Abstract
In mammals, both circadian rhythm and aging play important roles in regulating time-dependent homeostasis. We previously discovered an age-related increase element binding protein, hnRNP A3, which binds to the 3′-untranslated region (UTR) of blood coagulation factor IX (FIX). Here, we describe other members of this protein family, hnRNP C and hnRNP H, which bind to the 3′-UTR of the mouse circadian clock gene Period 2 (mPer2). RNA electrophoretic mobility shift assays using a 32P-labeled Per2 RNA probe coupled with two-dimensional gel electrophoresis followed by MALDI-TOF/MS peptide mass fingerprint analysis was used to analyze these proteins. Western blotting suggested that the total expression of these proteins in mouse liver cell nuclei does not increase with age. Two-dimensional gel electrophoresis analysis of age-related protein expression showed that many isoforms of these proteins exist in the liver and that each protein exhibits a complex age-related expression pattern. These results suggest that many isoforms of proteins are regulated by different aging systems and that many age regulation systems function in the liver.
Similar content being viewed by others
Introduction
In homeostatic systems, stable gene expression is maintained during aging via several factors, such as the age-related stability element (ASE)/age-related increase element (AIE) [1–4]. The AIE, originally identified in the hFIX gene, contains a 102-bp stretch of dinucleotide repeats (AT, GT, and CA) that have the potential to form distinct stem-loop (sl) [1–4] RNA structures (FIX-AIE RNA) in the 3′-untranslated region (3′-UTR) after the gene is transcribed [5]. We recently reported that heterogeneous nuclear ribonucleoprotein (hnRNP) A3 binds to the AIE and plays an important role in age-related gene expression [4]. HnRNP A3 is a member of the hnRNP family of proteins. These proteins, which are generally known to bind RNA, are most abundant in the nucleus of mammalian cells, where they play critical roles in the synthesis and processing of precursor RNAs in the nucleus and mRNA transportation from the nucleus to the cytosol. HnRNP A3 recognizes the overall three-dimensional sl structures formed by AIE RNA rather than specific nucleoside sequences [4].
In homeostatic systems, cell-based negative transcriptional feedback loop regulation is important for circadian expression of clock genes that are responsible for daily oscillations in physiology and behavior. In addition, posttranscriptional regulation via the 3′-UTR of genes appears to be important in the expression of circadian clock genes [6, 7]. HnRNP R, Q, and L reportedly bind to the 3′-UTR of serotonin N-acetyltransferase [arylalkylamine N-acetyltransferase (AANAT)], which is the key enzyme in melatonin synthesis and is regulated by the circadian system, resulting in rhythmic AANAT mRNA degradation [6]. LARK reportedly interacts with the 3′-UTR of the Per1 gene, regulating its expression in a posttranscriptional manner, recognizing the sl structure rather than the specific RNA sequence [7].
In the present study, we found that the hairpin sequence in the 3′-UTR of Per2 has the potential to form an sl RNA structure (hereafter mPer2 sl) after the gene is transcribed. In addition, we identified two mouse liver nuclear proteins, hnRNP H and C, which bind specifically to the 3′-UTR of Per2. We also describe the age-related expression profiles of these proteins.
Materials and methods
Construction of mPer2 sl-RNA probes
32P-labeled mPer2 sl-RNA was prepared by in vitro transcription for 2 h at 37 °C using a T7-MEGAshortscript high-yield kit (Ambion, Austin, TX, USA) with a template DNA fragment of mPer2 sl (39 bp). Template DNA fragments were generated using the sequence 5′-TAATACgACTCACTATAgg TACACTggCTTTTTTTgTTTTAggAAAAACAAAAAACAA-3′ (19 bp for the T7 promoter + 39 bp for mPer2), which corresponds to the sl region spanning nucleotides (nts) 4006–4043 of the mPer2 gene (Genbank accession number: AF036893).
Transcription using the mPer2 sl DNA fragments thus generated was carried out in a reaction mixture (final volume of 20 μL) composed of 2 μL of 10× transcription buffer provided in the Ambion kit, 2 μL each of 75 mM ATP, GTP, and CTP, 4 μL of α-[32P]UTP (800 Ci/mmol; GE Healthcare, Buckinghamshire, UK), 1 μg of mPer2 template DNA, and 2 μL of T7 MEGAshortscript enzyme mix. After 2 h incubation at 37 °C, the reaction mixture was added with 1 μl of RNase-free DNase solution, incubated for 15 min at 37 °C and subjected to electrophoresis using a 6 % polyacrylamide urea gel to remove template DNA fragments and unincorporated nucleotides. The gel was then exposed to an X-ray film for 10 s, and gel areas containing radioactivity were precisely located by matching with the autoradiogram and excised. The gel pieces recovered were then incubated in a probe elution buffer (Ambion) at 37 °C overnight. Non-radioactive mPer2 sl-RNA probes were prepared in a similar manner by using non-radioactive nucleosides in transcription reaction.
