Abstract
Purpose
Multi-modal brain imaging provides different in vivo windows into the human brain and thereby different ways to characterize brain disorders. Particularly, resting-state functional magnetic resonance imaging facilitates the study of macroscopic intrinsic brain networks, which are critical for development and spread of neurodegenerative processes in different neurodegenerative diseases. The aim of the current study is to present and highlight some paradigmatic findings in intrinsic network-based pathophysiology of neurodegenerative diseases and its potential for new network-based multimodal tools in imaging diagnostics.
Methods
Qualitative review of selected multi-modal imaging studies in neurodegenerative diseases particularly in Alzheimer’s disease (AD).
Results
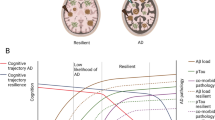
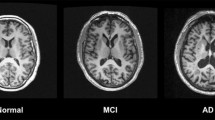
Functional connectivity of intrinsic brain networks is selectively and progressively impaired in AD, with changes likely starting before the onset of symptoms in fronto-parietal key networks such as default mode or attention networks. Patterns of distribution and development of both amyloid-β plaques and atrophy are linked with network connectivity changes, suggesting that start and spread of pathology interacts with network connectivity. Qualitatively similar findings have been observed in other neurodegenerative disorders, suggesting shared mechanisms of network-based pathophysiology across diseases.
Conclusion
Spread of neurodegeneration is intimately linked with the functional connectivity of intrinsic brain networks. These pathophysiological insights pave the way for new multi-modal network-based tools to detect and characterize neurodegeneration in individual patients.

Similar content being viewed by others
References
Fox MD, Raichle ME. Spontaneous fluctuations in brain activity observed with functional magnetic resonance imaging. Nat Rev Neurosci. 2007;8:700–11.
Vincent JL, Patel GH, Fox MD, Snyder AZ, Baker JT, Van Essen DC, Zempel JM, Snyder LH, Corbetta M, Raichle ME. Intrinsic functional architecture in the anaesthetized monkey brain. Nature. 2007;447:83–6.
Doria V, Beckmann CF, Arichi T, Merchant N, Groppo M, Turkheimer FE, Counsell SJ, Murgasova M, Aljabar P, Nunes RG, Larkman DJ, Rees G, Edwards AD. Emergence of resting state networks in the preterm human brain. Proc Natl Acad Sci U S A. 2010;107:20015–20.
Smith SM, Fox PT, Miller KL, Glahn DC, Fox PM, Mackay CE, Filippini N, Watkins KE, Toro R, Laird AR, Beckmann CF. Correspondence of the brain’s functional architecture during activation and rest. Proc Natl Acad Sci U S A. 2009;106:13040–5.
Pievani M, De Haan W, Wu T, Seeley WW, Frisoni GB. Functional network disruption in the degenerative dementias. Lancet Neurol. 2011;10:829–43.
Pievani M, Filippini N, Van Den Heuvel MP, Cappa SF, Frisoni GB. Brain connectivity in neurodegenerative diseases—from phenotype to proteinopathy. Nat Rev Neurol. 2014;10:620–33.
Greicius MD, Srivastava G, Reiss AL, Menon V. Default-mode network activity distinguishes Alzheimer’s disease from healthy aging: evidence from functional MRI. Proc Natl Acad Sci U S A. 2004;101:4637–42.
Sorg C, Riedl V, Mühlau M, Calhoun VD, Eichele T, Läer L, Drzezga A, Förstl H, Kurz A, Zimmer C, Wohlschläger AM. Selective changes of resting-state networks in individuals at risk for Alzheimer’s disease. Proc Natl Acad Sci U S A. 2007;104:18760–5.
Agosta F, Pievani M, Geroldi C, Copetti M, Frisoni GB, Filippi M. Resting state fMRI in Alzheimer’s disease: beyond the default mode network. Neurobiol Aging. 2012;33:1564–78.
Lim HK, Nebes R, Snitz B, Cohen A, Mathis C, Price J, Weissfeld L, Klunk W, Aizenstein HJ. Regional amyloid burden and intrinsic connectivity networks in cognitively normal elderly subjects. Brain. 2014;137:3327–38.
Hardy J, Selkoe DJ. The amyloid hypothesis of Alzheimer’s disease: progress and problems on the road to therapeutics. Science. 2002;297:353–6.
Zhou J, Greicius MD, Gennatas ED, Growdon ME, Jang JY, Rabinovici GD, Kramer JH, Weiner M, Miller BL, Seeley WW. Divergent network connectivity changes in behavioural variant frontotemporal dementia and Alzheimer’s disease. Brain. 2010;133:1352–67.
Luo C, Song W, Chen Q, Zheng Z, Chen K, Cao B, Yang J, Li J, Huang X, Gong Q, Shang HF. Reduced functional connectivity in early-stage drug-naive Parkinson’s disease: a resting-state fMRI study. Neurobiol Aging. 2014;35:431–41.
Seeley WW, Crawford RK, Zhou J, Miller BL, Greicius MD. Neurodegenerative diseases target large-scale human brain networks. Neuron. 2009;62:42–52.
Zhou J, Gennatas ED, Kramer JH, Miller BL, Seeley WW. Predicting regional neurodegeneration from the healthy brain functional connectome. Neuron. 2012;73:1216–27.
Sperling RA, Laviolette PS, O'Keefe K, O'Brien J, Rentz DM, Pihlajamaki M, Marshall G, Hyman BT, Selkoe DJ, Hedden T, Buckner RL, Becker JA, Johnson KA. Amyloid deposition is associated with impaired default network function in older persons without dementia. Neuron. 2009;63:178–88.
