Abstract
Key message
The white immature fruit color gene w was rapidly mapped to a 33.0-kb region to identify a valuable candidate gene that encodes peroxidase.
Abstract
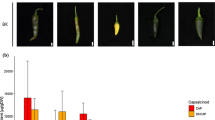
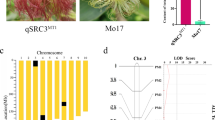
The skin color of immature fruit is a crucial external trait of cucumbers, and white skin is shared by limited numbers of commercial cultivars. Herein, one BC1 population and two F2 segregating populations were constructed using four inbred parental lines (WD3 × B-2-2 and Q30 × Q24) to investigate the inheritance patterns and chromosomal locations of immature fruit color genes in cucumbers. Consequently, a single recessive gene, w, was identified that controls white immature fruit color. A total of 526 markers, which were derived from published genetic maps, two reference cucumber genomes (“9930” and GY14), and two parents (Q30 and Q24) for which whole-genome sequence information is available, were used to map the target gene w to a 33.0-kb region flanked by two SNP-based markers, ASPCR39262 and ASPCR39229, which are physically located at 39262450 and 39229482 of chromosome 3 (“9930” draft genome assembly), respectively. Gene prediction indicated that four potential genes were located in the target region. One gene that encodes peroxidase is likely to be a valuable candidate gene because quantitative real-time PCR revealed an eightfold difference in its transcriptional level, and several amino acid variations were found when the deduced amino acid sequence was aligned. A co-segregating marker was used synergistically to test its ability to predict the skin colors of 83 dark green/white germplasms, and the validity of its utility in marker-assisted selection was confirmed. Fine mapping of this locus will assist in cloning the gene and in marker-assisted breeding to develop dark green/white cucumber cultivars.



Similar content being viewed by others
References
Abeles FB, Dunn LJ (1989) Role of peroxidase during ethylene-induced chlorophyll breakdown in Cucumis sativus cotyledons. J Plant Growth Regul 8:319–325
Beale SI (2005) Green genes gleaned. Trends Plant Sci 10:309–312
Bisognin DA (2002) Origin and evolution of cultivated cucurbits. Ciência Rural 32:715–723
Bollivar DW, Suzuki JY, Beatty JT, Dobrowolski JM, Bauer CE (1994) Directed mutational analysis of bacteriochlorophyll a biosynthesis in Rhodobacter capsulatus. J Mol Biol 237:622–640
Cavagnaro PF, Senalik DA, Yang LM et al (2010) Genome-wide characterization of simple sequence repeats in cucumber (Cucumis sativus L.). BMC Genom 11:569. doi:10.1186/1471-2164-11-569
Chang CC-C, Ball L, Fryer MJ, Baker NR, Karpinski S, Mullineaux PM (2004) Induction of ASCORBATE PEROXIDASE 2 expression in wounded Arabidopsis leaves does not involve known wound-signalling pathways but is associated with changes in photosynthesis. Plant J 38:499–511. doi:10.1111/j.1365-313X.2004.02066.x
Clark MS (ed) (1997) Plant molecular biology—a laboratory manual. Springer, Berlin. doi:10.1007/978-3-642-87873-2
Dong S, Miao H, Zhang S, Liu M, Wang Y, Gu X (2012) Genetic analysis and gene mapping of white fruit skin in cucumber (Cucumis sativus L.). Acta Bot Boreal Occident Sin 32:2177–2181
Drenkard E, Richter BG, Rozen S et al (2000) A simple procedure for the analysis of single nucleotide polymorphisms facilitates map-based cloning in Arabidopsis. Plant Physiol 124:1483–1492
Faris J, Laddomada B, Gill B (1998) Molecular mapping of segregation distortion loci in Aegilops tauschii. Genetics 149:319–327
Feller A, Machemer K, Braun EL, Grotewold E (2011) Evolutionary and comparative analysis of MYB and bHLH plant transcription factors. Plant J 66:94–116
Funamoto Y, Yamauchi N, Shigyo M (2003) Involvement of peroxidase in chlorophyll degradation in stored broccoli (Brassica oleracea L.) and inhibition of the activity by heat treatment. Postharvest Biol Technol 28:39–46
Howden R, Park SK, Moore JM, Orme J, Grossniklaus U, Twell D (1998) Selection of T-DNA-tagged male and female gametophytic mutants by segregation distortion in Arabidopsis. Genetics 149:621–631
