Abstract
White cypress pine (Callitris glaucophylla) is a drought-tolerant evergreen conifer, which is a member of the Australian C. columellaris species complex. The complex is comprised of five closely related morphospecies that occur in a wide range of bioclimatic regions in Australia. Ecological genomics of the complex provides an opportunity to identify markers associated with environmental adaptation and is expected to broaden our understanding of its speciation process. We adopted a single-tree linkage mapping approach combined with high-throughput restriction site associated DNA (RAD) sequencing and expressed sequence tag-simple sequence repeat (EST-SSR) genotyping to set up a baseline genetic map for C. glaucophylla. The generated linkage map consisted of 4284 markers positioned on 11 linkage groups, corresponding to the haploid chromosome number of Callitris (2n = 22). The spatial distribution of markers was uneven compared to random expectation with significant clustering in central positions of some linkage groups, which may be associated with recombination cold spots of pericentromere regions. Allelic segregation was shown to be distorted in particular regions of four linkage groups, where selection may have operated on viability genes, leaving allelic distortion in surrounding linked markers. We then tested RAD single nucleotide polymorphisms (RAD-SNP) marker recovery and transferability of the linkage map to population genomic data collected for a related species, Callitris gracilis. Of the linkage map markers, 1257 markers (ca. 30 %) were recovered in independent RAD sequencing of population samples of C. glaucophylla. Genetic diversity and differentiation evaluated using mapped markers reflected ascertainment bias slightly; a decrease in Hs (absolute difference of −0.018) for a related species (C. gracilis) and an increase in F ST between C. glaucophylla and C. gracilis (+0.018) were detected. Although care should be taken given such biases in cross-species transfer, this study demonstrated that the RAD-SNP-based linkage map is essentially useful when combined with population genomic analysis of this conifer lineage.



Similar content being viewed by others
References
Albrechtsen A, Nielsen FC, Nielsen R (2010) Ascertainment biases in SNP chips affect measures of population divergence Mol Biol Evol 27:2534–2547
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ (1990) Basic local alignment search tool. J Mol Biol 215:403–410
Andrew RL, Rieseberg LH (2013) Divergence is focused on few genomic regions early in speciation: incipient speciation of sunflower ecotypes. Evolution 67:2468–2482
Arnold B, Corbett‐Detig R, Hartl D, Bomblies K (2013) RADseq underestimates diversity and introduces genealogical biases due to nonrandom haplotype sampling. Mol Ecol 22:3179–3190
Baird NA et al (2008) Rapid SNP discovery and genetic mapping using sequenced RAD markers. PLoS One 3:e3376
Beissinger TM et al (2013) Marker density and read depth for genotyping populations using genotyping-by-sequencing. Genetics 193:1073–1081
Birol I et al (2013) Assembling the 20 Gb white spruce (Picea glauca) genome from whole-genome shotgun sequencing data. Bioinformatics 29:1492–1497. doi:10.1093/bioinformatics/btt178
Bishop D, Cannings C, Skolnick M, Williamson J, Weir B (1983) The number of polymorphic DNA clones required to map the human genome. In: Statistical analysis of DNA sequence data. Marcel-Dekker, New York, pp 181–200
Bolger AM, Lohse M, Usadel B (2014) Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30:2114–2120. doi:10.1093/bioinformatics/btu170
Bowman D, Harris S (1995) Conifers of Australia’s dry forests and open woodlands. In: Enright N, Hill R (eds) Ecology of southern conifers. University of Melbourne, Melbourne, pp 252–270
Bowman DMJS, MacDermott HJ, Nichols SC, Murphy BP (2014) A grass–fire cycle eliminates an obligate-seeding tree in a tropical savanna. Ecol Evol 4:4185–4194. doi:10.1002/ece3.1285
Brodribb TJ, Bowman DM, Grierson PF, Murphy BP, Nichols S, Prior LD (2013) Conservative water management in the widespread conifer genus Callitris. AoB Plants 5:plt052
