Abstract
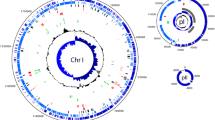
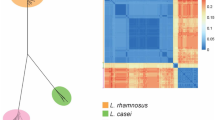
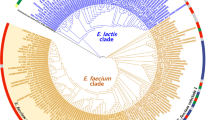
Lactobacillus casei, as a probiotic, has to survive the industrial production processes and endure the harsh physical–chemical environment of the human gastrointestinal tract. To explore genetic variability supporting improved adaptation of these bacteria to their environments, we performed a comparative genome analysis of L. casei. A total of 806 novel regions ≥500 bp were also found, as compared to the 12 recently sequenced L. casei strains, constituting 2,976,837 bp of genomic DNA not present in the seven reference complete genomes of L. casei. The widespread distribution of phage-related proteins among these novel regions suggests the transduction could be an important mechanism for genomic evolution in this species. The core- and accessory-based MP trees of L. casei had identical topology compared with MLST-based MP tree. The orthologues and phylogeny between L. casei and 35 human gut bacterial strains was analyzed; and their shared genes were involved in coenzyme transport and metabolism, which contributed perhaps to the adaptation of the human gastrointestinal tract for L. casei. These findings will be certainly helpful for the design of physiological studies which in turn will lead to a better understanding of genomic diversification of L. casei for niche expansion.


Similar content being viewed by others
References
de Vrese M, Schrezenmeir J (2008) Probiotics, prebiotics, and synbiotics. Adv Biochem Eng Biotechnol 111:1–66. doi:10.1007/10_2008_097
Corcoran BM, Stanton C, Fitzgerald G, Ross RP (2008) Life under stress: the probiotic stress response and how it may be manipulated. Curr Pharm Des 14:1382–1399. doi:10.2174/138161208784480225
Santivarangkna C, Kulozik U, Foerst P (2008) Inactivation mechanisms of lactic acid starter cultures preserved by drying processes. J Appl Microbiol 105:1–13. doi:10.1111/j.1365-2672.2008.03744.x
Kandler OWN (1986) Genus Lactobacillus. In: Sneath PHA, Mair NS, Sharpe ME, Holt JG (eds) Bergey’s manual of systematic bacteriology, 9th edn. Williams and Wilkins, Baltimore, pp 1063–1065
Makarova K, Slesarev A, Wolf Y, Sorokin A, Mirkin B, Koonin E, Pavlov A, Pavlova N, Karamychev V, Polouchine N, Shakhova V, Grigoriev I, Lou Y, Rohksar D, Lucas S, Huang K, Goodstein DM, Hawkins T, Plengvidhya V, Welker D, Hughes J, Goh Y, Benson A, Baldwin K, Lee JH, Diaz-Muniz I, Dosti B, Smeianov V, Wechter W, Barabote R, Lorca G, Altermann E, Barrangou R, Ganesan B, Xie Y, Rawsthorne H, Tamir D, Parker C, Breidt F, Broadbent J, Hutkins R, O’Sullivan D, Steele J, Unlu G, Saier M, Klaenhammer T, Richardson P, Kozyavkin S, Weimer B, Mills D (2006) Comparative genomics of the lactic acid bacteria. Proc Natl Acad Sci U S A 103:15611–15616. doi:10.1073/pnas.0607117103
Broadbent JR, Neeno-Eckwall EC, Stahl B, Tandee K, Cai H, Morovic W, Horvath P, Heidenreich J, Perna NT, Barrangou R, Steele JL (2012) Analysis of the Lactobacillus casei supragenome and its influence in species evolution and lifestyle adaptation. BMC Genom 13:533. doi:10.1186/1471-2164-13-533
Cai H, Thompson R, Budinich MF, Broadbent JR, Steele JL (2009) Genome sequence and comparative genome analysis of Lactobacillus casei: insights into their niche-associated evolution. Genome Biol Evol 1:239–257. doi:10.1093/gbe/evp019
Cai H, Rodriguez BT, Zhang W, Broadbent JR, Steele JL (2007) Genotypic and phenotypic characterization of Lactobacillus casei strains isolated from different ecological niches suggests frequent recombination and niche specificity. Microbiology 153:2655–2665. doi:10.1099/mic.0.2007/006452-0
Maze A, Boel G, Zuniga M, Bourand A, Loux V, Yebra MJ, Monedero V, Correia K, Jacques N, Beaufils S, Poncet S, Joyet P, Milohanic E, Casaregola S, Auffray Y, Perez-Martinez G, Gibrat JF, Zagorec M, Francke C, Hartke A, Deutscher J (2010) Complete genome sequence of the probiotic Lactobacillus casei strain BL23. J Bacteriol 192:2647–2648. doi:10.1128/Jb.00076-10
