Abstract
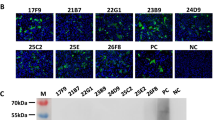
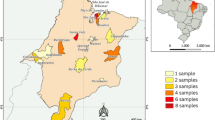
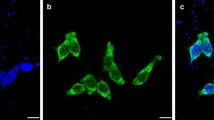
Puumala virus (PUUV) is the predominant hantavirus species in Germany causing large numbers of mild to moderate cases of haemorrhagic fever with renal syndrome (HFRS). During an outbreak in South-East Germany in 2004 a novel PUUV subtype designated Bavaria was identified as the causative agent of HFRS in humans [1]. Here we present a molecular characterization of this PUUV strain by investigating novel partial and almost entire nucleocapsid (N) protein-encoding small (S-) segment sequences and partial medium (M-) segment sequences from bank voles (Myodes glareolus) trapped in Lower Bavaria during 2004 and 2005. Phylogenetic analyses confirmed their classification as subtype Bavaria, which is further subdivided into four geographical clusters. The entire N protein, harbouring an amino-terminal hexahistidine tag, of the Bavarian strain was produced in yeast Saccharomyces cerevisiae and showed a slightly different reactivity with N-specific monoclonal antibodies, compared to the yeast-expressed N protein of the PUUV strain Vranica/Hällnäs. Endpoint titration of human sera from different parts of Germany and from Finland revealed only very slight differences in the diagnostic value of the different recombinant proteins. Based on the novel N antigen indirect and monoclonal antibody capture IgG-ELISAs were established. By using serum panels from Germany and Finland their validation demonstrated a high sensitivity and specificity. In summary, our investigations demonstrated the Bavarian PUUV strain to be genetically divergent from other PUUV strains and the potential of its N protein for diagnostic applications.





Similar content being viewed by others
References
S. Essbauer, J. Schmidt, F.J. Conraths, R. Friedrich, J. Koch, W. Hautmann, M. Pfeffer, R. Wölfel, J. Finke, G. Dobler, R. Ulrich, A new Puumala hantavirus subtype in rodents associated with an outbreak of Nephropathia epidemica in South-East Germany in 2004. Epidemiol. Infect. 134, 1333–1344 (2006)
S.T. Nichol, B.J. Beaty, R.M. Elliott, R. Goldbach, A. Plyusnin, C.S. Schmaljohn, R.B. Tesh, Family Bunyaviridae, in Virus Taxonomy, ed. by C.M. Fauquet, M.A. Mayo, J. Maniloff, U. Desselberger, L.A. Ball. 8th Report of the International Committee on Taxonomy of Viruses (Elsevier Academic Press, San Diego, CA, 2005), pp. 695–716
C.S. Schmaljohn, S.T. Nichol, Bunyaviridae, in Fields Virology, vol. 2, 5th edn., ed. by D.M. Knipe, P.M. Howley (Lippencott, Williams & Wilkins, Philadelphia, PA, 2007), pp. 1741–1789
A. Plyusnin, S.P. Morzunov, Virus evolution and genetic diversity of hantaviruses and their rodent hosts. Curr. Top. Microbiol. Immunol. 256, 47–75 (2001)
R. Ulrich, B. Hjelle, C. Pitra, D.H. Krüger, Emerging viruses: the case “Hantavirus”. Intervirology 45, 318–327 (2002)
K.M. Jääskeläinen, A. Plyusnina, Å. Lundkvist, A. Vaheri, A. Plyusnin, Tula hantavirus isolate with the full-length ORF for nonstructural protein NSs survives for more consequent passages in interferon-competent cells than the isolate having truncated NSs ORF. Virol. J. 5, 3 (2008)
G. Schönrich, A. Rang, N. Lütteke, M.J. Raftery, N. Charbonnel, R.G. Ulrich, Hantavirus-induced immunity in rodent reservoirs and humans. Immunol. Rev. 225, 163–189 (2008)