UV cross-linking and electrophoresis of RNA/protein complexes
Electrophoretic mobility shift assays (EMSAs) were carried out as previously described (Fig. S1) [4]. Nuclear extracts (NEs) were prepared from liver tissues of 6-month-old C57BL/6 mice (Charles River Laboratories, Yokohama, Japan). Animal care and use procedures were reviewed and approved by the committee for animal experimentation of the National Institute of Advanced Industrial Science and Technology (AIST) (Permission #36-07-009) and were performed in accordance with the institutional guidelines of the Committee for Animal Experimentation in the AIST. Some animal work was performed in accordance with the Hokkaido University Guidelines for the Care and Use of Laboratory Animals, under permission #14-0059 from the Hokkaido University Committee for Animal Experimentation.
Identification of liver nuclear proteins bound to the mPer2 sl-RNA probe
Identification of liver nuclear proteins bound to the mPer2 sl-RNA probe was carried out as previously described [4]. We showed a flowchart of the identification of mPer2 sl-RNA binding protein in supplemental Fig. 1. RNA electrophoretic mobility shift assays using a 32P-labeled Per2 RNA probe coupled with two-dimensional gel electrophoresis followed by MALDI-TOF/MS peptide mass fingerprint analysis was used to analyze these proteins.
Western blotting of liver nuclear proteins bound to the mPer2 sl-RNA probe
Liver NEs protein samples were prepared from C57BL/6J mice at various ages, as previously described [4]. Proteins were solubilized in 20 μL of SDS loading buffer. After adjusting the protein concentration using the BCA method, samples were subjected to 10 % SDS-PAGE. Western blotting analyses were carried out according to standard methods, using a horseradish peroxidase–conjugated anti-rabbit antibody for protein detection with the ECL assay system (GE Healthcare). Band intensity was measured using a Fujix Bio-imaging analyzer LAS 1000 v.3.4X software (Fujifilm, Tokyo, Japan). Mouse anti-hnRNP F/H (mouse monoclonal; 1G11; ImmuQuest, Cleveland, UK), anti-hnRNP C1/C2 (mouse monoclonal; 4F4; ImmuQuest), and anti-mouse HuR (mouse monoclonal; 19F12; Molecular Probes, Eugene, OR, USA) antibodies were used.
Two-dimensional gel electrophoresis (2DE) analysis of age-dependent expression of hnRNP C/C2 and hnRNP H1/H2 in the liver
Mice (C57BL/6J mice) were maintained for at least 1 week on a 12 h light/12 h dark (LD) cycle with lights on 0600–1800 hours. The sampling of mouse liver was performed during 0930–1200 hours. Liver NEs (300 μg each) prepared from 1-, 3-, 6-, 12-, 18-, 21-, and 24-month-old mice (n = 20 mice at each age) were subjected to isoelectric focusing (IEF) at pH 4–7 (acidic), pH 5–8 (neutral), and pH 6–11 (basic) using 24-cm Immobiline Dry Strips (Bio-Rad, Hercules, CA, USA), followed by SDS-PAGE on pre-cast 10–16 % gradient polyacrylamide gels (Bio-Rad). Each sampling was done once and each gel was used for analysis. The gels were then stained with colloidal Coomassie Brilliant Blue (CBB) and scanned using a GS-800 scanner (Bio-Rad). Protein spots were analyzed using PDQuest software (v.7.1, Bio-Rad). The gel images were normalized and the protein spots were excised, destained in 50 mM ammonium bicarbonate/50 % acetonitrile, and dried in a vacuum concentrator (Savant, Holbook, NY, USA). The dried gel pieces were then rehydrated with 5 μL of a solution of 20 ng/L trypsin in 10 mM ammonium bicarbonate and incubated at 30 °C overnight. Mass spectrometric analysis of the tryptic peptides was carried out as described previously [4]. All spots were subjected to matrix-assisted laser desorption/ionization-time-of-flight mass spectrometry (MALDI-TOF/MS) peptide mass fingerprint (PMF) analysis on an Axima® CFR MALDI-TOF mass spectrometer (Kratos, Manchester, UK/Shimazu Biotech, Kyoto, Japan). PMF data were searched against the National Center for Biotechnology Information (NCBI) database using the internet-available program Mascot (Matrix Science, London, UK). Each matched protein spot was assigned a unique sample spot protein (SSP) number prefixed by A (acidic), N (neutral), or B (basic) in the PDQuest software. Analysis of age-related expression of hnRNP C1/C2 and hnRNP H1/H2 was carried out using an age-related mouse liver nuclear protein database that we constructed (unpublished) [8, 9].
