Abstract
The development and cultivation of genetically modified crops is still increasing globally. Food and feed imports from outside the European Union (EU) will subsequently require more effort from the responsible authorities in monitoring the compliance with effective labelling regulations. The aim of this study was the development of the GMOfinder, a database for collection and interpretation of information related to the screening for genetically modified organisms (GMOs). Different genetic elements (e.g. promoters, terminators, structural genes) are artificially introduced into plants to establish new genetic modifications. The introduced elements may vary between GMO events, depending on the intended trait(s). Screening for such inserted elements with (real-time) polymerase chain reaction is a common first step to analyse samples for the presence of any genetical modification. From the pattern of detectable and nondetectable elements, valuable conclusions about the identity of putative present GMO event(s) can be drawn with the GMOfinder. Information about selected genetic elements from the literature, applications for authorisation and other (web) sources were systematically integrated in a tabular matrix format. Special care was taken to additionally record the sources of the information, thus facilitating evaluation of screening results, and tracing of possible errors in the matrix. The GMOfinder accesses data from the element matrix with implemented algorithms and facilitates to interpret the outcome of screenings. Such a preselection helps to systematically narrow down the candidates for subsequent identification reactions. Optional display of events with potentially masked elements completes the included features.





Similar content being viewed by others
Abbreviations
- BVL:
-
German Federal Office for Consumer Protection and Food Safety
- EFSA:
-
European Food Safety Authority
- EURL-GMFF:
-
European Reference Laboratory for GM Food and Feed
- GMO:
-
Genetically modified organism
- LGL:
-
Bavarian Health and Food Safety Authority
- QM:
-
Quality management
- UI:
-
Unique identifier
References
AGBIOS/CERA, GM crop database. http://cera-gmc.org/index.php?action=gm_crop_database. Accessed 23.06.2010
BATS (2003) Genetically modified (GM) crops: molecular and regulatory details - Version 2. Basel
Chaouchi M, Chupeau G, Berard A et al (2008) J Agric Food Chem 56:11596
Congen (unpublished) Detection method for pFMV; proposed for official German method collection ‘Amtliche Sammlung von Untersuchungsverfahren nach § 64 LFGB’
Convention on Biological Diversity (CBD) (2008) UNEP/CBD/BS/COP-MOP/4/INF/2/Add.1
CropScience-BioScience B (2006) Grain testing method for detection of rice GM events containing P35S::bar sequences using RT-PCR protocols PGS0494 and PGS0476. Protocol, Bayer Crop Science, Gent, Belgium
EFSA (2011) GMO requests and mandates. http://registerofquestions.efsa.europa.eu/roqFrontend/questionsListLoader?panel=GMO. Accessed 03.05.2011
Eurofins GeneScan (unpublished) Detection method for pFMV; proposed for official German method collection ‘Amtliche Sammlung von Untersuchungsverfahren nach § 64 LFGB’
Grohmann L, Brünen-Nieweler C, Nemeth A, Waiblinger HU (2009) J Agric Food Chem 57:8913
Hamels S, Glouden T, Gillard K et al (2009) Eur Food Res Technol 228:531
Heinze P (2008) J Verbr Lebensm 3:43
Holst-Jensen A (2009) Biotechnol Adv 27:1071
Holst-Jensen A, Bertheau Y, Loose MD et al (2012) Biotechnol Adv. doi:10.1016/j.biotechadv.2012.01.024
James C (2011) ISAAA Brief 42
Japanese Ministry of the Environment (2010) Japan Biosafety Clearing House. http://www.bch.biodic.go.jp/english/e_index.html, accessed 17.02.2010
Kuribara H, Shindo Y, Matsuoka T et al (2002) J AOAC Int 85:1077
Leimanis S, Hamels S, Nazé F et al (2008) Eur Food Res Technol 227:1621