R.M. Wells, S. Sosa Estani, Z.E. Yadon, D. Enria, P. Padula, N. Pini, J.N. Mills, C.J. Peters, E.L. Segura, An unusual hantavirus outbreak in southern Argentina: person-to-person transmission? Hantavirus Pulmonary Syndrome Study Group for Patagonia. Emerg. Infect. Dis. 3, 171–174 (1997)
P.J. Padula, A. Edelstein, S.D. Miguel, N.M. Lopez, C.M. Rossi, R.D. Rabinovich, Hantavirus pulmonary syndrome outbreak in Argentina: molecular evidence for person-to-person transmission of Andes virus. Virology 241, 323–330 (1998)
D.E. Carey, R. Reuben, K.N. Panicker, R.E. Shope, R.M. Myers, Thottapalayam virus: a presumptive arbovirus isolated from a shrew in India. Indian J. Med. Res. 59, 1758–1760 (1971)
J.W. Song, S.H. Gu, S.N. Bennett, S. Arai, M. Puorger, M. Hilbe, R. Yanagihara, Seewis virus, a genetically distinct hantavirus in the Eurasian common shrew (Sorex araneus). Virol. J. 4, 114 (2007)
J.W. Song, H.J. Kang, K.J. Song, T.T. Truong, S.N. Bennett, S. Arai, N.U. Truong, R. Yanagihara, Newfound hantavirus in Chinese mole shrew, Vietnam. Emerg. Infect. Dis. 13, 1784–1787 (2007)
B. Klempa, E. Fichet-Calvet, E. Lecompte, B. Auste, V. Aniskin, H. Meisel, P. Barriere, L. Koivogui, J. ter Meulen, D.H. Krüger, Novel hantavirus sequences in shrew, Guinea. Emerg. Infect. Dis. 13, 520–522 (2007)
S. Arai, J.W. Song, L. Sumibcay, S.N. Bennett, V.R. Nerurkar, C. Parmenter, J.A. Cook, T.L. Yates, R. Yanagihara, Hantavirus in northern short-tailed shrew, United States. Emerg. Infect. Dis. 13, 1420–1423 (2007)
H.J. Kang, S.N. Bennett, L. Sumibcay, S. Arai, A.G. Hope, G. Mocz, J.W. Song, J.A. Cook, R. Yanagihara, Evolutionary insights from a genetically divergent hantavirus harbored by the European common mole (Talpa europaea). PLoS One 4, 7 (2009)
H. Kariwa, S. Yoshizumi, J. Arikawa, K. Yoshimatsu, K. Takahashi, I. Takashima, N. Hashimoto, Evidence for the existence of Puumula-related virus among Clethrionomys rufocanus in Hokkaido. Japan. Am. J. Trop. Med. Hyg. 53, 222–227 (1995)
N.H. Daud, H. Kariwa, Y. Tanikawa, I. Nakamura, T. Seto, D. Miyashita, K. Yoshii, M. Nakauchi, K. Yoshimatsu, J. Arikawa, I. Takashima, Mode of infection of Hokkaido virus (Genus Hantavirus) among grey red-backed voles, Myodes rufocanus, in Hokkaido, Japan. Microbiol. Immunol. 51, 1081–1090 (2007)
A. Plyusnina, J. Laakkonen, J. Niemimaa, K. Nemirov, G. Muruyeva, B. Pohodiev, Å. Lundkvist, A. Vaheri, H. Henttonen, O. Vapalahti, A. Plyusnin, Genetic analysis of hantaviruses carried by Myodes and Microtus rodents in Buryatia. Virol. J. 5, 4 (2008)
J. Mustonen, O. Vapalahti, H. Henttonen, A. Pasternack, A. Vaheri, Epidemiology of hantavirus infections in Europe. Nephrol. Dial. Transplant. 13, 2729–2731 (1998)
H. Kallio-Kokko, N. Uzcategui, O. Vapalahti, O. Vaheri, Viral zoonoses in Europe. FEMS Microbiol. Rev. 29, 1051–1077 (2005)
S. Sandmann, H. Meisel, A. Razanskiene, A. Wolbert, B. Pohl, D.H. Krüger, K. Sasnauskas, R. Ulrich, Detection of human hantavirus infections in Lithuania. Infection 33(2), 66–72 (2005)
L. Fontana-Binard, D. Schultze, B.S. Rojanavisut, D.H. Krüger, G. Dollenmaier, G. Zanetti, P. Meylan, First case of nephropathia epidemica acquired in Switzerland. Rev. Med. Suisse. 4(163), 1572–1575 (2008)
S. Grygorczuk, S. Pancewicz, J. Zajkowska, M. Kondrusik, R. Swierzbińska, A. Moniuszko, W. Pawlak-Zalewska, Detection of anti-hantavirus antibodies in forest workers in the north-east of Poland. Przegl. Epidemiol. 62, 531–537 (2008)
