Abstract
Objective
The purpose of our study was to examine whether the H19 rs2839698, rs217727, and HOX transcript antisense RNA (HOTAIR) rs12826786 polymorphisms were associated with genetic susceptibility to rheumatoid arthritis (RA) in a Chinese population.
Methods
A total of 777 participants were enrolled in this study, including 328 RA patients and 449 healthy controls. The H19 rs2839698, rs217727, and HOTAIR rs12826786 polymorphisms were detected by the ligase detection reaction-polymerase chain reaction (LDR-PCR) technology.
Results
No significant difference in genotype distribution between RA patients and healthy controls was found (P = 0.38 for rs2839698; P = 0.79 for rs217727; P = 0.39 for rs12826786). The difference in allele frequencies between RA patients and controls was also non-significant (rs2839698 T versus C, P = 0.23, odds ratio (OR) = 1.15, 95% confidence interval (CI) = 0.92–1.43; rs217727 C versus T, P = 0.55, OR = 1.07, 95% CI = 0.87–1.32; and rs12826786 T versus C, P = 0.32, OR = 1.14, 95% CI = 0.88–1.47). We have also evaluated the relationships of above-mentioned polymorphisms with risk of RA under dominant model and recessive model, but non-significant evidence was found. No significant evidence was detected for the relationships of H19 rs2839698, rs217727, and HOTAIR rs12826786 polymorphisms with risk of different serotypes of RA.
Conclusions
Our results indicated that H19 rs2839698, rs217727, and HOTAIR rs12826786 polymorphisms might not be involved in the genetic background of RA in Chinese.
Similar content being viewed by others
References
McInnes IB, Schett G. The pathogenesis of rheumatoid arthritis. N Engl J Med. 2011;365(23):2205–19.
Li R, Sun J, Ren LM, Wang HY, Liu WH, Zhang XW, et al. Epidemiology of eight common rheumatic diseases in China: a large-scale cross-sectional survey in Beijing. Rheumatology (Oxford). 2012;51(4):721–9.
Terao C, Ikari K, Nakayamada S, Takahashi Y, Yamada R, Ohmura K, et al. A twin study of rheumatoid arthritis in the Japanese population. Mod Rheumatol. 2016;26(5):685–9.
Silman AJ, MacGregor AJ, Thomson W, Holligan S, Carthy D, Farhan A, et al. Twin concordance rates for rheumatoid arthritis: results from a nationwide study. Br J Rheumatol. 1993;32(10):903–7.
Bellamy N, Duffy D, Martin N, Mathews J. Rheumatoid arthritis in twins: a study of aetiopathogenesis based on the Australian Twin Registry. Ann Rheum Dis. 1992;51(5):588–93.
Frisell T, Holmqvist M, Källberg H, Klareskog L, Alfredsson L, Askling J. Familial risks and heritability of rheumatoid arthritis: role of rheumatoid factor/anti-citrullinated protein antibody status, number and type of affected relatives, sex, and age. Arthritis Rheum. 2013;65(11):2773–82.
Hemminki K, Li X, Sundquist J, Sundquist K. Familial associations of rheumatoid arthritis with autoimmune diseases and related conditions. Arthritis Rheum. 2009;60(3):661–8.
Grant SF, Thorleifsson G, Frigge ML, Thorsteinsson J, Gunnlaugsdóttir B, Geirsson AJ, et al. The inheritance of rheumatoid arthritis in Iceland. Arthritis Rheum. 2001;44(10):2247–54.
del Junco D, Luthra HS, Annegers JF, Worthington JW, Kurland LT. The familial aggregation of rheumatoid arthritis and its relationship to the HLA-DR4 association. Am J Epidemiol. 1984;119(5):813–29.
Okada Y, Wu D, Trynka G, Raj T, Terao C, Ikari K, et al. Genetics of rheumatoid arthritis contributes to biology and drug discovery. Nature. 2014;506(7488):376–81.
Ding J, Eyre S, Worthington J. Genetics of RA susceptibility, what comes next? RMD Open. 2015;1(1):e000028.
Erdmann VA, Szymanski M, Hochberg A, Groot N, Barciszewski J. Non-coding, mRNA-like RNAs database Y2K. Nucleic Acids Res. 2000;28(1):197–200.