Results
RNA-EMSAs of mPer2 sl-RNA with mouse liver NE
We found potential hairpin sequences in the 3′-UTR of the Per2 gene (Fig. 1a, b). GENETYX-MAC analysis predicted that the secondary structure of this region would form a sl structure (Fig. 1c). The 3′-UTRs of the rat Per2 (rPer2) (Genbank accession number: NM031678) and human Per2 (hPer2) (Genbank accession number: NM022817) genes also contain potential hairpin sequences (rPer2 sl and hPer2 sl) between nts 3974 and 3994 in rPer2 (5′-TTCACTgCTTCTTTTgTTTTAgAAAAAAAAACAAAACAC-3′) and between nts 4110 and 4147 in hPer2 (5′-CATgTTgCTTTTTTTgTTTTAgAAAAAAAAACAACATA-3′) (Fig. 1a). The mPer2, rPer2, and hPer2 sl structures are located after a stop codon about 100 bp in length (the mPer2, rPer2, and hPer2 stop codons are located at nts 3916–3918, 3876–3878, and 4003–4005, respectively). The mPer2 sl showed 86.8 and 90.6 % homology to rPer2 sl and hPer2 sl, respectively.
RNA-EMSA of 32P-labeled mPer2 sl-RNA with mouse liver NE. a Purine–pyridine sequence in the 3′-UTR of the Per2 gene. b Location of the potential hairpin sequence in the 3′-UTR (starting at nucleotide 4006) of the mPer2 gene. c The potential 3′-UTR hairpin (starting at nucleotide 4006) in the mPer2 gene. d RNA-EMSA of the 32P-labeled mPer2 sl-RNA probe followed by UV cross-linking and RNase treatment. RNA-EMSA of 32P-labeled mPer2 sl-RNA with NE. Arrowheads indicate the mPer2 sl-RNA probe–protein complexes. Lanes 1–5 contain 32P-labeled mPer2 sl-RNA and RNase T1. Lane 1 no NE; lanes 2– 4 with liver NE. Asterisk indicates that band intensity did not increase with increasing amount of NE added. e Competitive RNA-EMSA of 32P-labeled mPer2 sl-RNA with NE. Lanes 1–5 contain 32P-labeled mPer2 sl-RNA and RNase T1. Lanes 2–4 contain unlabeled mPer2 sl-RNA and NE. Lane 5 contains no NE
As shown in Fig. 1b, a 32P-labeled mPer2 sl-RNA probe was prepared from the region spanning nts 4006 and 4043 of the mPer2 gene. RNase treatment of the 32P-labeled mPer2 sl-RNA probe cross-linked by UV treatment to mouse liver nuclear proteins revealed some bands corresponding to RNA probe/nuclear protein complexes (Fig. 1d). Of these, two bands were identified with shifted mobility and for which the intensity increased with increasing amounts of added NE (Fig. 1d, lanes 2–5). These two bands were observed at approximately 37 and 50 kDa (Fig. 1d, lanes 2–5). These shifted bands effectively competed with 10-, 50-, and 100-fold excess amounts of non-radiolabeled (cold) mPer2 sl-RNA probe (Fig. 1e, lanes 2–4), indicating that they were specifically generated by the 32P-labeled mPer2 sl-RNA probe.
Identification of liver nuclear proteins bound to the mPer2 sl-RNA probe
An amount of 32P-labeled mPer2 sl-RNA/protein complex sufficient for preparative solution-phase IEF was obtained by repeating the RNA EMSA procedures with the 32P-labeled mPer2 sl-RNA probe, the UV-irradiation and RNase digestion of the extracted 32P-labeled mPer2 sl-RNA probe/protein complex, SDS-PAGE separation, and subsequent excision of the radioactive gel area and extraction of the treated complex using electroelution, as previously described [4]. Analytical 2DE of the complex after concentration and autoradiography showed that preparative solution-phase IEF efficiently concentrated most of the 32P-labeled mPer2 sl-RNA/protein complex within the pI 4-5 zone, consistent with a previous report (Fig. S1) [4]. The 2DE and subsequent autoradiographic analyses of the pooled and concentrated solutions of the pI 3–4.6 and pI 4.6–5.4 zones identified distinct multiple radioactive gel spots, including a major spot between approximately 40 and 50 kDa that exhibited the most intense radioactivity (data not shown). MALDI-TOF/MS and PMF analyses of the proteins extracted from this major radioactive gel spot at approximately 40 kDa identified hnRNP C as the protein component, with a Mascot score of 270 (Expect: 8.3e-023; Sequence Coverage: 46 %) (Table 1, Fig. S3). Similar analyses of 32P-labeled mPer2 sl-RNA probe/nuclear protein complexes at approximately 50 kDa identified hnRNP H2 and H1, with Mascot significant scores of 171 (Expect: 6.6e-013; Sequence Coverage: 56 %) and 159 (Expect: 1e-011; Sequence Coverage: 65 %), respectively (Table 1, Fig. S4). No other proteins were identified by the MALDI-TOF/MS and PMF analyses (Table 1).
