenella. Mar Genomics 7:51–56. https://doi.org/10.1016/j.margen.2012.03.002
Nakamura H, Ichikawa H, Nishina A (2003) Kuroshio path meander between the continental slope and the Tokara Strait in the East China Sea. J Geophys Res 108:3360. https://doi.org/10.1029/2002JC001450
Nakamura H, Hiranaka R, Ambe D, Saito T (2015) Local wind effect on the Kuroshio path state off the southeastern coast of Kyushu. J Oceanogr 71:575–596. https://doi.org/10.1007/s10872-015-0309-1
Newman SP, Handy RD, Gruber SH (2010) Diet and prey preference of juvenile lemon sharks Negaprion brevirostris. Mar Ecol Prog Ser 398:221–234. https://doi.org/10.3354/meps08334
Okazaki Y, Miyamoto H, Suzuki K, Saito H, Hidaka K, Ichikawa T (2019) Diverse trophic pathways from zooplankton to larval and juvenile fishes in the Kuroshio ecosystem. In: Nagai T, Saito H, Suzuki K, Takahashi M (eds) Kuroshio current: physical, biogeochemical and ecosystem dynamics, geophysical monograph, vol 243, pp 245−256. doi: https://doi.org/10.1002/9781119428428.ch15
Østergaard P, Munk P, Janekarn V (2005) Contrasting feeding patterns among species of fish larvae from the tropical Andaman Sea. Mar Biol 146:595–606. https://doi.org/10.1007/s00227-004-1458-8
Pompanon F, Deagle BE, Symondson WOC, Brown DS, Jarman SN, Taberlet P (2012) Who is eating what: diet assessment using next generation sequencing. Mol Ecol 21:1931–1950. https://doi.org/10.1111/j.1365-294X.2011.05403.x
Post DM (2002) Using stable isotopes to estimate trophic position: models, methods, and assumptions. Ecology 83:703–718. https://doi.org/10.1890/0012-9658(2002)083[0703:USITET]2.0.CO;2
Prokopowich CD, Gregory TR, Crease TJ (2003) The correlation between rDNA copy number and genome size in eukaryotes. Genome 46:48–50. https://doi.org/10.1139/G02-103
Quast C, Pruesse E, Yilmaz P, Gerken J, Schweer T, Yarza P, Peplies J, Glöckner FO (2013) The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucleic Acids Res 41:D590–D596. https://doi.org/10.1093/%2Fnar%2Fgks1219
Riemann L, Alfredsson H, Hansen MM, Als TD et al (2010) Qualitative assessment of the diet of European eel larvae in the Sargasso Sea resolved by DNA barcoding. Biol Lett 6:819–822. https://doi.org/10.1098/rsbl.2010.0411
Robert D, Murphy HM, Jenkins GP, Fortier L (2014) Poor taxonomical knowledge of larval fish prey preference is impeding our ability to assess the existence of a “critical period” driving year-class strength. ICES J Mar Sci 71:2042–2052. https://doi.org/10.1093/icesjms/fst198
Sakaguchi SO, Shimamura S, Shimizu Y, Ogawa G, Yamada Y, Shimizu K, Kasai H, Kitazato H, Fujiwara Y, Fujikura K, Takishita K (2017) Comparison of morphological and DNA-based techniques for stomach content analyses in juvenile chum salmon Oncorhynchus keta: a case study on diet richness of juvenile fishes. Fish Sci 83:47–56. https://doi.org/10.1007/s12562-016-1040-6
Sassa C, Kawaguchi K (2005) Larval feeding habits of Diaphus theta, Protomyctophum thompsoni, and Tarletonbeania taylori (Pisces: Myctophidae) in the transition region of the western North Pacific. Mar Ecol Prog Ser 298:261–276. https://doi.org/10.3354/meps298261
Sassa C, Takahashi M (2018) Comparative larval growth and mortality of mesopelagic fishes and their predatory impact on zooplankton in the Kuroshio region. Deep Sea Res 131:121–132. https://doi.org/10.1016/j.dsr.2017.11.007
Sassa C, Tsukamoto Y (2012) Inter-annual comparison of diet and daily ration of larval jack mackerel Trachurus japonicus in the southern East China Sea. J Plankton Res 34:173–187. https://doi.org/10.1093/plankt/fbr089
Sassa C, Moser HG, Kawaguchi K (2002) Horizontal and vertical distribution patterns of larval myctophid fishes in the Kuroshio Current region. Fish Oceanogr 11:1–10. https://doi.org/10.1046/j.1365-2419.2002.00182.x
Sassa C, Kawaguchi K, Mori K (2004) Late winter larval mesopelagic fish assemblage in the Kuroshio waters of the western North Pacific. Fish Oceanogr 13:121–133. https://doi.org/10.1046/j.1365-2419.2003.00275.x