Drzezga A, Becker JA, Van Dijk KR, Sreenivasan A, Talukdar T, Sullivan C, Schultz AP, Sepulcre J, Putcha D, Greve D, Johnson KA, Sperling RA. Neuronal dysfunction and disconnection of cortical hubs in non-demented subjects with elevated amyloid burden. Brain. 2011;134:1635–46.
Myers N, Pasquini L, Göttler J, Grimmer T, Koch K, Ortner M, Neitzel J, Mühlau M, Förster S, Kurz A, Förstl H, Zimmer C, Wohlschläger AM, Riedl V, Drzezga A, Sorg C. Within-patient correspondence of amyloid-beta and intrinsic network connectivity in Alzheimer’s disease. Brain. 2014;137:2052–64.
Iturria-Medina Y, Sotero RC, Toussaint PJ, Evans AC. Epidemic spreading model to characterize misfolded proteins propagation in aging and associated neurodegenerative disorders. PLoS Comput Biol. 2014;10:e1003956.
Raj A, Kuceyeski A, Weiner M. A network diffusion model of disease progression in dementia. Neuron. 2012;73:1204–15.
Raj A, Locastro E, Kuceyeski A, Tosun D, Relkin N, Weiner M; for the Alzheimer’s Disease Neuroimaging Initiative (ADNI). Network diffusion model of progression predicts longitudinal patterns of atrophy and metabolism in Alzheimer’s disease. Cell Rep. 2015; 10:359–69.
Deco G, Jirsa VK, Mcintosh AR. Resting brains never rest: computational insights into potential cognitive architectures. Trends Neurosci. 2013;36:268–74.
Honey CJ, Kotter R, Breakspear M, Sporns O. Network structure of cerebral cortex shapes functional connectivity on multiple time scales. Proc Natl Acad Sci U S A. 2007;104:10240–5.
Hahn K, Myers N, Prigarin S, Rodenacker K, Kurz A, Förstl H, Zimmer C, Wohlschläger AM, Sorg C. Selectively and progressively disrupted structural connectivity of functional brain networks in Alzheimer’s disease—revealed by a novel framework to analyze edge distributions of networks detecting disruptions with strong statistical evidence. Neuroimage. 2013;81:96–109.
Buckner RL, Sepulcre J, Talukdar T, Krienen FM, Liu H, Hedden T, Andrews-Hanna JR, Sperling RA, Johnson KA. Cortical hubs revealed by intrinsic functional connectivity: mapping, assessment of stability, and relation to Alzheimer’s disease. J Neurosci. 2009;29:1860–73.
Zhang Y, Schuff N, Du AT, Rosen HJ, Kramer JH, Gorno-Tempini ML, Miller BL, Weiner MW. White matter damage in frontotemporal dementia and Alzheimer’s disease measured by diffusion MRI. Brain. 2009;132:2579–92.
Shao J, Myers N, Yang Q, Feng J, Plant C, Böhm C, Förstl H, Kurz A, Zimmer C, Meng C, Riedl V, Wohlschläger A, Sorg C. Prediction of Alzheimer’s disease using individual structural connectivity networks. Neurobiol Aging. 2012;33:2756–65.
Wee CY, Yap PT, Zhang D, Denny K, Browndyke JN, Potter GG, Welsh-Bohmer KA, Wang L, Shen D. Identification of MCI individuals using structural and functional connectivity networks. Neuroimage. 2012;59:2045–56.
Dyrba M, Grothe M, Kirste T, Teipel SJ. Multimodal analysis of functional and structural disconnection in Alzheimer’s disease using multiple kernel SVM. Hum Brain Mapp. 2015;6:2118–31.
Fox MD, Greicius M. Clinical applications of resting state functional connectivity. Front Syst Neurosci. 2010;4:19.
Biswal BB, Mennes M, Zuo XN, Gohel S, Kelly C, Smith SM, Beckmann CF, Adelstein JS, Buckner RL, Colcombe S, Dogonowski AM, Ernst M, Fair D, Hampson M, Hoptman MJ, Hyde JS, Kiviniemi VJ, Kötter R, Li SJ, Lin CP, Lowe MJ, Mackay C, Madden DJ, Madsen KH, Margulies DS, Mayberg HS, McMahon K, Monk CS, Mostofsky SH, Nagel BJ, Pekar JJ, Peltier SJ, Petersen SE, Riedl V, Rombouts SA, Rypma B, Schlaggar BL, Schmidt S, Seidler RD, Siegle GJ, Sorg C, Teng GJ, Veijola J, Villringer A, Walter M, Wang L, Weng XC, Whitfield-Gabrieli S, Williamson P, Windischberger C, Zang YF, Zhang HY, Castellanos FX, Milham MP. Toward discovery science of human brain function. Proc Natl Acad Sci U S A. 2010;107:4734–9.
Smith SM, Beckmann CF, Andersson J, Auerbach EJ, Bijsterbosch J, Douaud G, Duff E, Feinberg DA, Griffanti L, Harms MP, Kelly M, Laumann T, Miller KL, Moeller S, Petersen S, Power J, Salimi-Khorshidi G, Snyder AZ, Vu AT, Woolrich MW, Xu J, Yacoub E, Uğurbil K, Van Essen DC, Glasser MF; WU-Minn HCP Consortium. Resting-state fMRI in the human connectome project. Neuroimage. 2013;80:144–68.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Sorg, C., Göttler, J. & Zimmer, C. Imaging Neurodegeneration: Steps Toward Brain Network-Based Pathophysiology and Its Potential for Multi-modal Imaging Diagnostics. Clin Neuroradiol 25 (Suppl 2), 177–181 (2015). https://doi.org/10.1007/s00062-015-0438-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00062-015-0438-3