L. Zöller, M. Faulde, H. Meisel, B. Ruh, P. Kimmig, U. Schelling, M. Zeier, P. Kulzer, C. Becker, M. Roggendorf, E.K.F. Bautz, D.H. Krüger, G. Darai, Seroprevalence of hantavirus antibodies in Germany as determined by a new recombinant enzyme immunoassay. Eur. J. Clin. Microbiol. Infect. Dis. 14, 305–313 (1995)
R. Ulrich, H. Meisel, M. Schütt, J. Schmidt, A. Kunz, B. Klempa, M. Niedrig, P. Kimmig, G. Pauli, D.H. Krüger, J. Koch, Verbreitung von Hantavirusinfektionen in Deutschland. Bundesgesundheitsbl. Gesundheitsforsch. Gesundheitsschutz 47, 661–670 (2004)
S.S. Essbauer, J. Schmidt-Chanasit, E.L. Madeja, W. Wegener, R. Friedrich, R. Petraityte, K. Sasnauskas, J. Jacob, J. Koch, G. Dobler, F.J. Conraths, M. Pfeffer, C. Pitra, R.G. Ulrich, Nephropathia epidemica outbreak in a metropolitan area, Germany. Emerg. Infect. Dis. 13, 1271–1273 (2007)
J. Koch, S.O. Brockmann, C. Winter, P. Kimmig, K. Stark, Significant increase of hantavirus infections in Germany since the beginning of 2007. Euro Surveill. 12, 5 (2007)
J. Hofmann, H. Meisel, B. Klempa, S.M. Vesenbeckh, R. Beck, D. Michel, J. Schmidt-Chanasit, R.G. Ulrich, S. Grund, G. Enders, D.H. Krüger, Molecular epidemiology of a large hantavirus outbreak in Germany, 2007. Emerg. Infect. Dis. 14, 850–852 (2008)
S. Schilling, P. Emmerich, B. Klempa, B. Auste, E. Schnaith, H. Schmitz, D.H. Krüger, S. Günther, H. Meisel, Hantavirus outbreak in Germany: limitations of routine serological diagnostics and clustering of virus sequences of human and rodent origin. J. Clin. Microbiol. 45, 3008–3014 (2007)
H.W. Lee, C. Calisher, C.S. Schmaljohn, Manual of Hemorrhagic Fever with Renal Syndrome and Hantavirus Pulmonary Syndrome (WHO Collaborating Center for Virus Reference and Research, Seoul, 1999)
D.H. Krüger, R. Ulrich, Å. Lundkvist, Hantavirus infections and their prevention. Microbes. Infect. 3, 1129–1144 (2001)
M. Mertens, R. Wölfel, K. Ullrich, K. Yoshimatsu, J. Blumhardt, I. Römer, J. Esser, J. Schmidt-Chanasit, M.H. Groschup, G. Dobler, S.S. Essbauer, R.G. Ulrich, Seroepidemiological study in a Puumala virus outbreak area in South-East Germany. Med. Mircobiol. Immunol. 198, 83–91 (2009)
H. Feldmann, A. Sanchez, S. Morzunov, C.F. Spiropoulou, P.E. Rollin, T.G. Ksiazek, C.J. Peters, S.T. Nichol, Utilization of autopsy RNA for the synthesis of the nucleocapsid antigen of a newly recognized virus associated with hantavirus pulmonary syndrome. Virus Res. 30, 351–367 (1993)
F. Elgh, M. Linderholm, G. Wadell, A. Tarnvik, P. Juto, Development of humoral cross-reactivity to the nucleocapsid protein of heterologous hantaviruses in nephropathia epidemica. FEMS Immunol. Med. Microbiol. 22, 309–315 (1998)
K.B. Sjolander, Å. Lundkvist, Dobrava virus infection: serological diagnosis and cross-reactions to other hantaviruses. J. Virol. Methods 80(2), 137–143 (1999)
J. Schmidt, H. Meisel, B. Hjelle, D.H. Krüger, R. Ulrich, Development and evaluation of serological assays for detection of human hantavirus infections caused by Sin Nombre virus. J. Clin. Virol. 33, 247–253 (2005)
J. Schmidt, H. Meisel, S.G. Capria, R. Petraityte, Å. Lundkvist, B. Hjelle, P.A. Vial, P. Padula, D.H. Krüger, R. Ulrich, Serological assays for the detection of human Andes hantavirus infections based on its yeast-expressed nucleocapsid protein. Intervirology 49, 173–184 (2006)