Bryant D, Moulton V (2004) Neighbor-Net: an agglomerative method for the construction of phylogenetic networks. Mol Biol Evol 21:255–265
Buetow KH (1991) Influence of aberrant observations on high-resolution linkage analysis outcomes. Am J Hum Genet 49:985
Catchen JM, Amores A, Hohenlohe P, Cresko W, Postlethwait JH (2011) Stacks: building and genotyping loci de novo from short-read sequences G3-genes genomes. Genetics 1:171–182. doi:10.1534/g3.111.000240
Chancerel E et al (2013) High-density linkage mapping in a pine tree reveals a genomic region associated with inbreeding depression and provides clues to the extent and distribution of meiotic recombination. BMC Biol 11:50
Choi K et al (2013) Arabidopsis meiotic crossover hot spots overlap with H2A. Z nucleosomes at gene promoters. Nat Genet 45:1327
Chutimanitsakun Y et al (2011) Construction and application for QTL analysis of a restriction site associated DNA (RAD) linkage map in barley. BMC Genomics 12:4
Clark AG, Hubisz MJ, Bustamante CD, Williamson SH, Nielsen R (2005) Ascertainment bias in studies of human genome-wide polymorphism. Genome Res 15:1496–1502
Debreczy Z, Racz I (2006) Conifers around the world vol II. DendroPress Limited, Budapest
Eckert AJ, van Heerwaarden J, Wegrzyn JL, Nelson CD, Ross-Ibarra J, Gonzalez-Martinez SC, Neale DB (2010) Patterns of population structure and environmental associations to aridity across the range of loblolly pine (Pinus taeda L., Pinaceae). Genetics 185:969–982. doi:10.1534/genetics.110.115543
Enright N, Hill R (1995) Ecology of southern conifers. University of Melbourne, Melbourne
Falush D, Stephens M, Pritchard JK (2003) Inference of population structure using multilocus genotype data: linked loci and correlated allele frequencies. Genetics 164:1567–1587
Gaut BS, Wright SI, Rizzon C, Dvorak J, Anderson LK (2007) Recombination: an underappreciated factor in the evolution of plant genomes. Nat Rev Genet 8:77–84
Gautier M et al (2013) The effect of RAD allele dropout on the estimation of genetic variation within and between populations. Mol Ecol 22:3165–3178
Gernandt D, Willyard A, Syring JV, Liston A (2011) The conifers (Pinophyta). In: Plomion C, Bousquet J, Kole C (eds) Genetics, genomics and breeding of conifers. CRC, pp 1–39
Gillet E, Gregorius HR (1992) What can be inferred from open-pollination progenies about the source of observed segregation distortion?–a case study in Castanea sativa Mill. Silvae Genetica 41:82–87
González-Martínez S et al. (2011) Patterns of nucleotide diversity and association mapping. In: Genetics, genomics and breeding of conifers. pp 239–275
Gouded J (2014) hierfstat: Estimation and tests of hierarchical F-statistics. R-package
Guo F et al (2015) Construction of a SNP-based high-density genetic map for pummelo using RAD sequencing. Tree Genetics & Genomes 11:1–11. doi:10.1007/s11295-014-0831-0
Hackett C, Broadfoot L (2003) Effects of genotyping errors, missing values and segregation distortion in molecular marker data on the construction of linkage maps. Heredity 90:33–38
Huson DH (1998) SplitsTree: analyzing and visualizing evolutionary data. Bioinformatics 14:68–73
Isagi Y, Saito D, Kawaguchi H, Tateno R, Watanabe S (2007) Effective pollen dispersal is enhanced by the genetic structure of an Aesculus turbinata population. J Ecol 95:983–990. doi:10.1111/j.1365-2745.2007.01272.x
Iwata H, Ninomiya S (2006) AntMap: constructing genetic linkage maps using an ant colony optimization algorithm. Breed Sci 56:371–377. doi:10.1270/jsbbs.56.371
Kakioka R, Kokita T, Kumada H, Watanabe K, Okuda N (2013) A RAD-based linkage map and comparative genomics in the gudgeons (genus Gnathopogon, Cyprinidae). BMC Genomics 14:32
Kang B-Y, Major JE, Rajora OP (2011) A high-density genetic linkage map of a black spruce (Picea mariana)×red spruce (Picea rubens) interspecific hybrid. Genome 54:128–143
Karam MJ, Lefèvre F, Dagher‐Kharrat MB, Pinosio S, Vendramin G (2015) Genomic exploration and molecular marker development in a large and complex conifer genome using RADseq and mRNAseq Mol Ecol Resour 15:601–612
Kosambi DD (1943) The estimation of map distances from recombination values. Ann Eug 12:172–175