Mercer TR, Dinger ME, Mattick JS. Long non-coding RNAs: insights into functions. Nat Rev Genet. 2009;10(3):155–9.
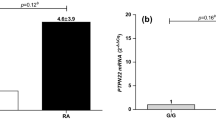
Stuhlmüller B, Kunisch E, Franz J, Martinez-Gamboa L, Hernandez MM, Pruss A, et al. Detection of oncofetal h19 RNA in rheumatoid arthritis synovial tissue. Am J Pathol. 2003;163(3):901–11.
Song J, Kim D, Han J, Kim Y, Lee M, Jin EJ. PBMC and exosome-derived Hotair is a critical regulator and potent marker for rheumatoid arthritis. Clin Exp Med. 2015;15(1):121–6.
Rinn JL, Kertesz M, Wang JK, Squazzo SL, Xu X, Brugmann SA, et al. Functional demarcation of active and silent chromatin domains in human HOX loci by noncoding RNAs. Cell. 2007;129(7):1311–23.
Raveh E, Matouk IJ, Gilon M, Hochberg A. The H19 Long non-coding RNA in cancer initiation, progression and metastasis - a proposed unifying theory. Mol Cancer. 2015;14:184.
Hajjari M, Salavaty A. HOTAIR: an oncogenic long non-coding RNA in different cancers. Cancer Biol Med. 2015;12(1):1–9.
Verhaegh GW, Verkleij L, Vermeulen SH, den Heijer M, Witjes JA, Kiemeney LA. Polymorphisms in the H19 gene and the risk of bladder cancer. Eur Urol. 2008;54(5):1118–26.
Yang C, Tang R, Ma X, Wang Y, Luo D, Xu Z, et al. Tag SNPs in long non-coding RNA H19 contribute to susceptibility to gastric cancer in the Chinese Han population. Oncotarget. 2015;6(17):15311–20.
Li S, Hua Y, Jin J, Wang H, Du M, Zhu L, et al. Association of genetic variants in lncRNA H19 with risk of colorectal cancer in a Chinese population. Oncotarget. 2016;7(18):25470–7.
Xia Z, Yan R, Duan F, Song C, Wang P, Wang K. Genetic polymorphisms in long noncoding RNA H19 are associated with susceptibility to breast cancer in Chinese population. Medicine (Baltimore). 2016;95(7):e2771.
Hua Q, Lv X, Gu X, Chen Y, Chu H, Du M, et al. Genetic variants in lncRNA H19 are associated with the risk of bladder cancer in a Chinese population. Mutagenesis. 2016;31(5):531–8.
Guo W, Dong Z, Bai Y, Guo Y, Shen S, Kuang G, et al. Associations between polymorphisms of HOTAIR and risk of gastric cardia adenocarcinoma in a population of north China. Tumour Biol. 2015;36(4):2845–54.
Bayram S, Sümbül AT, Dadaş E. A functional HOTAIR rs12826786 C > T polymorphism is associated with breast cancer susceptibility and poor clinicopathological characteristics in a Turkish population: a hospital-based case-control study. Tumour Biol. 2016;37(4):5577–84.
Ohmura K, Terao C, Maruya E, Katayama M, Matoba K, Shimada K, et al. Anti-citrullinated peptide antibody-negative RA is a genetically distinct subset: a definitive study using only bone-erosive ACPA-negative rheumatoid arthritis. Rheumatology (Oxford). 2010;49(12):2298–304.
Viatte S, Plant D, Bowes J, Lunt M, Eyre S, Barton A, et al. Genetic markers of rheumatoid arthritis susceptibility in anti-citrullinated peptide antibody negative patients. Ann Rheum Dis. 2012;71(12):1984–90.
Han B, Diogo D, Eyre S, Kallberg H, Zhernakova A, Bowes J, et al. Fine mapping seronegative and seropositive rheumatoid arthritis to shared and distinct HLA alleles by adjusting for the effects of heterogeneity. Am J Hum Genet. 2014;94(4):522–32.
Viatte S, Massey J, Bowes J, Duffus K, arcOGEN Consortium, Eyre S, et al. Replication of associations of genetic loci outside the HLA region with susceptibility to anti-cyclic citrullinated peptide-negative rheumatoid arthritis. Arthritis Rheumatol. 2016;68(7):1603–13.