H. Meisel, A. Wolbert, A. Razanskiene, A. Marg, A. Kazaks, K. Sasnauskas, G. Pauli, R. Ulrich, D.H. Krüger, Development of novel immunoglobulin G (IgG), IgA and IgM enzyme immunoassays based on recombinant Puumala and Dobrava hantavirus nucleocapsid proteins. Clin. Vaccine Immunol. 13, 1349–1357 (2006)
A. Zvirbliene, L. Samonskyte, A. Gedvilaite, T. Voronkova, R. Ulrich, K. Sasnauskas, Generation of monoclonal antibodies of desired specificity using chimeric polyomavirus-derived virus-like particles. J. Immunol. Methods 311, 57–70 (2006)
Å. Lundkvist, A. Fatouros, B. Niklasson, Antigenic variation of European haemorrhagic fever with renal syndrome virus strains characterized using bank vole monoclonal antibodies. J. Gen. Virol. 72, 2097–2103 (1991)
Å. Lundkvist, B. Niklasson, Bank vole monoclonal antibodies against Puumala virus envelope glycoproteins: identification of epitopes involved in neutralization. Arch. Virol. 126, 93–105 (1992)
L.G. Zöller, S. Yang, P. Gött, E.K. Bautz, G. Darai, A novel μ-capture enzyme-linked immunosorbent assay based on recombinant proteins for sensitive and specific diagnosis of hemorrhagic fever with renal syndrome. J. Clin. Microbiol. 31, 1194–1199 (1993)
K. Yoshimatsu, Y.C. Yoo, R. Yoshida, C. Ishihara, I. Azuma, J. Arikawa, Protective immunity of Hantaan virus nucleocapsid and envelope protein studied using baculovirus-expressed proteins. Arch. Virol. 130, 365–376 (1993)
I. Kucinskaite-Kodze, R. Petraityte-Burneikiene, A. Zvirbliene, B. Hjelle, R.A. Medina, A. Gedvilaite, A. Razanskiene, J. Schmidt-Chanasit, M. Mertens, P. Padula, K. Sasnauskas, R.G. Ulrich, Characterization of monoclonal antibodies against hantavirus nucleocapsid protein and their use for immunohistochemistry on rodent and human samples. Arch. Virol. 156, 443–456 (2011)
A. Plyusnin, J. Hörling, M. Kanerva, J. Mustonen, Y. Cheng, J. Partanen, O. Vapalahti, S.K. Kukkonen, J. Niemimaa, H. Henttonen, B. Niklasson, Å. Lundkvist, A. Vaheri, Puumala hantavirus genome in patients with nephropathia epidemica: correlation of PCR positivity with HLA haplotype and link to viral sequences in local rodents. J. Clin. Microbiol. 35, 1090–1096 (1997)
A. Razanskiene, J. Schmidt, A. Geldmacher, A. Ritzi, M. Niedrig, Å. Lundkvist, D.H. Krüger, H. Meisel, K. Sasnauskas, R. Ulrich, High yields of stable and highly pure nucleocapsid proteins of different hantaviruses can be generated in the yeast Saccharomyces cerevisiae. J. Biotechnol. 111, 319–333 (2004)
A. Dargeviciute, K. Brus Sjölander, K. Sasnauskas, D.H. Krüger, H. Meisel, R. Ulrich, Å. Lundkvist, Yeast-expressed Puumala hantavirus nucleocapsid protein induces protection in a bank vole model. Vaccine 20, 3523–3531 (2002)
J.D. Thompson, T.J. Gibson, F. Plewniak, F. Jeanmougin, D.G. Higgins, The CLUSTAL_X windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res. 25, 4876–4882 (1997)
T.A. Hall, BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acid Symp. Ser. 41, 95–98 (1999)
S. Braaker, G. Heckel, Transalpine colonisation and partial phylogeographic erosion by dispersal in the common vole Microtus arvalis. Mol. Ecol. 18, 2518–2531 (2009)
N. Saitou, M. Nei, The neighbor-joining method: a new method for reconstructing phylogentic trees. Mol. Biol. Evol. 4, 406–425 (1987)
S. Kumar, K. Tamura, M. Nei, Mega3: integrated software for molecular evolutionary genetics analysis and sequence alignment. Brief. Bioinform. 5, 150–163 (2004)