Mano J, Shigemitsu N, Futo S et al (2009) J Agric Food Chem 57:26
OECD (2009) BioTrack product database. http://www2.oecd.org/biotech/byorganism.aspx. Accessed 21.07.2009
Official German method collection ‘Amtliche Sammlung von Untersuchungsverfahren nach § 64 LFGB’ (2008) Untersuchung von Lebensmitteln - Nachweis von bestimmten, häufig in gentechnisch veränderten Organismen (GVO) verwendeten DNA-Sequenzen aus dem Blumenkohlmosaikvirus (CaMV 35S-Promotor, P35S) sowie aus Agrobacterium tumefaciens (T-nos) in Lebensmitteln: Screening-Verfahren, vol 00.00-122
Official German method collection ‘Amtliche Sammlung von Untersuchungsverfahren nach § 64 LFGB’ (2008) Untersuchung von Lebensmitteln - Nachweis der CTP2-CP4-EPSPS-Gensequenz zum Screening auf Bestandteile aus gentechnisch veränderten Organismen (GVO) in Lebensmitteln; Konstrukt-spezifisches Verfahren, vol L 00.00–125
Official German method collection ‘Amtliche Sammlung von Untersuchungsverfahren nach § 64 LFGB’ (2008) Untersuchung von Lebensmitteln - Nachweis einer bestimmten, häufig in gentechnisch veränderten Organismen (GVO) verwendeten DNA-Sequenz aus dem bar-Gen von Streptomyces hygroscopicus in Lebensmitteln: Screening-Verfahren, vol 00.00-122
Official German method collection ‘Unterausschuss Methodenentwicklung der Bund/Länder-Arbeitsgemeinschaft Gentechnik (LAG)’ (2006) Real-time PCR zur quantitativen Bestimmung gentechnisch veränderter Rapslinien mit dem 35S/pat-Genkonstrukt, vol 03/2006
Official German method collection ‘Unterausschuss Methodenentwicklung der Bund/Länder-Arbeitsgemeinschaft Gentechnik (LAG)’ (2006) Konzept zur Untersuchung von Saatgut auf Anteile gentechnisch veränderter Pflanzen, vol 03/2006
Querci M, Van den Bulcke M, Zel J, Van den Eede G, Broll H (2010) Anal Bioanal Chem 396
Reiting R (2010) J Verbr Lebensm 5:377
Reiting R (unpublished) Unpublished method for: real-time PCR-detection of pTA29/barnase construct (29bar-Tm). LHL Hessen, Germany
Reiting R (unpublished) Real-time PCR-detection of pSSUAra/bar construct (Pbar-FP2/Tq-Pbar; modification of the PCR method of the German network LAG). LHL Hessen, Germany
Remacle J, Bertheau Y (2001) New technology in food science facing the multiplicity of new released GMO (GMOchips). University of Namur. http://www.fundp.ac.be/en/research/projects/page_view/01275404
Stein AJ, Rodríguez-Cerezo E (2009) The global pipeline of new GM crops: implications of asynchronous approval for international trade. JRC Sci Tech Rep
The Food and Environment Research Agency (FERA) (2010) GM Test Matrices—GM lines test elements search (v_web2). http://www.gm-inspectorate.gov.uk/seedAuditProgramme/GMTestMatrices.cfm. Accessed 04.03.2010
Van den Bulcke M, Lievens A, Barbau-Piednoir E et al (2010) Anal Bioanal Chem 396:2113
Waiblinger H-U, Boernsen B, Pietsch K (2008) DLR 104:261
Waiblinger HU, Grohmann L, Mankertz J, Engelbert D, Pietsch K (2010) Anal Bioanal Chem 396:2065
Working Group Biochemical and Molecular Biological Analysis (2011) Screening table for the detection of authorised and unauthorised genetically modified plants. http://www.gdch.de/strukturen/fg/lm/ag/bioanal/gvonachweis.xls. Accessed 24.03.2011
Zeitler R (2003) Standard operating procedure (SOP): ‘Qualitativer Nachweis von DNA-Sequenzen mit Hilfe der TaqMan-PCR’. Bavarian Environment Agency (LfU), Augsburg, Germany
Zeitler R, Pietsch K, Waiblinger H-U (2002) Eur Food Res Technol 214:346
Acknowledgements
The presented work was funded by the Bavarian State Ministry of the Environment and Public Health (UGV04090803099). We thank Krimhilde Posthoff and Patrick Guertler for testing the database and for useful discussions. Andrea Harwardt, Claudia Bujotzek, Roswitha Dorfner, Ulrike Mulats, Melina Mehmedovic, and Manuela Hillen provided excellent technical assistance.
