Abstract
Key message
This manuscript provides a genetic map of Raphanus sativus that has been used as a reference genetic map for an ongoing genome sequencing project. The map was constructed based on genotyping by whole-genome resequencing of mapping parents and F 2 population.
Abstract
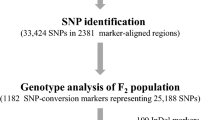
Raphanus sativus is an annual vegetable crop species of the Brassicaceae family and is one of the key plants in the seed industry, especially in East Asia. Assessment of the R. sativus genome provides fundamental resources for crop improvement as well as the study of crop genome structure and evolution. With the goal of anchoring genome sequence assemblies of R. sativus cv. WK10039 whose genome has been sequenced onto the chromosomes, we developed a reference genetic map based on genotyping of two parents (maternal WK10039 and paternal WK10024) and 93 individuals of the F2 mapping population by whole-genome resequencing. To develop high-confidence genetic markers, ~83 Gb of parental lines and ~591 Gb of mapping population data were generated as Illumina 100 bp paired-end reads. High stringent sequence analysis of the reads mapped to the 344 Mb of genome sequence scaffolds identified a total of 16,282 SNPs and 150 PCR-based markers. Using a subset of the markers, a high-density genetic map was constructed from the analysis of 2,637 markers spanning 1,538 cM with 1,000 unique framework loci. The genetic markers integrated 295 Mb of genome sequences to the cytogenetically defined chromosome arms. Comparative analysis of the chromosome-anchored sequences with Arabidopsis thaliana and Brassica rapa revealed that the R. sativus genome has evident triplicated sub-genome blocks and the structure of gene space is highly similar to that of B. rapa. The genetic map developed in this study will serve as fundamental genomic resources for the study of R. sativus.





Similar content being viewed by others
References
Al-Shehbaz I, Beilstein M, Kellogg E (2006) Systematics and phylogeny of the Brassicaceae (Cruciferae): an overview. Plant Syst Evol 259:89–120
Bancroft I, Morgan C, Fraser F, Higgins J, Wells R, Clissold L, Baker D, Long Y, Meng J, Wang X, Liu S, Trick M (2011) Dissecting the genome of the polyploid crop oilseed rape by transcriptome sequencing. Nat Biotechnol 29:762–766
Bett K, Lydiate D (2003) Genetic analysis and genome mapping in Raphanus. Genome 46:423–430
Budahn H, Peterka H, Mousa M, Ding Y, Zhang S, Li J (2009) Molecular mapping in oil radish (Raphanus sativus L.) and QTL analysis of resistance against beet cyst nematode (Heterodera schachtii). Theor Appl Genet 118:775–782
Cao J, Schneeberger K, Ossowski S, Günther T, Bender S, Fitz J, Koenig D, Lanz C, Stegle O, Lippert C, Wang X, Ott F, Müller J, Alonso-Blanco C, Borgwardt K, Schmid K, Weigel D (2011) Whole-genome sequencing of multiple Arabidopsis thaliana populations. Nat Genet 43:956–963
Celton J-M, Christoffels A, Sargent D, Xu X, Rees D (2010) Genome-wide SNP identification by high-throughput sequencing and selective mapping allows sequence assembly positioning using a framework genetic linkage map. BMC Biol 8:155
Cheng F, Mandáková T, Wu J, Xie Q, Lysak M, Wang X (2013) Deciphering the diploid ancestral genome of the Mesohexaploid Brassica rapa. Plant Cell 25:1541–1554
Chung H, Jeong Y-M, Mun J-H, Lee S-S, Chung W-H, Yu H-J (2014) Construction of a genetic map based on high throughput SNP genotyping and genetic mapping of a TuMV resistance locus in Brassica rapa. Mol Genet Genomics 289:149–160
Deschamps S, Llaca V, May G (2012) Genotyping-by-sequencing in plants. Biology 1:460–483
Edenberg H, Liu Y (2009) Laboratory methods for high-throughput genotyping. Cold Spring Harb Protoc 2009:pdb-top62
Elshire R, Glaubitz J, Sun Q, Poland J, Kawamoto K, Buckler E, Mitchell S (2011) A robust, simple genotyping-by-sequencing (GBS) approach for high diversity species. PLoS ONE 6:e19379
Hall A, Fiebig A, Preuss D (2002) Beyond the Arabidopsis genome: opportunities for comparative genomics. Plant Physiol 129:1439–1447
Huang X, Feng Q, Qian Q, Zhao Q, Wang L, Wang A, Guan J, Fan D, Weng Q, Huang T, Dong G, Sang T, Han B (2009) High-throughput genotyping by whole-genome resequencing. Genome Res 19:1068–1076