Age-related expression of mPer2 sl-RNA–binding proteins in the liver
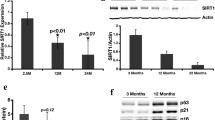
Using monoclonal anti-hnRNP F/H and anti-hnRNP C1/C2 antibodies, western blot analysis was carried out with liver NEs obtained from 1-, 3-, 8-, 16-, and 21-month-old C57BL/6 mice (Figs. 2a, 2S). HnRNP H and its variants are closely related to hnRNP F [10]. The major hnRNP H protein bands, at approximately 50 kDa, were identified according to a previous report [10]. The major hnRNP C1/C2 protein bands, at approximately 40 kDa, were also identified [11]. Expression of HnRNP F/H peaked at 3 months of age and remained stable thereafter (Fig. 2b). Expression of HnRNP C1/C2 peaked at 8 months of age (Fig. 2b). The expression of these proteins differed from that of both HuR (an AU-rich RNA domain binding protein that showed an age-stable expression pattern; Fig. 2a, b) and hnRNP A3, the expression of which has been shown to increase with age [4].
Age-related expression of mPer2 sl-RNA–binding proteins in the liver. a Western blotting analysis of age-related hnRNP H and hnRNP C expression using monoclonal antibodies and 10 μg of male mouse liver NE. Molecular weight is indicated on the right, and the age of the animals is shown at the top. b Graphical illustration of the patterns of hnRNP C1/C2 and hnRNP H/F expression as shown in (a). Protein expression is shown as the total mean intensity of all bands at each age. c 2DE analysis of age-related expression of hnRNP C1/C2 in the liver. Representative images of 2DE gels for 3-month-old mice. CBB stained spots are shown in the images from immobilized pH 4–7, 5–8, and 6–11 gradient IEF gels and 10–16 % gradient SDS-PAGE gels. Spots identified as hnRNP C1/C2 are shown with red circles. d Age-related expression profiles of spots identified as hnRNP C1/C2. e 2DE analysis of age-related expression of hnRNP H1/H2 in the liver. Spots identified as hnRNP H1/H2 are shown with a red circle. The location of each spot is shown with red circle in Figs. S6, S7 and S8. f Age-related expression profiles of spots identified as hnRNP H1/H2. Spot N5525 in Table 2 is not shown because the intensity of spot was very low. Mouse liver NEs were sampled between 0930 and 1200 hours (lights on between 0600 and 1800 hours) (color figure online)
2DE analysis of age-related expression hnRNP C1/C2 and hnRNP H1/H2 in the liver
To examine the age-related expression of hnRNP C1/C2 and hnRNP H1/H2 in detail, we analyzed the liver nuclear proteins of 1-, 3-, 6-, 12-, 18-, 21-, and 24-month-old mice using 2DE combined with MALDI-TOF/MS (Fig. 2c–f). All 2DE gel spots (total number of spots, 3113; number of single spots, 2557; number of mixed spots, 556) of proteins from 3-month-old mice were examined using MALDI-TOF/MS analysis. HnRNP C1/C2 was identified in 7 single spots and 3 mixed spots (Table 2; Figs. 2c, S6). Many spots were distributed between pI 5 and pI 6 and between 37 and 39 kDa. The age-related expression of 4 single spots (A2319, A2427, A2432, and A2433) is illustrated in Fig. 2d. Spots B6819, B8328, and B9112 spots in Table 2 were disregarded due to their very different molecular weight from the band corresponding to the RNA/nuclear protein complex (Fig. 1e) or because the intensity of the spot was very low. Spot A2432 exhibited the highest expression intensity, followed by spots A2319, A2433, and A2427, in that order. The expression of these proteins tended to decline after 18 months of age (Fig. 2d).
HnRNP H1 was identified in 3 single spots and 4 mixed spots, whereas hnRNP H2 was identified in 1 single spot and 4 mixed spots (Table 2; Figs. 2e, S7, S8). The expression of both hnRNP H1 and H2 was relatively lower than that of both hnRNP C1 and C2 (Fig. 2d, f). The age-related expression of 4 single spots (A1520, A4520, A6405, and A514) is illustrated in Fig. 2f. The expression patterns of these proteins were unique. Spot N6405, identified as hnRNP H1, peaked at 12 months of age. After 21 months, the expression of spot N4520 decreased. The expression of spot N6514 remained constant. The expression of spot A1520, identified as hnRNP H2, decreased after 3 months of age.
Discussion
In the present study, we found that the proteins hnRNP C and hnRNP H bind to the 3′-UTR region of the Per2 gene, which has the potential to form distinct sl RNA structures after transcription. Both hnRNP C and hnRNP H are members of the hnRNP family, which is composed of more than 20 proteins, including hnRNP A1, A2, and A3 [10, 11]. As previously reported, we found that hnRNP A3 binds to the AIE in 3′-UTR of the Factor IX gene and recognizes the overall three-dimensional sl structures of AIE RNAs rather than specific nucleoside sequences [4]. In contrast to hnRNP A3, neither hnRNP C nor hnRNP H exhibited age-related differences in expression in the liver in this study. These results indicate that hnRNP C and hnRNP H play different roles than hnRNP A3 in regulating gene expression in the liver.