Zhang WY, Yu DL, Sun ZH, Wu RN, Chen X, Chen W, Meng H, Hu SN, Zhang HP (2010) Complete genome sequence of Lactobacillus casei Zhang, a new probiotic strain isolated from traditional homemade Koumiss in Inner Mongolia, China. J Bacteriol 192:5268–5269. doi:10.1128/Jb.00802-10
Chen C, Ai LZ, Zhou FF, Wang L, Zhang H, Chen W, Guo BH (2011) Complete genome sequence of the probiotic bacterium Lactobacillus casei LC2W. J Bacteriol 193:3419–3420. doi:10.1128/Jb.05017-11
Hochwind K, Weinmaier T, Schmid M, van Hemert S, Hartmann A, Rattei T, Rothballer M (2012) Draft genome sequence of Lactobacillus casei W56. J Bacteriol 194:6638. doi:10.1128/Jb.01386-12
Ai LZ, Chen C, Zhou FF, Wang L, Zhang H, Chen W, Guo BH (2011) Complete genome sequence of the probiotic strain Lactobacillus casei BD-II. J Bacteriol 193:3160–3161. doi:10.1128/Jb.00421-11
Koryszewska-Baginska A, Aleksandrzak-Piekarczyk T, Bardowski J (2013) Complete genome sequence of the probiotic strain Lactobacillus casei (formerly Lactobacillus paracasei) LOCK919. Genome Announc 1:e00120. doi:10.1128/genomeA.00758-13
Laing C, Buchanan C, Taboada EN, Zhang Y, Kropinski A, Villegas A, Thomas JE, Gannon VP (2010) Pan-genome sequence analysis using Panseq: an online tool for the rapid analysis of core and accessory genomic regions. BMC Bioinformatics 11:461. doi:10.1186/1471-2105-11-461
Aziz RK, Bartels D, Best AA, DeJongh M, Disz T, Edwards RA, Formsma K, Gerdes S, Glass EM, Kubal M, Meyer F, Olsen GJ, Olson R, Osterman AL, Overbeek RA, McNeil LK, Paarmann D, Paczian T, Parrello B, Pusch GD, Reich C, Stevens R, Vassieva O, Vonstein V, Wilke A, Zagnitko O (2008) The RAST server: rapid annotations using subsystems technology. BMC Genom 9:75. doi:10.1186/1471-2164-9-75
Felsenstein J (1989) PHYLIP-Phylogeny inference package (version 3.2). Cladistics 5:164–166
Darling AE, Mau B, Perna NT (2010) progressiveMauve: multiple genome alignment with gene gain, loss and rearrangement. PLoS ONE 5:e11147. doi:10.1371/journal.pone.0011147
Dereeper A, Guignon V, Blanc G, Audic S, Buffet S, Chevenet F, Dufayard JF, Guindon S, Lefort V, Lescot M, Claverie JM, Gascuel O (2008) Phylogeny.fr: robust phylogenetic analysis for the non-specialist. Nucleic Acids Res 36:W465–W469. doi:10.1093/nar/gkn180 (Web Server issue)
Barrangou R, Fremaux C, Deveau H, Richards M, Boyaval P, Moineau S, Romero DA, Horvath P (2007) CRISPR provides acquired resistance against viruses in prokaryotes. Science 315:1709–1712. doi:10.1126/science.1138140
Garneau JE, Dupuis ME, Villion M, Romero DA, Barrangou R, Boyaval P, Fremaux C, Horvath P, Magadan AH, Moineau S (2010) The CRISPR/Cas bacterial immune system cleaves bacteriophage and plasmid DNA. Nature 468:67–71. doi:10.1038/nature09523
Bottacini F, Medini D, Pavesi A, Turroni F, Foroni E, Riley D, Giubellini V, Tettelin H, van Sinderen D, Ventura M (2010) Comparative genomics of the genus Bifidobacterium. Microbiology 156:3243–3254. doi:10.1099/mic.0.039545-0
de Vos WM (2011) Systems solutions by lactic acid bacteria: from paradigms to practice. Microb Cell Fact 10(Suppl 1):S2. doi:10.1186/1475-2859-10-S1-S2
Marco ML, Pavan S, Kleerebezem M (2006) Towards understanding molecular modes of probiotic action. Curr Opin Biotech 17:204–210. doi:10.1016/j.copbio.2006.02.005
Acknowledgments
This work was jointly supported by the Open Project Program of State Key Laboratory of Dairy Biotechnology, Bright Dairy & Food Co. Ltd., (No. SKLDB2012-007), and the Project of Young Teacher of Ordinary Undergraduate Colleges and Universities for Visiting Abroad by Department of Education of Jiangxi Province, P. R. China, (No. 132 [2012]).
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Yu, S., Peng, Y., Zheng, Y. et al. Comparative Genome Analysis of Lactobacillus casei: Insights into Genomic Diversification for Niche Expansion. Indian J Microbiol 55, 102–107 (2015). https://doi.org/10.1007/s12088-014-0496-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12088-014-0496-2