Huang S, Deng L, Guan M, Li J, Lu K, Wang H, Fu D, Mason A, Liu S, Hua W (2013) Identification of genome-wide single nucleotide polymorphisms in allopolyploid crop Brassica napus. BMC Genomics 14:717
Hwang Y-J, Yu H-J, Mun J-H, Ryu K, Park B-S, Lim K-B (2012) Centromere repeat DNA originated from Brassica rapa is detected in the centromere region of Raphanus sativus chromosomes. Korean J Hortic Sci Technol 30:751–756
Jeong Y-M, Chung W-H, Chung H, Kim N, Park B-S, Lim K-B, Yu H-J, Mun J-H (2014) Comparative analysis of the radish genome based on a conserved ortholog set (COS) of Brassica. Theor Appl Genet 127:1975–1989
Johnston J, Pepper A, Hall A, Chen Z, Hodnett G, Drabek J, Lopez R, Price H (2005) Evolution of genome size in Brassicaceae. Ann Bot 95:229–253
Jones E, Sullivan H, Bhattramakki D, Smith J (2007) A comparison of simple sequence repeat and single nucleotide polymorphism marker technologies for the genotypic analysis of maize (Zea mays L.). Theor Appl Genet 115:361–371
Kamei A, Tsuro M, Kobo N, Hayashi T, Wang N, Fujimura T, Hirai M (2010) QTL mapping of clubroot resistance in radish (Raphanus sativus L.). Theor Appl Genet 120:1021–1027
Kiss G, Kereszt A, Kiss P, Endre G (1998) Colormapping: a non-mathematical procedure for genetic mapping. Acta Biol Hung 49:125–142
Kitashiba H, Li F, Hirakawa H, Kawanabe T, Zou Z, Hasegawa Y, Tonosaki K, Shirasawa S, Fukushima A, Yokoi S, Takahata Y, Kakizaki T, Ishida M, Okamoto S, Sakamoto K, Shirasawa K, Tabata S, Nishio T (2014) Draft sequences of the radish (Raphanus sativus L.) genome. DNA Res. doi:10.1093/dnares/dsu014
Kosambi D (1944) The estimation of map distances from recombination values. Ann Eugen 12:172–175
Lai K, Duran C, Berkman P, Lorenc M, Stiller J, Manoli S, Hayden M, Forrest K, Fleury D, Baumann U, Zander M, Mason A, Batley J, Edwards D (2012) Single nucleotide polymorphism discovery from wheat next-generation sequence data. Plant Biotechnol J 10:743–749
Lee S-S, Lee S-A, Yang J, Kim J (2011) Developing stable progenies of xBrassicoraphanus, an intergeneric allopolyploid between Brassica rapa and Raphanus sativus, through induced mutation using microspore culture. Theor Appl Genet 122:885–891
Li H, Durbin R (2009) Fast and accurate short read alignment with Burrows–Wheeler Transform. Bioinformatics 25:1754–1760
Li F, Hasegawa Y, Saito M, Shirasawa S, Fukushima A, Ito T, Fujii H, Kishitani S, Kitashiba H, Nishio T (2011) Extensive chromosome homoeology among Brassiceae species were revealed by comparative genetic mapping with high-density EST-based SNP markers in radish (Raphanus sativus L.). DNA Res 18:401–411
Lim K, de Jong H, Yang T, Park J, Kwon S, Kim J, Lim M, Kim J, Jin M, Jin Y, Kim S, Lim Y, Bang J, Kim H, Park B (2005) Characterization of rDNA and tandem repeats in the heterochromatin of Brassica rapa. Mol Cell 19:436–444
Lysak M, Koch M, Pecinka A, Schubert I (2005) Chromosome triplication found across the tribe Brassiceae. Genome Res 15:516–525
McKenna A, Hanna M, Banks E, Sivachenko A, Cibulskis K, Kernytsky A, Garimella K, Altshuler D, Gabriel S, Daly M, DePristo M (2010) The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res 20:1297–1303
Mun J-H, Kim D-J, Choi H-K, Gish J, Debellé F, Mudge J, Denny R, Endré G, Saurat O, Dudez A, Kiss G, Roe B, Young N, Cook D (2006) Distribution of microsatellites in the genome of Medicago truncatula: a resource of genetic markers that integrate genetic and physical maps. Genetics 172:2541–2555
Mun J-H, Kwon SJ, Yang TJ, Seol YJ, Jin M, Kim JA, Lim MH, Kim JS, Lee SI, Baek S, Choi BS, Kim DS, Kim N, Yu HJ, Lim KB, Lim YP, Bancroft I, Hahn JH, Park BS (2009) Genome-wide comparative analysis of the Brassica rapa gene space reveals genome shrinkage and differential loss of duplicated genes after whole genome triplication. Genome Biol 10:R111
Nussbaumer T, Martis M, Roessner S, Pfeifer M, Bader K, Sharma S, Gundlach H, Spannagl M (2013) MIPS PlantsDB: a database framework for comparative plant genome research. Nucleic Acids Res 41:D1144–D1151
Ossowski S, Schneeberger K, Clark R, Lanz C, Warthmann N, Weigel D (2008) Sequencing of natural strains of Arabidopsis thaliana with short reads. Genome Res 18:2024–2033
Park S, Yu HJ, Mun JH, Lee SC (2010) Genome-wide discovery of DNA polymorphism in Brassica rapa. Mol Genet Genomics 283:135–145