Sassa C, Kawaguchi K, Hirota Y, Ishida M (2007) Distribution depth of the transforming stage larvae of myctophid fishes in the subtropical–tropical waters of the western North Pacific. Deep Sea Res 54:2181–2193. https://doi.org/10.1016/j.dsr.2007.09.006
Sato R, Suzuki Y (2010) Carbon and nitrogen stable isotope analysis by EA/IRMS. Res Org Geochem 26:21–29. https://doi.org/10.20612/rog.26.0_21
Schloss PD, Westcott SL, Ryabin T, Hall JP et al (2009) Introducing mothur: open-source, platform-independent, community-supported software for describing and comparing microbial communities. Appl Environ Microbiol 75:7537–7541. https://doi.org/10.1128/AEM.01541-09
Sugisaki H, Kodama M, Ichikawa T, Yamada K, Wada E, Watanabe T (2013) A new aspect on food web analyses using a nitrogen and carbon isotope map—guidance for interpretations of values and estimation model for stable isotope values of primary producers of food web. J Fish Technol 6:57–68
Uotani I (1985) The relation between feeding mode and feeding habit of the anchovy larvae. Bull Jpn Soc Sci Fish 51:1057–1065. https://doi.org/10.2331/suisan.51.1057
Vaslet A, Phillips DL, France CAM, Feller IC, Baldwin CC (2015) Trophic behaviour of juvenile reef fishes inhabiting interlinked mangrove–seagrass habitats in offshore mangrove islets. J Fish Biol 87:256–273. https://doi.org/10.1111/jfb.12715
Wang Q, Garrity GM, Tiedje JM, Cole JR (2007) Naïve Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl Environ Microbiol 73:5261–5267. https://doi.org/10.1128/AEM.00062-07
Watanabe H, Sassa C, Ishida M (2010) Late winter vertical distribution of mesopelagic fish larvae in the Kuroshio Current region of the western North Pacific. Bull Jpn Soc Fish Oceanogr 74:153–158
Yamashiro T, Shigehara T, Asano T (2008) Influence of the Osumi Branch Current on oceanic water inflow into Kagoshima Bay. Proc Civil Engineer Ocean 24:921–926. https://doi.org/10.2208/prooe.24.921
Yasue N, Doiuchi R, Yoshimoto Y, Takeuchi T (2010) Diet of late larval Japanese anchovy Engraulis japonicus in the Kii Channel, Japan. Fish Sci 76:63–73. https://doi.org/10.1007/s12562-009-0181-2
Acknowledgements
We thank the captain and crew of the research vessel Nansei-maru for their help in oceanographic observations and fish and zooplankton samplings. We also thank the anonymous reviewers for their constructive feedback on our manuscript and Dr. Clive Trueman for editorial comments.
Funding
This work was supported by 16H02947, 17K00522, 18H04920, 20K06186, 20K12151 (JSPS), and the Study of Kuroshio Ecosystem Dynamics for Sustainable Fisheries: JPMXD0511102330 (MEXT).
Author information
Authors and Affiliations
Contributions
KG, KT, KH, TT, IM, and KT led field surveys. KG and KH led all gut contents analyses for fish larvae. KT and HJ led DNA metabarcoding analysis for larval gut contents. A-NM and HF led stable isotope analysis for fish larvae and prey components. KG and HJ conducted the literature research.
Corresponding author
Ethics declarations
Conflicts of interest
The authors declare that they have no conflicts of interest.
Ethics approval
This work followed all applicable international, national, and institutional guidelines for the care and use of animals.
Availability of data and material
Raw sequencing data are available from NCBI/EBI/DDBJ Sequence Read Archive (BioProject accession number PRJDB9655).
Additional information
Responsible Editor: C. Trueman.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Reviewed by C. Harrod and M. R. Jensen
Supplementary information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Kume, G., Kobari, T., Hirai, J. et al. Diet niche segregation of co-occurring larval stages of mesopelagic and commercially important fishes in the Osumi Strait assessed through morphological, DNA metabarcoding, and stable isotope analyses. Mar Biol 168, 6 (2021). https://doi.org/10.1007/s00227-020-03810-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00227-020-03810-x