Similar to other hnRNP family proteins, various protein modifications in addition to alternative splicings might give rise to many hnRNP C and hnRNP H proteins spots detected in 2DE [12–18]. Supplementary Fig. 3 show 2DE analysis of the mouse liver NEs along the age axis and spots originated from hnRNP A3 protein. HnRNP A3 is regulated by various types of modifications including phosphorylation [4, 17, 18], methylation [19, 20] and sumoylation [21]. Our data show HnRNP A3 proteins received modification including the phosphorylation at its Ser359 and the level of this phosphorylation increased with age. In contrast, the number of spots of hnRNP C and hnRNP H proteins were stable along the age axis (Fig. 2). Unfortunately, we could not detect any phosphorylation sites of HnRNP C or hnRNP H in the present experiment by MS analysis. Further studies are needed to analyze the modification sites of HnRNP C and hnRNP H proteins.
The present study showed that there are a number of isoforms of hnRNP C and hnRNP H, and that these isoforms exhibit complex age-related expression patterns. Our data suggest that many protein isoforms are regulated differentially during aging and that many mechanisms function in the liver to regulate protein expression during aging. HnRNPs also reportedly play important roles in the posttranslational regulation of circadian clock gene expression. In the cryptochrome (Cry) gene, hnRNP D binds to the middle part of the 3′-UTR, which contains destabilizing cis-acting elements and contributes to Cry1 mRNA turnover and modulation of the circadian rhythm [22]. AANAT expression is also post-transcriptionally regulated by hnRNP R, Q, and L via the 3′-UTR of the gene [23]. The expression of hnRNP U in the mouse SCN (suprachiasmatic nucleus), in which the master circadian pacemaker is located, shows circadian rhythm [24]. Using the full-length 3′-UTR, Woo et al. showed that hnRNP I in the cytoplasm of CHO-K1 cells binds to the CU-rich portion of 3′-UTR of the mouse Per2 gene and that many RNA-binding proteins interact with Per2 mRNA, including two proteins of approximately 40 and 50 kDa [25]. These two proteins may correspond to hnRNP C and hnRNP H, which were identified in the present study (Fig. 1d). Post-transcriptional regulation via the 3′-UTR of genes appears to be an important system for maintaining circadian clock gene expression patterns in a number of species. CCTR and CHLAMY1 in Gonyaulax polyedra and Chlamydomonas, AtGRP7 in Arabidopsis, and LARK in Drosophila, are all reportedly involved in regulating the circadian rhythm of translation [26–29]. After transcription, Per1 forms a sl RNA structure in its 3′-UTR. In mammals, the RNA-binding protein LARK interacts with the 3′-UTR of Per1 and post-transcriptionally regulates Per1 expression [7].
In this study, we could not detect the spot arising from the PER2 protein in our 2DE/MALDI-TOF/MS analyses of age-related mouse liver nuclear protein expression. To obtain a better understanding of the relationship between age and PER2 expression, it is important that future studies identify which isoform of hnRNP C1/C2 or H1/H2 binds to the Per2 sl structure.
References
Kurachi S, Deyashiki Y, Takeshita J, Kurachi K (1999) Genetic mechanisms of age regulation of human blood coagulation factor IX. Science 285:739–743
Zhang K, Kurachi S, Kurachi K (2002) Genetic mechanisms of age regulation of protein c and blood coagulation. J Biol Chem 277:4532–4540
Kurachi S, Huo JS, Ameri A, Zhang K, Yoshizawa AC, Kurachi K (2009) An age-related homeostasis mechanism is essential for spontaneous amelioration of hemophilia B Leyden. Proc Natl Acad Sci USA 106:7921–7926
Hamada T, Kurachi S, Kurachi K (2010) HnRNP A3 is the liver nuclear protein binding to the age-related increase element derived RNA of the factor IX gene. PLoS ONE 5(9):e12971
Yoshitake S, Schach BG, Foster DC, Davie EW, Kurachi K (1985) Nucleotide sequence of the gene for human factor IX (antihemophilic factor B). Biochemistry 24:3736–3750
Kim TD, Woo KC, Cho S, Ha DC, Jang SK, Kim KT (2007) Rhythmic control of AANAT translation by hnRNP Q in circadian melatonin production. Genes Dev 21:797–810
Kojima S, Matsumoto K, Hirose M, Shimada M, Nagano M, Shigeyoshi Y, Hoshino S, Ui-Tei K, Saigo K, Green CB, Sakaki Y, Tei H (2007) LARK activates posttranscriptional expression of an essential mammalian clock protein, PERIOD1. Proc Natl Acad Sci USA 104:1859–1864
Kurachi K, Kurachi S, Hamada T, Suenaga E, Yoshizawa AC (2008) Aging and gerontological diseases in relation to age-related homeostasis and age dimension technology. Nihon Ronen Igakkai Zasshi 45:126–131