Prakash S, Bhat S, Quiros C, Kirti P, Chopra V (2009) Brassica and its close allies: cytogenetics and evolution. In: Jules J (ed) Plant Breed Reviews, vol 31. Wiley, London, pp 21–187
Rozen S, Skaletsky H (1999) Primer3 on the WWW for General Users and for Biologist Programmers. In: Misener S, Krawetz S (eds) Bioinformatics Methods and Protocols. Humana Press, Totowa, pp 365–386
Santosh K, Travis W, Sylvie C (2012) SNP Discovery through next-generation sequencing and its applications. Int J Plant Genomics 2012:ID 831460
Shen D, Sun H, Huang M, Zheng Y, Li X, Fei Z (2013) RadishBase: a database for genomics and genetics of radish. Plant Cell Physiol 54:e3
Shirasawa K, Oyama M, Hirakawa H, Sato S, Tabata S, Fujioka T, Kimizuka-Takagi C, Sasamoto S, Watanabe A, Kato M, Kishida Y, Kohara M, Takahashi C, Tsuruoka H, Wada T, Sakai T, Isobe S (2011) An EST-SSR linkage map of Raphanus sativus and comparative genomics of the Brassicaceae. DNA Res 18:221–232
Sonah H, Bastien M, Iquira E, Tardivel A, Légaré G, Boyle B, Normandeau É, Laroche J, Larose S, Jean M, Belzile F (2013) An improved genotyping by sequencing (GBS) approach offering increased versatility and efficiency of SNP discovery and genotyping. PLoS ONE 8:e54603
Song K, Osborn T, Williams P (1990) Brassica taxonomy based on nuclear restriction fragment length polymorphisms (RFLPs): 3. Genomic relationships in Brassica and related genera and the origin of B. oleracea and B. rapa (syn. campestris). Theor Appl Genet 79:497–506
Subbaiyan G, Waters D, Katiyar S, Sadananda A, Vaddadi S, Henry R (2012) Genome-wide DNA polymorphisms in elite indica rice inbreds discovered by whole-genome sequencing. Plant Biotechnol J 10:623–634
Trick M, Adamski N, Mugford S, Jiang C-C, Febrer M, Uauy C (2012) Combining SNP discovery from next-generation sequencing data with bulked segregant analysis (BSA) to fine-map genes in polyploidy wheat. BMC Plant Biol 12:14
Tsuro M, Suwabe K, Kobo N, Matsumoto S, Hirai M (2005) Construction of a molecular linkage map of radish (Raphanus sativus L.), based on AFLP and Brassica-SSR markers. Breed Sci 55:107–111
Uitdewilligen J, Wolters A, D’hoop B, Borm T, Visser R, van Eck H (2013) A next-generation sequencing method for genotyping-by-sequencing of highly heterozygous autotetraploid potato. PLoS ONE 8:e62355
van Ooijen JW (2006) JoinMap® 4, Software for the calculation of genetic linkage maps in experimental populations. Kyazma B. V, Wageningen
Wang S, Wang X, He Q, Liu X, Xu W, Li L, Gao J, Wang F (2012) Transcriptome analysis of the roots at early and late seedling stages using Illumina paired-end sequencing and development of EST-SSR markers in radish. Plant Cell Rep 31:1437–1447
Xu X, Liu X, Ge S, Jensen J, Hu F, Li X, Dong Y, Gutenkunst R, Fang L, Huang L, Li J, He W, Zhang G, Zheng X, Zhang F, Li Y, Yu C, Kristiansen K, Zhang X, Wang J, Wright M, McCouch S, Nielsen R, Wang J, Wang W (2011) Resequencing 50 accessions of cultivated and wild rice yields markers for identifying agronomically important genes. Nat Biotechnol 30:105–111
Yang Y, Tai P, Chen Y, Li W (2002) A study of the phylogeny of Brassica rapa, B. nigra, Raphanus sativus, and their related genera using noncoding regions of chloroplast DNA. Mol Phylogenet Evol 23:268–275
Yu H-J, Park S-G, Oh M, Hwang H-J, Kim N, Chung H, Sohn S-H, Park B-S, Mun J-H (2011) The Brassica rapa tissue-specific EST database. Korean J Hortic Sci Technol 29:633–640
Acknowledgments
This work was supported by grants from the National Academy of Agricultural Science (PJ009795), RDA to JHM and the Next-Generation Biogreen21 program (PJ008019), RDA to HJY.
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical standard
The authors declare that the experiments complied with current laws of the country in which they were performed.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by Isobel Parkin.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Mun, JH., Chung, H., Chung, WH. et al. Construction of a reference genetic map of Raphanus sativus based on genotyping by whole-genome resequencing. Theor Appl Genet 128, 259–272 (2015). https://doi.org/10.1007/s00122-014-2426-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-014-2426-4