Kuang H, Richardson T, Carson S, Wilcox P, Bongarten B (1999) Genetic analysis of inbreeding depression in plus tree 850.55 of Pinus radiata D. Don. I. Genetic map with distorted markers. Theor Appl Genet 98:697–703
Langmead B, Salzberg SL (2012) Fast gapped-read alignment with Bowtie 2. Nat Methods 9:357–U354. doi:10.1038/nmeth.1923
Larsson H, De Paoli E, Morgante M, Lascoux M, Gyllenstrand N (2013) The hypomethylated partial restriction (HMPR) method reduces the repetitive content of genomic libraries in Norway spruce (Picea abies). Tree Genetics & Genomes 9:601–612
Li S, Chen Y, Gao H, Yin T (2010) Potential chromosomal introgression barriers revealed by linkage analysis in a hybrid of Pinus massoniana and P. hwangshanensis. BMC Plant Biol 10:37. doi:10.1186/1471-2229-10-37
Luca F, Hudson RR, Witonsky DB, Di Rienzo A (2011) A reduced representation approach to population genetic analyses and applications to human evolution. Genome Res 21:1087–1098
Mao KS et al (2012) Distribution of living Cupressaceae reflects the breakup of Pangea. Proc Natl Acad Sci U S A 109:7793–7798. doi:10.1073/pnas.1114319109
Martínez-García PJ, Stevens KA, Wegrzyn JL, Liechty J, Crepeau M, Langley CH, Neale DB (2013) Combination of multipoint maximum likelihood (MML) and regression mapping algorithms to construct a high-density genetic linkage map for loblolly pine (Pinus taeda L.). Tree Genetics & Genomes 9:1529–1535
Moriguchi Y et al (2012) The construction of a high-density linkage map for identifying SNP markers that are tightly linked to a nuclear-recessive major gene for male sterility in Cryptomeria japonica D. Don BMC Genomics 13:95
Moritsuka E, Hisataka Y, Tamura M, Uchiyama K, Watanabe A, Tsumura Y, Tachida H (2012) Extended linkage disequilibrium in noncoding regions in a conifer, Cryptomeria japonica. Genetics 190:1145–1148
Murray MG, Thompson WF (1980) Rapid isolation of high molecular weight plant DNA. Nucleic Acids Res 8:4321–4325
Naito Y et al (2005) Selfing and inbreeding depression in seeds and seedlings of Neobalanocarpus heimii (Dipterocarpaceae). J Plant Res 118:423–430
Neale DB et al (2014) Decoding the massive genome of loblolly pine using haploid DNA and novel assembly strategies. Genome Biol 15:R59
Neves LG, Davis JM, Barbazuk WB, Kirst M (2014) A high-density gene map of loblolly pine (Pinus taeda L.) based on exome sequence capture genotyping G3–Genes Genomes Genetics 4:29–37 g3. 113.008714
Nystedt B et al (2013) The Norway spruce genome sequence and conifer genome evolution. Nature 497:579–584. doi:10.1038/nature12211
Ohri D, Khoshoo T (1986) Genome size in gymnosperms. Plant Syst Evol 153:119–132
Pavy N, Pelgas B, Laroche J, Rigault P, Isabel N, Bousquet J (2012) A spruce gene map infers ancient plant genome reshuffling and subsequent slow evolution in the gymnosperm lineage leading to extant conifers. BMC Biol 10:84
Pavy N et al (2013) Development of high‐density SNP genotyping arrays for white spruce (Picea glauca) and transferability to subtropical and nordic congeners. Mol Ecol Resour 13:324–336
Pegadaraju V, Nipper R, Hulke B, Qi L, Schultz Q (2013) De novo sequencing of sunflower genome for SNP discovery using RAD (restriction site associated DNA) approach. BMC Genomics 14:556
Peterson BK, Weber JN, Kay EH, Fisher HS, Hoekstra HE (2012) Double Digest RADseq: an inexpensive method for de novo SNP discovery and genotyping in model and non-model species. PLoS One 7:e37135. doi:10.1371/journal.pone.0037135
Petes TD (2001) Meiotic recombination hot spots and cold spots. Nat Rev Genet 2:360–369
Pool JE, Hellmann I, Jensen JD, Nielsen R (2010) Population genetic inference from genomic sequence variation. Genome Res 20:291–300
Prior LD, McCaw WL, Grierson PF, Murphy BP, Bowman DMJS (2011) Population structures of the widespread Australian conifer Callitris columellaris are a bio-indicator of continental environmental change. For Ecol Manag 262:252–262. doi:10.1016/j.foreco.2011.03.030
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959
Purcell S et al (2007) PLINK: a tool set for whole-genome association and population-based linkage analyses. Am J Hum Genet 81:559–575. doi:10.1086/519795
R Development Core Team (2014) R version 3.1.0: A language and environment for statistical computing