Kurachi K, Kurachi S, Hamada T, Suenaga E, Bolotova T, Solovieva E (2008) Age-related homeostasis and hemostaic system. In: Tanaka K, Davie EW (eds) Recent advances in thrombosis and hemostasis, pp 427–438
Honoré B, Rasmussen HH, Vorum H, Dejgaard K, Liu X, Gromov P, Madsen P, Gesser B, Tommerup N, Celis JE (1995) Heterogeneous nuclear ribonucleoproteins H, H′, and F are members of a ubiquitously expressed subfamily of related but distinct proteins encoded by genes mapping to different chromosomes. J Biol Chem 270:28780–28789
Shetty S (2005) Regulation of urokinase receptor mRNA stability by hnRNP C in lung epithelial cells. Mol Cell Biochem 272:107–118
Weighardt F, Biamonti G, Riva S (1995) Nucleo-cytoplasmic distribution of human hnRNP proteins: a search for the targeting domains in hnRNP A1. J Cell Sci 108:545–555
Ostareck DH, Ostareck-Lederer A, Wilm M, Thiele BJ, Mann M, Hentze MW (1997) mRNA silencing in erythroid differentiation: hnRNP K and hnRNP E1 regulate 15-lipoxygenase translation from the 3′ end. Cell 89:597–606
Ostareck-Lederer A, Ostareck DH, Cans C, Neubauer G, Bomsztyk K, Superti-Furga G, Hentze MW (2002) c-Src-mediated phosphorylation of hnRNP K drives translational activation of specifically silenced mRNAs. Mol Cell Biol 22:4535–4543
Buxadé M, Parra JL, Rousseau S, Shpiro N, Marquez R, Morrice N, Bain J, Espel E, Proud CG (2005) The Mnks are novel components in the control of TNF alpha biosynthesis and phosphorylate and regulate hnRNP A1. Immunity 23:177–189
Hüttelmaier S, Zenklusen D, Lederer M, Dictenberg J, Lorenz M, Meng X, Bassell GJ, Condeelis J, Singer RH (2005) Spatial regulation of beta-actin translation by Src-dependent phosphorylation of ZBP1. Nature 438:512–515
Yamaoka K, Imajoh-Ohmi S, Fukuda H, Akita Y, Kurosawa K, Yamamoto Y, Sanai Y (2006) Identification of phosphoproteins associated with maintenance of transformed state in temperature-sensitive Rous sarcoma-virus infected cells by proteomic analysis. Biochem Biophys Res Commun 345:1240–1246
Villén J, Beausoleil SA, Gerber SA, Gygi SP (2007) Large-scale phosphorylation analysis of mouse liver. Proc Natl Acad Sci USA 104:1488–1493
Liu Q, Dreyfuss G (1995) In vivo and in vitro arginine methylation of RNA-binding proteins. Mol Cell Biol 15:2800–2808
Bedford MT, Richard S (2005) Arginine methylation an emerging regulator of protein function. Mol Cell 18:263–272
Li T, Evdokimov E, Shen RF, Chao CC, Tekle E, Wang T, Stadtman ER, Yang DC, Chock P (2004) Sumoylation of heterogeneous nuclear ribonucleoproteins, zinc finger proteins, and nuclear pore complex proteins: a proteomic analysis. Proc Natl Acad Sci USA 101:8551–8556
Woo KC, Ha DC, Lee KH, Kim DY, Kim TD, Kim KT (2010) Circadian amplitude of cryptochrome 1 is modulated by mRNA stability regulation via cytoplasmic hnRNP D oscillation. Mol Cell Biol 30:197–205
Kim TD, Kim JS, Kim JH, Yung J, Chae HD, Woo KC, Jang SK, Koh DS, Kim KT (2005) Rhythmic serotonin N-acetyltransferase mRNA degradation is essential for the maintenance of its circadian oscillation. Mol Cell Biol 25:3232–3246
Tamaru T, Isojima Y, Nagai K, Takamatsu KC (2003) Circadian expression of hnRNP U, a nuclear multi-potent regulatory protein, in the murine suprachiasmatic nucleus. Neurosci Lett 341:111–114
Woo KC, Kim TD, Lee KH, Kim DY, Kim W, Lee KY, Kim KT (2009) Mouse period 2 mRNA circadian oscillation is modulated by PTB-mediated rhythmic mRNA degradation. Nucleic Acids Res 37:26–37
Ziemienowicz A, Haasen D, Staiger D, Merkle T (2003) Arabidopsis transportin1 is the nuclear import receptor for the circadian clock-regulated RNA-binding protein AtGRP7. Plant Mol Biol 53:201–212
Mittag M, Lee DH, Hastings JW (1994) Circadian expression of the luciferin-binding protein correlates with the binding of a protein to the 3′ untranslated region of its mRNA. Proc Natl Acad Sci USA 91:5257–5261
Mittag M (1996) Conserved circadian elements in phylogenetically diverse algae. Proc Natl Acad Sci USA 93:14401–14404
Heintzen C, Nater M, Apel K, Staiger D (1997) AtGRP7, a nuclear RNA-binding protein as a component of a circadian-regulated negative feedback loop in Arabidopsis thaliana. Proc Natl Acad Sci USA 94:8515–8520
Acknowledgments
This research was supported in part by an internal research fund from AIST, JSPS KAKENHI (Grant Number 24621001), and by a research fund from The Suhara Memorial Foundation. We thank Dr. Kenneth Sutherland for comments on drafts of this manuscript.