Huang S, Li R, Zhang Z et al (2009) The genome of the cucumber, Cucumis sativus L. Nat Genet 41:1275–1281. doi:10.1038/ng.475
Hwang J, Oh J, Kim Z, Staub JE, Chung S-M, Park Y (2014) Fine genetic mapping of a locus controlling short internode length in melon (Cucumis melo L.). Mol Breeding 34:949–961. doi:10.1007/s11032-014-0088-1
Jacob-Wilk D, Holland D, Goldschmidt EE, Riov J, Eyal Y (1999) Chlorophyll breakdown by chlorophyllase: isolation and functional expression of the Chlase1 gene from ethylene-treated Citrus fruit and its regulation during development. Plant J 20:653–661. doi:10.1046/j.1365-313X.1999.00637.x
Kooistra E (1971) Inheritance of fruit flesh and skin colours in powdery mildew resistant cucumbers (Cucumis sativus L.). Euphytica 20:521–523. doi:10.1007/bf00034206
Ky C-L, Barre P, Lorieux M et al (2000) Interspecific genetic linkage map, segregation distortion and genetic conversion in coffee (Coffea sp.). Theor Appl Genet 101:669–676
Li Y (2008) SRAP markers linked to the green skin trait of cucumber. Dissertation. Northwest A&F University, Yangling
Li Y, Wen C, Weng Y (2013) Fine mapping of the pleiotropic locus B for black spine and orange mature fruit color in cucumber identifies a 50 kb region containing a R2R3-MYB transcription factor. Theor Appl Genet 126:2187–2196. doi:10.1007/s00122-013-2128-3
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2− ΔΔCT method. Methods 25:402–408
Lorieux M, Goffinet B, Perrier X, de León DG, Lanaud C (1995a) Maximum-likelihood models for mapping genetic markers showing segregation distortion. 1. Backcross populations. Theor Appl Genet 90:73–80. doi:10.1007/bf00220998
Lorieux M, Perrier X, Goffinet B, Lanaud C, de León DG (1995b) Maximum-likelihood models for mapping genetic markers showing segregation distortion. 2. F2 populations. Theor Appl Genet 90:81–89. doi:10.1007/bf00220999
Lu H, Romero-Severson J, Bernardo R (2002) Chromosomal regions associated with segregation distortion in maize. Theor Appl Genet 105:622–628. doi:10.1007/s00122-002-0970-9
McCouch S, Kochert G, Yu Z, Wang Z, Khush G, Coffman W, Tanksley S (1988) Molecular mapping of rice chromosomes. Theor Appl Genet 76:815–829. doi:10.1007/BF00273666
Meglic V, Staub JE (1996) Inheritance and linkage relationships of isozyme and morphological loci in cucumber (Cucumis sativus L.). Theor Appl Genet 92:865–872. doi:10.1007/bf00221899
Miao H, Zhang S, Wang X et al (2011) A linkage map of cultivated cucumber (Cucumis sativus L.) with 248 microsatellite marker loci and seven genes for horticulturally important traits. Euphytica 182:167–176. doi:10.1007/s10681-011-0410-5
Michaels SD, Amasino RM (1998) A robust method for detecting single-nucleotide changes as polymorphic markers by PCR. Plant J 14:381–385. doi:10.1046/j.1365-313X.1998.00123.x
Motamayor JC, Mockaitis K, Schmutz J et al (2013) The genome sequence of the most widely cultivated cacao type and its use to identify candidate genes regulating pod color. Genome Biol 14:r53. doi:10.1186/gb-2013-14-6-r53
Neff MM, Neff JD, Chory J, Pepper AE (1998) dCAPS, a simple technique for the genetic analysis of single nucleotide polymorphisms: experimental applications in Arabidopsis thaliana genetics. Plant J 14:387–392
Nie J, He H, Peng J et al (2015) Identification and fine mapping of pm5. 1: a recessive gene for powdery mildew resistance in cucumber (Cucumis sativus L.). Mol Breeding 35:1–13
Paterson AH, Lander ES, Hewitt JD, Peterson S, Lincoln SE, Tanksley SD (1988) Resolution of quantitative traits into mendelian factors by using a complete linkage map of restriction fragment length polymorphisms. Nature 335:721–726
Phadnis N, Orr HA (2009) A single gene causes both male sterility and segregation distortion in Drosophila hybrids. Science 323:376–379
Pierce LK, Wehner TC (1990) Review of genes and linkage groups in cucumber. HortScience 25:605–615
Ren Y, Zhang ZH, Liu JH et al (2009) Integrated genetic and cytogenetic map of the cucumber genome. PLoS One 4:e5795. doi:10.1371/journal.pone.0005795