Rabinowicz PD et al (2005) Differential methylation of genes and repeats in land plants. Genome Res 15:1431–1440
Ritland K, Krutovsky KV, Tsumura Y, Pelgas B, Isabel N, Bousquet J (2011) Genetic mapping in conifers. In: Genetics, genomics and breeding of conifers. CRC, pp 196–238
Sakaguchi S et al (2011) Isolation and characterization of 52 polymorphic EST-SSR markers for Callitris columellaris (Cupressaceae). Am J Bot 98:E363–E368. doi:10.3732/ajb.1100276
Sakaguchi S, Bowman DMJS, Prior LD, Crisp MD, Linde CC, Tsumura Y, Isagi Y (2013) Climate, not Aboriginal landscape burning, controlled the historical demography and distribution of fire-sensitive conifer populations across Australia. P Roy Soc B–Biol Sci 28:1–9 doi:10.1098/rspb.2013.2182
Savolainen O, Lascoux M, Merila J (2013) Ecological genomics of local adaptation. Nat Rev Genet 14:807–820. doi:10.1038/nrg3522
Siregar IZ, Yunanto T (2008) Inference on the possible causes of segregation distortion from open pollination progenies of merkus pine (Pinus merkusii). HAYATI J Biosci 15:173
Slavov GT et al (2014) Genome-wide association studies and prediction of 17 traits related to phenology, biomass and cell wall composition in the energy grass Miscanthus sinensis. New Phytol 201:1227–1239. doi:10.1111/nph.12621
Studer B et al (2012) A transcriptome map of perennial ryegrass (Lolium perenne L.). BMC Genomics 13:140
Talukder ZI, Gong L, Hulke BS, Pegadaraju V, Song Q, Schultz Q, Qi L (2014) A high-density SNP map of sunflower derived from RAD-sequencing facilitating fine-mapping of the rust resistance gene R12. PLoS One 9:e98628
Tsumura Y, Kado T, Takahashi T, Tani N, Ujino-Ihara T, Iwata H (2007) Genome scan to detect genetic structure and adaptive genes of natural populations of Cryptomeria japonica. Genetics 176:2393–2403
Tsumura Y, Uchiyama K, Moriguchi Y, Kimura MK, Ueno S, Ujino-Ihara T (2014) Genetic differentiation and evolutionary adaptation in Cryptomeria japonica. G3: Genes| Genomes| Genetics 4:2389–2402
Tulsieram LK, Glaubitz JC, Kiss G, Carlson JE (1992) Single tree genetic linkage mapping in conifers using haploid DNA from megagametophytes. Nat Biotechnol 10:686–690
Voorrips RE (2002) MapChart: software for the graphical presentation of linkage maps and QTLs. J Hered 93:77–78. doi:10.1093/jhered/93.1.77
Ward JA et al (2013) Saturated linkage map construction in Rubus idaeus using genotyping by sequencing and genome-independent imputation. BMC Genomics 14:2. doi:10.1186/1471-2164-14-2
Warren RL et al (2015) Improved white spruce (Picea glauca) genome assemblies and annotation of large gene families of conifer terpenoid and phenolic defense metabolism. Plant J 83:189–212. doi:10.1111/tpj.12886
Wu J et al (2003) Physical maps and recombination frequency of six rice chromosomes. Plant J 36:720–730
Wu Y, Bhat PR, Close TJ, Lonardi S (2008) Efficient and accurate construction of genetic linkage maps from the minimum spanning tree of a graph. PLoS Genet 4:e1000212
Wu J et al (2014) High-density genetic linkage map construction and identification of fruit-related QTLs in pear using SNP and SSR markers. J Exp Bot 65:5771–5781 doi:10.1093/jxb/eru311
Acknowledgments
We are grateful to NSW State Forests, NSW Department of Environment and Climate Change, Queensland Department of Environment and Resource Management, SA Department of Environment and Heritage, Victorian Department of Sustainability and Environment, the Australian Wildlife Conservancy, and many private landholders for help with site selection and permission to sample on their land. We thank Lynda D. Prior for collecting population samples, N. Nakahama and Y. Unno for their assists in genotyping EST-SSRs and a preliminary analysis, M. Yamasaki for his insightful discussions on statistical analysis, and Y. Moriguchi for kindly providing unpublished data. RAD-sequencing experiment was conducted using Joint Usage/Research Program of Center for Ecological Research, Kyoto University. Funding was provided by Japan Society for the Promotion of Science Grant-in-Aid for JSPS Fellows (13 J06059), Grant-in-Aid for Scientific Research (JSPS KAKENHI 24248028 and 26850098), and the Environment Research and Technology Development Fund of the Ministry of the Environment (4-1403).