F. Ronquist, J.P. Huelsenbeck, MrBayes 3: Bayesian phylogenetic inference under mixed models. Bioinformatics 19, 1572 (2003)
G.E. Schwarz, Estimating the dimension of a model. Ann. Stat. 6(2), 461–464 (1978)
D. Posada, iModelTest: phylogenetic model averaging. Mol. Biol. Evol. 25, 1253–1256 (2008)
K. Tamura, M. Nei, P.S. Kumar, Prospects for inferring very large phylogenies by using the neighbor-joining method. Proc. Natl Acad. Sci. USA 101, 11030–11035 (2004)
M.A. Miller, M.T. Holder, R. Vos, P.E. Midford, T. Liebowitz, L. Chan, P. Hoover, T. Warnow, The CIPRES Portals. CIPRES, http://www.phylo.org/sub_sections/portal. Accessed 4 Aug 2009 (Archived by WebCite(r) at http://www.webcitation.org/5imQlJeQa)
J.C. Wilgenbusch, D.L. Warren, D.L. Swofford, AWTY: a system for graphical exploration of MCMC convergence in Bayesian phylogenetic inference (2004), http://ceb.csit.fsu.edu/awty
E.R. Kallio, J. Klingström, E. Gustafsson, T. Manni, A. Vaheri, H. Henttonen, O. Vapalahti, Å. Lundkvist, Prolonged survival of Puumala hantavirus outside the host: evidence for indirect transmission via the environment. J. Gen. Virol. 87, 2127–2134 (2006)
J. Hardestam, M. Simon, K.O. Hedlund, A. Vaheri, J. Klingström, Å. Lundkvist, Ex vivo stability of the rodent-borne Hantaan virus in comparison to that of arthropod-borne members of the Bunyaviridae family. Appl. Environ. Microbiol. 73, 2547–2551 (2007)
S. Schex, G. Dobler, J. Riehm, J. Müller, S. Essbauer, Rickettsia spp. in wild small mammals in Lower Bavaria, South-Eastern Germany. Vector Borne Zoonotic Dis. (2010) [Epub ahead of print]
R.G. Ulrich, J. Schmidt-Chanasit, M. Schlegel, J. Jacob, H.-J. Pelz, M. Mertens, M. Wenk, T. Büchner, D. Masur, K. Sevke, M.H. Groschup, F.-W. Gerstengarbe, M. Pfeffer, R. Oehme, W. Wegener, M. Bemmann, L. Ohlmeyer, R. Wolf, H. Zoller, J. Koch, S. Brockmann, G. Heckel, S.S. Essbauer, Network “Rodent-borne pathogens” in Germany: longitudinal studies on the geographical distribution and prevalence of hantavirus infections. Parasitol. Res. 103(Suppl. 1), 121–129 (2008)
M. Razzauti, A. Plyusnina, H. Henttonen, A. Plyusnin, Accumulation of point mutations and reassortment of genomic RNA segments are involved in the microevolution of Puumala hantavirus in a bank vole (Myodes glareolus) population. J. Gen. Virol. 89, 1649–1660 (2008)
A. Dekonenko, V. Yakimenko, A. Ivanov, V. Morozov, P. Nikitin, S. Khasanova, T. Dzagurova, E. Tkachenko, C. Schmaljohn, Genetic similarity of Puumala viruses found in Finland and Western Siberia and of the mitochondrial DNA of their rodent hosts suggests a common evolutionary origin. Infect. Genet. Evol. 3, 245–257 (2003)
J. Schmidt-Chanasit, S. Essbauer, R. Petraityte, K. Yoshimatsu, K. Tackmann, F.J. Conraths, K. Sasnauskas, J. Arikawa, A. Thomas, M. Pfeffer, J.J. Scharninghausen, W. Splettstoesser, M. Wenk, G. Heckel, R.G. Ulrich, Extensive host sharing of Central European Tula virus. J. Virol. 84, 459–474 (2010)
P. Kotlik, V. Deffontaine, S. Mascheretti, J. Zima, J.R. Michaux, J.B. Searle, A northern glacial refugium for bank voles (Clethrionomys glareolus). Proc. Natl Acad. Sci. USA 103, 14860–14864 (2006)
G. Gerlach, K.F. Musolf, Fragmentation of landscape as a cause for genetic subdivision in bank voles. Conserv. Biol. 14(4), 1066–1074 (2000)
