Abstract
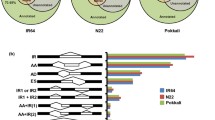
Osmotic stress induced by dehydration and salinity, is among the major abiotic stresses that adversely impacts crop productivity and plants often display cultivar-dependent response against osmotic imbalance. To better understand the molecular mechanisms underlying differential responses to dehydration, transcriptome changes of two contrasting wheat (Triticum aestivum L.) cultivars were evaluated in plants grown under unfavorable osmotic conditions. A total of 107 non-redundant transcripts were identified. Of these, most had unknown functions (31; ~30 %) signifying the existence of putative stress-specific genes in wheat, reported here for the first time. Upon comparing with previous transcriptomic studies, 43 (40 %) of the osmotically-responsive transcripts were found not to be documented. These new transcripts may therefore signify unexplored gene sources for specific responses towards short-term osmotic stress in wheat. Through macroarray analysis, 69 (~64 %) transcripts were found to be differentially expressed (≥3-fold) and expression of 14 transcripts (with known or unknown functions) was further confirmed by quantitative real time PCR. Expression analysis of the seven unknown transcripts also revealed their tissue- and stress-specific regulation. Comparative in silico mapping of these 107 wheat transcripts against available mapping data for rice (40; ~37 %), maize (34; ~32 %), and sorghum (33; ~31 %) revealed presence of wheat orthologous sequences in these cereal crops. This study provides an interesting account on several novel genes, besides those with known functions, which may regulate stress response dynamics and thus, may be used as potential candidates to improve stress adaptability through genetic and molecular studies.




Similar content being viewed by others
Change history
15 February 2024
This article has been retracted. Please see the Retraction Notice for more detail: https://doi.org/10.1007/s11240-024-02702-y
References
Baisakh N, Subudhi PK, Varadwaj P (2008) Primary responses to salt stress in a halophyte, smooth cordgrass (Spartina alterniflora Loisel.). Funct Integr Genomics 8:287–300
Bartels D, Sunkar R (2005) Drought and salt tolerance in plants. Crit Rev Plant Sci 24:23–58
Bartels D, Schneider K, Terstappen G, Piatkowski D, Salamini F (1990) Molecular cloning of abscisic acid-modulated genes which are induced during desiccation of the resurrection plant Craterostigma plantagineum. Planta 181:27–34
Boominathan P, Shukla R, Kumar A, Manna D, Negi D, Verma PK, Chattopadhyay D (2004) Long term transcript accumulation during the development of dehydration adaptation in Cicer arietinum. Plant Physiol 135:1–13
Dallaire S, Houde M, Gagne Y, Saini HS, Boileau S, Chevrier N, Sarhan F (1994) ABA and low temperature induce freezing tolerance via distinct regulatory pathways in wheat. Plant Cell Physiol 35:1–9
Eisen MB, Spellman PT, Brown PO, Botstein D (1998) Cluster analysis and display of genome-wide expression patterns. Proc Natl Acad Sci USA 95:14863–14868
Feuillet C, Keller B (2002) Comparative genomics in the grass family: molecular characterization of grass genome structure and evolution. Ann Bot 89:3–10
Finn RD, Mistry J, Tate J, Coggill P, Heger A, Pollington JE, Gavin OL, Gunesekaran P, Ceric G, Forslund K, Holm L, Sonnhammer EL, Eddy SR, Bateman A (2010) The Pfam protein families database. Nucleic Acids Res 38:D211–D222
Garg B, Jaiswal JP,Misra S, Tripathi BN,Prasad M (2012) A comprehensive study on dehydration-induced antioxidative responses during germination of Indian bread wheat (Triticum aestivum L. em Thell) cultivars collected from different agroclimatic zones. Physiol Mol Biol Plants. doi:10.1007/s12298-012-0117-7
Gasteiger E, Hoogland C, Gattiker A, Duvaud S, Wilkins MR, Appel RD, Bairoch A (2005) Protein identification and analysis tools on the ExPASy server. In: John MW (ed) The proteomics protocols handbook. Humana Press, New Jersey, pp 571–607
Gill BS, Appels R, Botha-Oberholster AM et al (2004) A workshop report on wheat genome sequencing: international genome research on wheat consortium. Genetics 168:1087–1096
Guan B, Jiang GQ, Wang YX, Wang ZC, Haxim Y, Bao Q, Hu YZ, Zhang FC, Wang Y (2010) Identification of differentially expressed transcripts involved in the salt-stress response of Salsola ferganica by suppression subtractive hybridization. Plant Cell Tissue Organ Cult 103:343–352
Gupta S, Kumari K, Sahu PP, Vidapu S, Prasad M (2012) Sequence-based novel genomic microsatellite markers for robust genotyping purposes in foxtail millet [Setaria italica (L.) P. Beauv.]. Plant Cell Rep 31:323–337