Data archiving statement
Raw RAD sequence data is deposited at DDBJ Sequence Read Archive (DRA) with accession numbers of DRA003554 (submission), PRJDB3893 (BioProject), SAMD00029742 (BioSample), DRX031648 (Experiment), and DRR035015 (Run). Details of each sequence read file can be found in supplementary material 4.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by W. Ratnam
This article is part of the Topical Collection on Germplasm Diversity
Electronic supplementary material
Below is the link to the electronic supplementary material.
Supplementary material 1
Oligonucleotide sequences used as adaptor and index primers in RAD-sequencing library construction. (DOC 31 kb)
Supplementary material 2
Description of RAD reference assembly of Callitris columellaris species complex, which includes information of the individual samples used for Miseq (Illumina, San Diego, USA) sequencing and analysis protocol for short read assembly. (DOC 38 kb)
Supplementary material 3
Information of genetic markers mapped on the linkage groups. (CSV 889 kb)
Supplementary material 4
Information of RAD-sequencing read files. (CSV 9 kb)
Supplementary figure 1
(a) Distribution of Callitris columellaris species complex. Ranges for C. glaucophylla (GD lineage in Sakaguchi et al. (2013)) and C. gracilis are colored by ochre and pink, respectively. The locality where the seed samples for linkage map construction were collected is indicated by a black square on the smaller map. Superimposed are pie charts illustrating the two genetic clusters, corresponding to the two species, which were detected by STRUCTURE analysis. (b) A split network for 31 individuals of C. glaucophylla and C. gracilis analyzed in this study. Genetic membership estimated from STRUCTURE analysis is placed on the tips. A genetically intermediate individual is indicated by a green triangle. (GIF 134 kb)
Supplementary figure 2
Graphical results of GAM analysis of genetic marker segregation. Partial effects of genomic position are shown for each linkage group, expressed as fitted loess functions with 95 % boot-strapped confidence intervals (gray in color). Ticks in the x-axis represent the location of observations along the predictor. (GIF 127 kb)
Supplementary figure 3
Relationship between marker positions estimated from genotype data sets with and without imputation and error correction procedures. (GIF 97 kb)
Supplementary figure 4
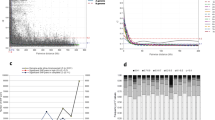
Distribution of genetic marker density calculated based on different bin widths (1, 5, 10 cM). Expected probability curves are estimated using a Poisson distribution (blue) and a negative binomial distribution (red) (GIF 78 kb)
Supplementary table 1
Results of GAM analysis of genetic marker segregation for each linkage group, as a function of missing rate in genotype data and map position. (DOCX 27 kb)
Rights and permissions
About this article
Cite this article
Sakaguchi, S., Sugino, T., Tsumura, Y. et al. High-throughput linkage mapping of Australian white cypress pine (Callitris glaucophylla) and map transferability to related species. Tree Genetics & Genomes 11, 121 (2015). https://doi.org/10.1007/s11295-015-0944-0
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11295-015-0944-0