B. Walser, G. Heckel, Microsatellite markers for the common vole (Microtus arvalis) and their cross-species utility. Conserv. Genet. 9, 479–481 (2008)
M. Schweizer, L. Excoffier, G. Heckel, Fine-scale genetic structure and dispersal patterns in the common vole (Microtus arvalis). Mol. Ecol. 16, 2463–2473 (2007)
G. Heckel, R. Burri, S. Fink, J.-F. Desmet, L. Excoffier, Genetic structure and colonization processes in European populations of the common vole, Microtus arvalis. Evolution 59, 2231–2242 (2005)
V. Deffontaine, R. Libois, P. Kotlik, R. Sommer, C. Nieberding, E. Paradis, J.B. Searle, J.R. Michaux, Beyond the Mediterranean peninsulas: evidence of central European glacial refugia for a temperate forest mammal species, the bank vole (Clethrionomys glareolus). Mol. Ecol. 14, 1727–1739 (2005)
V. Deffontaine, R. Ledevin, M.C. Fontaine, J.P. Quéré, S. Renaud, R. Libois, J.R. Michaux, A relict bank vole lineage highlights the biogeographic history of the Pyrenean region in Europe. Mol. Ecol. 18, 2489–2502 (2009)
Å. Lundkvist, H. Meisel, D. Koletzki, H. Lankinen, F. Cifire, A. Geldmacher, C. Sibold, P. Gött, A. Vaheri, D.H. Krüger, R. Ulrich, Mapping of B-cell epitopes in the nucleocapsid protein of Puumala hantavirus. Viral Immunol. 15, 177–192 (2002)
K. Yoshimatsu, J. Arikawa, M. Tamura, R. Yoshida, Å. Lundkvist, B. Niklasson, H. Kariwa, I. Azuma, Characterization of the nucleocapsid protein of Hantaan virus strain 76–118 using monoclonal antibodies. J. Gen. Virol. 77, 695–704 (1996)
K.B. Sundström, M. Stoltz, N. Lagerqvist, Å. Lundkvist, K. Nemirov, J. Klingström, Characterization of two substrains of Puumala virus that show phenotypes that are different from each other and from the original strain. J. Virol. 85, 1747–1756 (2011)
D. Samuel, K. Sasnauskas, L. Jin, A. Gedvilaite, R. Slibinskas, S. Beard, A. Zvirbliene, S.A. Oliveira, J. Staniulis, B. Cohen, D. Brown, Development of a measles specific IgM ELISA for use with serum and oral fluid samples using recombinant measles nucleoprotein produced in Saccharomyces cerevisiae. J. Clin. Virol. 28, 121–129 (2003)
R. Slibinskas, D. Samuel, A. Gedvilaite, J. Staniulis, K. Sasnauskas, Synthesis of the measles virus nucleoprotein in yeast Pichia pastoris and Saccharomyces cerevisiae. J. Biotechnol. 107, 115–124 (2004)
M. Juozapaitis, A. Serva, A. Zvirbliene, R. Slibinskas, J. Staniulis, K. Sasnauskas, B.J. Shiell, L.F. Wang, W.P. Michalski, Generation of henipavirus nucleocapsid proteins in yeast Saccharomyces cerevisiae. Virus Res. 124, 95–102 (2007)
M. Juozapaitis, A. Serva, I. Kucinskaite, A. Zvirbliene, R. Slibinskas, J. Staniulis, K. Sasnauskas, B.J. Shiell, T.R. Bowden, W.P. Michalski, Generation of menangle virus nucleocapsid-like particles in yeast Saccharomyces cerevisiae. J. Biotechnol. 130, 441–447 (2007)
M. Juozapaitis, A. Zvirbliene, I. Kucinskaite, I. Sezaite, R. Slibinskas, M. Coiras, F. de Ory Manchon, M.R. López-Huertas, P. Pérez-Breña, J. Staniulis, I. Narkeviciute, K. Sasnauskas, Synthesis of recombinant human parainfluenza virus 1 and 3 nucleocapsid proteins in yeast Saccharomyces cerevisiae. Virus Res. 133, 178–186 (2008)
R. Petraityte, P.L. Tamosiunas, M. Juozapaitis, A. Zvirbliene, K. Sasnauskas, B. Shiell, G. Russell, J. Bingham, W.P. Michalski, Generation of Tioman virus nucleocapsid-like particles in yeast Saccharomyces cerevisiae. Virus Res. 145, 92–96 (2009)