Conflict of interest
None of the authors have any conflict of interest.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
12576_2015_373_MOESM1_ESM.pdf
Supplemental Fig ure 1. Flowchart of the identification of mPer2 sl-RNA binding protein. For identification of the mPer2 sl -RNA binding protein(s), preparative RNA EMSA using 20 μg NEs were repeated in a similar condition as for analytical scale RNA EMSA [4]. UV cross -linking and electrophoresis of RNA/protein complexes: A 32P-labeled mPer2 sl-RNA probe was incubated on ice for 50 min with 20 μg of NEs in the presence of 0.2 μg of yeast RNA and 15 U RNase inhibitor in a final volume of 30 μl of EMSA binding buffer (10 mM Tris–HCl, 50 mM NaCl, 5 % glycerol, 1 mM DTT and 1 mM EDTA) (Fig. 1D, lane 5). The reaction mixture was then exposed to UV light for 30 min on ice (UV Stratalinker 1800, Stratagene), followed by incubation with 1.5 μl RNase T1 (5 U/μl) and incubated for 20 min at 25 °C. The mixture was added with 8 μl SDS loading buffer, boiled for 2 min and subjected to a preparative SDS-PAGE using NuPAGE® Novex Bis–Tris 4–12 % ZOOM® Gel (Invitrogen, Carlsbad, CA, USA). The gel was exposed to an X-ray film at 4 °C for 1 day. Radioactive areas of the gel containing the nuclear protein/32P-labeled mPer2 sl-RNA complex observed in Fig. 1D were excised (yellow square area and red square area indicated by arrow in Fig. S1, step1) and subjected to electro-elution using a Centriruter microelectroeluter (Amicon, Beverly, MA, USA). Identification of liver nuclear proteins bound to the m Per2 sl-RNA probe: 32P-labeled mPer2 sl-RNA probe/nuclear protein complex recovered from the gel pieces were concentrated with a Centricon apparatus (Millipore, Billerica, MA, USA). 32P-labeled mPer2 sl/protein complex preparations obtained from repeated procedures were pooled, and subjected to preparative solution phase 1st isoelectric focusing (IEF) by using a ZOOM® IEF Fractionator (Invitrogen) which can divide the solution at each pI (Fig. S1, step2). The sample solution with radio-activity (pI 3-5.4) separated in the ZOOM Disks apparatus was then recovered, concentrated by a Centricon apparatus (Amicon) and subjected to two-dimensional electrophoresis (2DE) using an IPGRunner system (Invitrogen) (Fig. S1, step 3) [4]. In step 3, for solid phase IEF (the first dimension of separation), a pre-casted ZOOM strip of an appropriate pH range gradient (Invitrogen), which was selected according to the solution phase IEF, was used. After completing IEF, the gel was subjected to SDS-PAGE (2nd dimension separation) using ZOOM gels (Invitrogen), followed by silver staining and exposure to X-ray film at 4 °C for 1–6 days depending on the radioactivity level [4]. The gel area precisely matched with the radioactive area of autoradiogram was excised, in-gel trypsin digested and subjected to peptide mass fingerprint (PMF) analysis using a Axima® CFR matrix-assisted laser desorption/ionization-time-of-flight mass-spectrophtometer (Axima® CFR MALDI-TOF/MS) (Kratos, Manchester, UK/Shimazu Biotech, Kyoto, Japan). A database search analyses were performed with the NCBI database using the internet-available program Mascot (Matrix Science) (Fig. S1, step 4). (pdf 68 kb)
12576_2015_373_MOESM2_ESM.pdf
Supplemental Figure 2. Western blotting of HnRNP C, HnRNP H and HuR. Western blot analysis of age-related HnRNP C, HnRNP H and HuR proteins levels in the mouse liver. Full size images of Fig. 2A are shown. (pdf 43 kb)
12576_2015_373_MOESM3_ESM.pdf
Supplemental Figure 3 Amino acid sequences of mouse hnRNP C in complex with mPer2 sl- RNA.Amino acid sequences of mouse liver hnRNP C bound with mPer2 sl- RNA identified by MALDI-TOF/MS and subsequent Mascot search analyses are shown in Red letters of its known complete sequence.(pdf 34 kb)
12576_2015_373_MOESM4_ESM.pdf
Supplemental Fig ure 4. Amino acid sequences of mouse hnRNP H1 and H2 in complex with mPer2 sl- RNA.Amino acid sequences of mouse liver hnRNP H1 and H2 bound with mPer2 sl- RNA identified by MALDI-TOF/MS and subsequent Mascot search analyses are shown in red letters of its known complete sequence. (pdf 36 kb)
12576_2015_373_MOESM5_ESM.pdf