Conflict of Interest
None
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Fig. S6
Query algorithms in the GMOfinder. Schematic overview and partial screenshots of the dataset selection procedure in the GMOfinder. The user decides on the form by adjusting option groups; the value from the option group is assigned via visual basic for applications (VBA) to a global variable; a public function is set via VBA according to the global variable; the public function is then used directly as query parameter in a query (JPEG 580 kb)
Fig. S7
Query parameters for the species selection. Schematic overview of the species selection procedure in the GMOfinder. The user enters experimental screening results by adjusting the species option groups (e.g. ‘Maize’) on the form; the value from the option group (e.g. ‘1’ for ‘detectable’) is assigned via VBA to a global variable (e.g. ‘MaisVorh’: ‘-1’ for ‘detectable’); a public function (e.g. ‘SpeziesMais()’) is set via VBA according to the global variable (e.g. ‘Mais’ for ‘detectable’); the public function is then used directly as parameter in the query. Query parameters: SpeziesMais() Or SpeziesSoja()… Or SpeziesTorenie(). Each of the parameters is either the German name of the species (e.g. ‘Mais’) or an empty string (‘“”’). Selection of ‘detectable’ or ‘not tested’ yields the corresponding German name, selection of ‘not detectable’ or ‘exclude’ yields the empty string as query parameter for the corresponding species (JPEG 166 kb)
Fig. S8
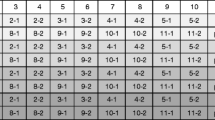
Query parameters for the selection of genetic elements/constructs. Schematic overview of the genetic elements/constructs selection procedure in the GMOfinder. The user enters experimental screening results by adjusting the element/construct option groups (e.g. ‘p35S (a)’); the value from the option group (e.g. ‘1’ for ‘detectable’) is assigned via VBA to a global variable (e.g. ‘aVorh’: ‘1’ for ‘detectable’); two public functions [e.g. ‘Eamin()’ and ‘Eamax()’] are set via VBA according to the global variable (e.g. ‘0’ and ‘9’ for ‘detectable’); the public functions are then used as limiting parameters in the query. Query parameters: Between Exxxmin() And Exxxmax(). Each of the parameters Exxxmin and Exxxmax is -9, 9, or 0. The database value for the given element/construct is for each GMO an integer value between -9 and 9, or 0 (compare table in Fig. 2). The selection of the option with/without masking affects the public function Exxxmin() that is set to -9 independently of the selection in the option group (JPEG 194 kb)
Fig. S9
Query parameters for the selection of empty datasets. Schematic overview of the selection procedure of empty datasets in the GMOfinder. The user decides on the inclusion/exclusion of empty datasets by adjusting the corresponding option group; the value from the option group (e.g. ‘1’ for ‘exclude’) is assigned via VBA to the global variable ‘leereDS’: ‘1’ for ‘exclude’; the public function ‘leerDS()’ is set via VBA according to the global variable (e.g. ‘“”’ for ‘exclude’); the public function is then used as comparison parameter in the query. Query parameters: Abs([a])+Abs([b])+…+Abs([m])>0 Or leerDS(). The database values for all elements/constructs are integer values between −9 and 9 or 0. Datasets with only 0 as value and accordingly 0 as sum of the absolute values are excluded by the presented query if the option ‘exclude’ is chosen (JPEG 137 kb)
Rights and permissions
About this article
Cite this article
Gerdes, L., Busch, U. & Pecoraro, S. GMOfinder—A GMO Screening Database. Food Anal. Methods 5, 1368–1376 (2012). https://doi.org/10.1007/s12161-012-9378-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12161-012-9378-6