P.J. Padula, C.M. Rossi, M.O. Della Valle, P.V. Martinez, S.B. Colavecchia, A. Edelstein, S.D.L. Miguel, R.D. Rabinovich, E.L. Segura, Development and evaluation of a solid-phase enzyme immunoassay based on Andes hantavirus recombinant nucleoprotein. J. Med. Microbiol. 49, 149–155 (2000)
R. Petraityte, L. Jin, R. Hunjan, A. Razanskiene, A. Zvirbliene, K. Sasnauskas, Use of Saccharomyces cerevisiae-expressed recombinant nucleocapsid protein to detect Hantaan virus-specific immunoglobulin G (IgG) and IgM in oral fluid. Clin. Vaccine Immunol. 14, 1603–1608 (2007)
R. Petraitytė, H. Yang, R. Hunjan, R. Ražanskienė, P. Dhanilall, R.G. Ulrich, K. Sasnauskas, L. Jin, Development and evaluation of serological assays for detection of Hantaan virus-specific antibodies in human sera using yeast-expressed nucleocapsid protein. J. Virol. Methods 148, 89–95 (2008)
K. Brus Sjölander, F. Elgh, H. Kallio-Kokko, O. Vapalahti, M. Hägglund, V. Palmcrantz, P. Juto, A. Vaheri, B. Niklasson, Å. Lundkvist, Evaluation of serological methods for diagnosis of Puumala hantavirus infection (nephropathia epidemica). J. Clin. Microbiol. 35, 3264–3268 (1997)
Acknowledgements
We would like to thank Ausra Razanskiene and Kestutis Sasnauskas (Vilnius) and Bernd Köllner (Riems) for continous support and helpful discussions, Åke Lundkvist (Stockholm), Kumiko Yoshimatsu and Jiro Arikawa (Sapporo) for provision of N protein-specific mAbs and Olli Vapalahti (Helsinki), Regina Allwinn and Holger F. Rabenau (Frankfurt) for provision of human serum panels. We kindly acknowledge Harald Weber, Aileene Lorber, Rahime Terzioglu, Elmar Schröpfer, Ralf Hagen, Mirko Köhler, Anne Grumbach, Peter Klein and Kerstin Weiss for their support during rodent trapping, rodent necropsy and collection of human sera in South-East Germany. The excellent technical lab assistance of Jana Blumhardt, Ina Römer, Dörte Kaufmann, Sabine Fink and Astrid Thomas is gratefully acknowledged. The work was supported by Research-Project M/SAB1/5/A017 for the Bundeswehr Medical Service to RGU.
Author information
Authors and Affiliations
Corresponding author
Additional information
Marc Mertens and Eveline Kindler contributed equally to this paper.
Electronic supplementary material
Below is the link to the electronic supplementary material.
11262_2011_620_MOESM1_ESM.pptx
Supplementary Figure. Bayesian reconstruction of phylogenetic relationships based on 968 bp mtDNA (cyt b) from 120 Myodes glareolus with M. rufocanus as outgroup. Only support values for main nodes that connect major evolutionary lineages are displayed. Posterior probabilities are indicated above the major branches and percentage of bootstrap support for neighbour-joining (NJ) algorithms below the branches. * indicates a different topology based on NJ algorithms, ns refers to posterior probabilities <0.50 and percentages of bootstrap support <50%. Arrows depict cyt b sequences from bank voles from Bavaria where PUUV sequences were detected in (PPTX 64 kb)
Rights and permissions
About this article
Cite this article
Mertens, M., Kindler, E., Emmerich, P. et al. Phylogenetic analysis of Puumala virus subtype Bavaria, characterization and diagnostic use of its recombinant nucleocapsid protein. Virus Genes 43, 177–191 (2011). https://doi.org/10.1007/s11262-011-0620-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11262-011-0620-x