Summary
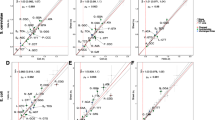
A characteristic profile of the fluctuations of codon usage is observed in bacteriophages and mitochondria. By following the DNA in the direction of transcription, one moves slowly from a region where selective pressure favours codons ending with C to a region where the bias is in favour of codons ending with T; then, abruptly, one again enters a region of codons ending in C. The transcription end point takes place in the area of abrupt change in codon usage. By comparing Drosophila yakuba and mouse mitochondrial genomes, it is possible to show that the strategy of codon usage for a given gene depends on its location along the transcription unit and not on the encoded protein. The choice of codons ending in T or C allows large scale variations of DNA stability which could regulate the speed of propagation of the RNA polymerase.
Similar content being viewed by others
References
Anderson S, Bankier AT, Barrell BG, de Bruijn MHL, Coulson AR, Drouin J, Eperon IC, Nierlich DP, Roe BA, Sanger F, Schreier PH, Smith AJH, Staden R, Young IG (1981) Nature 290:457–465
Anderson S, de Bruijn MHL, Coulson AR, Eperon IC, Sanger F, Young IG (1982) J Mol Biol 156:683–717
Attardi G (1981) Trends Biochem Sci 6:100–103
Attardi G, Chomyn A, Doolittle RF, Mariottini P, Ragan CI (1986) Cold Spring Harbor Symp Quant Biol 51:103–114
Beck E, Sommer R, Auerswald EA, Kurz C, Zink B, Osterburg G, Schaller H, Sugimoto K, Sugisaki H, Okamoto T, Takanami M (1978) Nucleic Acids Res 5:4495–4503
Benzécri JP (1973) L'analyse des données. Dunod, Paris
Bernardi G, Olofsson B, Filipski J, Zerial M, Salinas J, Cuny G, Meunier-Rotival M, Rodier F (1985) Science 228:953–958
Bernardi G, Mouchiroud D, Gautier C, Bernardi G (1988) J Mol Evol 28:7–18
Bibb MJ, van Etten RA, Wright CT, Walberg MW, Clayton DA (1981) Cell 26:167–180
Clary DO, Wolstenholme DR (1985) J Mol Evol 22:252–271
Gadaleta G, Pepe G, de Candia G, Quagliariello C, Sbisa E, Saccone C (1989) J Mol Evol 28:497–516
Godson GN, Barrell BG, Staden R, Fiddes JC (1978) Nature 276:236–247
Gouy M, Gautier C (1982) Nucleic Acids Res 10:7055–7073
Grantham R, Gautier C (1980) Naturwissenschaften 67:93–94
Grantham R, Gautier C, Gouy M, Mercier R, Pavé A (1980) Nucleic Acids Res 8:49–62
Hénaut A, Limaiem J, Vigier P (1985) Biochemie 67:475–483
Hill MO (1974) Appl Statist 23:340–354
Jacobs HT, Elliott DJ, Math VB, Farquharson A (1988) J Mol Biol 202:185–217
Kendall M (1975) Rank correlation methods. 4th edn. Charles Griffin and Co., High Wycombe
Lébart L, Morineau A, Warwick KA (1984) Multivariate descriptive statistical analysis. John Wiley and Sons, New York
Limaiem J, Hénaut A (1984a) C R Acad Sc Paris 298 Série III:279–286
Limaiem J, Hénaut A (1984b) C R Acad Sc Paris 299 Série III:275–280
Mouchiroud D, Gautier C, Bernardi G (1988) J Mol Evol 27:311–320
Roe BA, Ma DP, Wilson RK, Wong JF (1985) J Biol Chem 260:9759–9774
Sanger F, Coulson AR, Friedmann T, Air GM, Barrell BG, Brown NL, Fiddes JC, Hutchinson CA, Slocombe PM, Smith M (1978) J Mol Biol 125:225–246
Sharp PM, Devine KM (1989) Nucleic Acids Res 17:5029–5039
Sharp PM, Cowe E, Higgins DG, Shields DC, Wolfe HK, Wright F (1988) Nucleic Acids Res 16:8207–8211
Shields DC, Sharp PM (1987) Nucleic Acids Res 15:3023–3040
Wezenbeek P van, Hulsebos T, Schoenmakers J (1980) Gene 11:129–148
Zar J (1972) J Am Statist Assoc 67:578–580
Author information
Authors and Affiliations
Additional information
Communicated by K. Wolf
Rights and permissions
About this article
Cite this article
Delorme, MO., Hénaut, A. Codon usage is imposed by the gene location in the transcription unit. Curr Genet 20, 353–358 (1991). https://doi.org/10.1007/BF00317061
Received:
Issue Date:
DOI: https://doi.org/10.1007/BF00317061