Salamov AA, Solovyev VV (2000) Ab initio gene finding in Drosophila genomic DNA. Genome Res 10:516–522
Sebastian P, Schaefer H, Telford IR, Renner SS (2010) Cucumber (Cucumis sativus) and melon (C. melo) have numerous wild relatives in Asia and Australia, and the sister species of melon is from Australia. Proc Natl Acad Sci 107:14269–14273
Siegel BZ (1993) Plant peroxidases—an organismic perspective. Plant Growth Regul 12:303–312. doi:10.1007/bf00027212
Singh R, Low E-TL, Ooi LC-L et al (2014) The oil palm VIRESCENS gene controls fruit colour and encodes a R2R3-MYB. Nat Commun 5:4106. doi:10.1038/ncomms5106
Tripathi BN, Bhatt I, Dietz K-J (2009) Peroxiredoxins: a less studied component of hydrogen peroxide detoxification in photosynthetic organisms. Protoplasma 235:3–15
Wang J, Fang X, Li X, Chen Y, Fang Z, Xu Y (2013) Genetic study on immature fruit color of cucumber. Acta Hortic Sin 40:479–486
Weng Y (2014) Molecularly tagged genes and quantitative trait loci in cucumber. Paper presented at the CUCURBITACEAE 2014, Bay Harbor, October 12–16, 2014
Weng Y, Li W, Devkota RN, Rudd JC (2005) Microsatellite markers associated with two Aegilops tauschii-derived greenbug resistance loci in wheat. Theor Appl Genet 110:462–469. doi:10.1007/s00122-004-1853-z
Weng Y, Johnson S, Staub JE, Huang S (2010) An extended intervarietal microsatellite linkage map of cucumber, Cucumis sativus L. HortScience 45:882–886
Xie J, Wehner TC (2001) Gene list 2001 for cucumber. Cucurbit Genet Coop Rep 24:110–136
Yamauchi N, Funamoto Y, Shigyo M (2004) Peroxidase-mediated chlorophyll degradation in horticultural crops. Phytochem Rev 3:221–228. doi:10.1023/b:phyt.0000047796.98784.06
Yang L, Koo D-H, Li Y et al (2012) Chromosome rearrangements during domestication of cucumber as revealed by high-density genetic mapping and draft genome assembly. Plant J 71:895–906. doi:10.1111/j.1365-313X.2012.05017.x
Yang L, Li D, Li Y, Gu X, Huang S, Garcia-Mas J, Weng Y (2013) A 1,681-locus consensus genetic map of cultivated cucumber including 67 NB-LRR resistance gene homolog and ten gene loci. BMC Plant Biol 13:53
Yang X, Li Y, Zhang W, He H, Pan J, Cai R (2014a) Fine mapping of the uniform immature fruit color gene u in cucumber (Cucumis sativus L.). Euphytica 196:341–348
Yang X, Zhang W, Li Y et al (2014b) High-resolution mapping of the dull fruit skin gene D in cucumber (Cucumis sativus L.). Mol Breed 33:15–22
Yuan X, Pan J, Cai R et al (2008) Genetic mapping and QTL analysis of fruit and flower related traits in cucumber (Cucumis sativus L.) using recombinant inbred lines. Euphytica 164:473–491
Zhang WW, Pan JS, He HL et al (2012) Construction of a high density integrated genetic map for cucumber (Cucumis sativus L.). Theor Appl Genet 124:249–259. doi:10.1007/s00122-011-1701-x
Zivy M, Devaux P, Blaisonneau J, Jean R, Thiellement H (1992) Segregation distortion and linkage studies in microspore-derived double haploid lines of Hordeum vulgare L. Theor Appl Genet 83:919–924. doi:10.1007/bf00226716
Acknowledgments
We are grateful for the assistance provided by the laboratory of Yiqun Weng (USDA-ARS, Vegetable Crops Research Unit and Horticultural Department, University of Wisconsin, 1575 Linden Drive, Madison, WI 53706). We also thank Zheng Li for ideas and Dr. Sikandar Hayat and Husain Ahmad for critical reading of the manuscript. This research was supported by a grant from the National Natural Science Foundation of China (Project #31372074).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical standards
The authors declare that this study complies with the current laws of the countries in which the experiments were performed.
Additional information
Communicated by S. Huang.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Liu, H., Meng, H., Pan, Y. et al. Fine genetic mapping of the white immature fruit color gene w to a 33.0-kb region in cucumber (Cucumis sativus L.). Theor Appl Genet 128, 2375–2385 (2015). https://doi.org/10.1007/s00122-015-2592-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-015-2592-z