Supplemental Fig ure 5. 2DE analysis of the mouse liver NEs along the age axis and spots originated from hnRNP A3 protein.The images of 2DE gel electrophoresis using mice liver nuclear protein at 6 month age (A) and 21 month age (B) are shown. The spots of hnRNP A3 protein are detected within the square window in (A) and (B), and magnified in (C). Mouse liver NEs (300 μg) were subjected to IEF using a 24 cm immobiline DryStrip (pH 6-11), followed by SDS-PAGE using pre-casted 10–16 % gradient polyacrylamide gels. Gels were then stained with Colloidal Coomassie Brilliant Blue (CBB) and scanned by GS-800 scanner (BIO-RAD). (pdf 56 kb)
12576_2015_373_MOESM6_ESM.pdf
Supplemental Fig ure 5 C. Age-related expression change of spots originated from hnRNP A3 protein on the 2DE gel. Protein spots were analyzed with a PDQuest software (BIO-RAD, v.7.1), and excised, destained in 50 mM ammonium bicarbonate/50 % acetonitrile and dried in a vacuum concentrator (Savant, Holbook, NY, USA). Dried gel pieces were then rehydrated with 5 μl of 20 ng/L trypsin in 10 mM ammonium bicarbonate, and incubated at 30 °C overnight. The peptide containing fractions collected and dried were mixed with CHCA matrix before applying to an Axima® CFR MALDI-TOF/MS (Shimazu Biotech, Kyoto, Japan) or an AXIMA® MALDI-TOF2 (Shimadzu Biotech). PMF analysis was performed with a MASCOT server version 2.1 (Matrix Science, London, UK) taking UniProt or NCBI databases as a reference database. Age-related expression analysis of hnRNP A3 proteins was carried out using an age-related mouse liver nuclear protein database which we made (data not published) [8, 9]. In the present study, we found 20 spots of hnRNP A3 protein in this area at 3 and 6 months old samples. MS/MS analysis only detected mass-spectra m/z 1990.8, indicating that Ser359 in the peptide fragment (aa 356 through 377, SSGSPYGGGYGSGGGGSGGYGSR) [4] as the post-translational modification site of hnRNP A3. Four of the 14 hnRNP A3 single protein spots contained hnRNP A3 with its Ser359 phosphorylated, while others contained hnRNP A3 with its Ser359 unphosphorylated (3 and 6 months old samples). The single protein spots contain only hnRNP A3 protein and the mixture spots contain with the other proteins identified besides hnRNP A3 protein. Single and Mixture spots of Ser359 phoshorylated hnRNP A3 protein are marked by red and blue circles respectively. Unphosphorylated hnRNP A3 protein spots are marked by yellow and purple circles which mean single and mixture spots, respectively. At 3 and 6 months old, the number and expression pattern of all spots of hnRNP A3 protein were the same (4 single spots of Ser359 phosphorylated hnRNP A3 protein were widely distributed along the PI axis). Interestingly, the numbers of spots of Ser359 phosphorylated hnRNP A3 protein increased in 18 months (data not shown) and 21 months old mice. Two spots of Ser359 phoshorylated hnRNP A3 protein and 1 spot of unphosphorylated hnRNP A3 protein were newly shown at the area close to Spots 3 and 4. a, Ornithine carbamoyltransferase, and mitochondrial precursor protein; b, Fructose-bisphosphate aldolase B protein; c, Fructose-bisphosphate aldolase B protein; d, WD repeat-containing protein 57 protein; e, Cytochrome b-c1 complex subunit 2, and mitochondrial precursor protein. Symbols a ~ e show proteins as position marker of the spot in 2DE gels at 3, 6, and 21 months of age.(pdf 27 kb)
12576_2015_373_MOESM7_ESM.pdf
Supplemental Fig ure 6. Each spot of hnRNP C1/C2 proteins in the image of 2DE gel electrophoresis observed in Fig. 2C. (pdf 135 kb)
12576_2015_373_MOESM8_ESM.pdf
Supplemental Fig ure 7. Each spot of hnRNP H1 proteins in the image of 2DE gel electrophoresis observed in Fig. 2E. (pdf 96 kb)
12576_2015_373_MOESM9_ESM.pdf
Supplemental Figure 8. Each spot of hnRNP H2 proteins in the image of 2DE gel electrophoresis observed in Fig. 2E. (pdf 84 kb)
About this article
Cite this article
Hamada, T., Miyakawa, K., Kushige, H. et al. Age-related expression analysis of mouse liver nuclear protein binding to 3′-untranslated region of Period2 gene. J Physiol Sci 65, 349–357 (2015). https://doi.org/10.1007/s12576-015-0373-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12576-015-0373-8