Arnett FC, Edworthy SM, Bloch DA, McShane DJ, Fries JF, Cooper NS, et al. The American Rheumatism Association 1987 revised criteria for the classification of rheumatoid arthritis. Arthritis Rheum. 1988;31(3):315–24.
Aletaha D, Neogi T, Silman AJ, Funovits J, Felson DT, Bingham CO, et al. Rheumatoid arthritis classification criteria: an American College of Rheumatology/European League Against Rheumatism collaborative initiative. Arthritis Rheum. 2010;62(9):2569–2581.
Brannan CI, Dees EC, Ingram RS, Tilghman SM. The product of the H19 gene may function as an RNA. Mol Cell Biol. 1990;10(1):28–36.
Gabory A, Jammes H, Dandolo L. The H19 locus: role of an imprinted non-coding RNA in growth and development. Bioessays. 2010;32(6):473–80.
Keniry A, Oxley D, Monnier P, Kyba M, Dandolo L, Smits G, et al. The H19 lincRNA is a developmental reservoir of miR-675 that suppresses growth and Igf1r. Nat Cell Biol. 2012;14(7):659–65.
Kallen AN, Zhou XB, Xu J, Qiao C, Ma J, Yan L, et al. The imprinted H19 lncRNA antagonizes let-7 microRNAs. Mol Cell. 2013;52(1):101–12.
Xia T, Liao Q, Jiang X, Shao Y, Xiao B, Xi Y, et al. Long noncoding RNA associated-competing endogenous RNAs in gastric cancer. Sci Rep. 2014;4:6088.
Matouk IJ, Halle D, Raveh E, Gilon M, Sorin V, Hochberg A. The role of the oncofetal H19 lncRNA in tumor metastasis: orchestrating the EMT-MET decision. Oncotarget. 2016;7(4):3748–65.
Wang WT, Ye H, Wei PP, Han BW, He B, Chen ZH, et al. LncRNAs H19 and HULC, activated by oxidative stress, promote cell migration and invasion in cholangiocarcinoma through a ceRNA manner. J Hematol Oncol. 2016;9(1):117.
Gupta RA, Shah N, Wang KC, Kim J, Horlings HM, Wong DJ, et al. Long non-coding RNA HOTAIR reprograms chromatin state to promote cancer metastasis. Nature. 2010;464(7291):1071–6.
Tsai MC, Manor O, Wan Y, Mosammaparast N, Wang JK, Lan F, et al. Long noncoding RNA as modular scaffold of histone modification complexes. Science. 2010;329(5992):689–93.
Ma MZ, Li CX, Zhang Y, Weng MZ, Zhang MD, Qin YY, et al. Long non-coding RNA HOTAIR, a c-Myc activated driver of malignancy, negatively regulates miRNA-130a in gallbladder cancer. Mol Cancer. 2014;13:156.
Liu XH, Sun M, Nie FQ, Ge YB, Zhang EB, Yin DD, et al. Lnc RNA HOTAIR functions as a competing endogenous RNA to regulate HER2 expression by sponging miR-331-3p in gastric cancer. Mol Cancer. 2014;13:92.
Acknowledgements
This work was supported by National Natural Science Foundation of China (Grant No. 81602921), Nature Science Foundation of Ningbo city (Grant No. 2015A610205), Zhejiang Provincial Key Laboratory of Pathophysiology, Ningbo University Talent Project (Grant No. 421504530), School Research Foundation of Ningbo University (Grant No. XKL14D2094), Ningbo Scientific Innovation Team for Environmental Hazardous Factor Control and Prevention (Grant No. 2016C51001), and K.C. Wong Magna Fund in Ningbo University.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that there is no conflict of interest associated with this manuscript.
Additional information
Communicated by John Di Battista.
J.-Z. Zhou, J.-J. Li and D.-J. Hua contributed equally to this work and should be considered as co-first author.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Zhou, JZ., Li, JJ., Hua, DJ. et al. A study on associations of single-nucleotide polymorphisms within H19 and HOX transcript antisense RNA (HOTAIR) with genetic susceptibility to rheumatoid arthritis in a Chinese population. Inflamm. Res. 66, 515–521 (2017). https://doi.org/10.1007/s00011-017-1035-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00011-017-1035-5