Guyot R, Yahiaoui N, Feuillet C, Keller B (2004) In silico comparative analysis reveals a mosaic conservation of genes within a novel colinear region in wheat chromosome 1AS and rice chromosome 5S. Funct Integr Genomics 4:47–58
Horton P, Park K-J, Obayashi T, Fujita N, Harada H, Adams-Collier CJ, Nakai K (2007) WoLF PSORT: protein localization predictor. Nucleic Acids Res 35:W585–W587
Jain D, Chattopadhyay D (2010) Analysis of gene expression in response to water deficit of chickpea (Cicer arietinum L.) varieties differing in drought tolerance. BMC Plant Biol. 10–24
Jayaraman A, Puranik S, Rai NK, Vidapu S, Sahu PP, Lata C, Prasad M (2008) cDNA-AFLP analysis reveals differential gene expression in response to salt stress in foxtail millet (Setaria italica L.). Mol Biotechnol 40:241–251
La Rota M, Sorrells ME (2004) Comparative DNA sequence analysis of mapped wheat ESTs reveals the complexity of genome relationships between rice and wheat. Funct Integr Genomics 4:34–46
Lata C, Prasad M (2012) Validation of an allele specific marker associated with dehydration stress tolerance in a core set of foxtail millet accessions. Plant Breed. doi:10.1111/j.1439-0523.2012.01983.x
Lata C, Sahu PP, Prasad M (2010) Comparative transcriptomic analysis of differential expressed genes in foxtail millet (Setaria italica L.) during dehydration stress. Biochem Biophys Res Commun 393:720–727
Letunic I, Doerks T, Bork P (2012) SMART 7: recent updates to the protein domain annotation resource. Nucleic Acids Res. doi:10.1093/nar/gkr931
Linder P (2006) Dead-box proteins: a family affair active and passive players in RNP-remodeling. Nucleic Acids Res 34:4168–4180
Liu L, Wang Y, Zeng Y, Haxim Y, Zhang F (2012a) Identification and characterization of differentially expressed genes in the halophyte Halostachys caspica under salt stress. Plant Cell Tissue Organ Cult 110:1–12
Liu W, Glunde K, Bhujwalla ZM, Raman V, Sharma A, Phang JM (2012b) Proline oxidase promotes tumor cell survival in hypoxic tumor microenvironments. Cancer Res 72:3677–3686
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using realtime quantitative PCR and the 2−∆∆Ct method. Methods 25:402–408
Lokko Y, Anderson JV, Rudd S, Horvath D, Mikel MA, Kim R, Liu L, Hernandez A, Dixon AGO, Ingelbrecht IL (2007) Characterization of an 18,166 EST dataset for cassava (Manihot esculenta Crantz) enriched for drought-responsive genes. Plant Cell Rep 26:1605–1618
Mare C, Mazzucotelli E, Crosatti C, Francia E, Stanca AM, Cattivelli L (2004) Hv-WRKY38: a new transcription factor involved in cold and drought-response in barley. Plant Mol Biol 55:399–416
Matsumoto T, Tanaka T, Sakai H, Amano N, Kanamori H, Kurita K, Kikuta A, Kamiya K, Yamamoto M, Ikawa H, Fujii N, Hori K, Itoh T, Sato K (2011) Comprehensive sequence analysis of 24,783 barley full-length cDNAs derived from 12 clone libraries. Plant Physiol 15:20–28
Mehta PA, Sivaprakash K, Parani M, Venkataraman G, Parida AK (2005) Generation and analysis of expressed sequence tags from the salt-tolerant mangrove species Avicennia marina (Forsk) Vierh. Theor Appl Genet 110:416–424
Mohammadi M, Kav NNV, Deyholos MK (2007) Transcriptional profiling of hexaploid wheat (Triticum aestivum L.) roots identifies novel, dehydration-responsive genes. Plant, Cell Environ 30:630–645
Moller S, Croning MDR, Apweiler R (2001) Evaluation of methods for the prediction of membrane spanning regions. Bioinformatics 17:646–653
Natarajan SK, Becker DF (2012) Role of apoptosis-inducing factor, proline dehydrogenase, and NADPH oxidase in apoptosis and oxidative stress. Cell Health Cytoskelet 4:11–27
Nayyar H (2003) Accumulation of osmolytes and osmotic adjustment in water-stressed wheat (Triticum aestivum L.) and maize (Zea mays) affected by calcium and its antagonists. Environ Exp Bot 50:253–264
Owttrim GW (2006) RNA helicases and abiotic stress. Crit Rev Nucleic Acids Res 34:3220–3230
Pandhare J, Donald SP, Cooper SK, Phang JM (2009) Regulation and function of proline oxidase under nutrient stress. J Cell Biochem 107:759–768
Puranik S, Jha S, Srivastava PS, Sreenivasulu N, Prasad M (2011) Comparative transcriptome analysis of contrasting foxtail millet cultivars in response to short-term salinity stress. J Plant Physiol 168:280–287
Sambrook J, Russell D (2001) Molecular cloning: a laboratory manual, 3rd edn. Cold Spring Harbour Laboratory Press, Cold Spring Harbour
Shinozaki K, Yamaguchi-Shinozaki K (2000) Molecular responses to dehydration and low temperature: differences and cross talk between two stress signaling pathways. Curr Opin Plant Biol 3:217–223
Singh NK, LaRosa PC, Handa AK, Hasegawa PM, Bressan RA (1987) Hormonal regulation of protein synthesis associated with salt tolerance in plant cells. Proc Natl Acad Sci USA 84:739–743
Stone SL, Williams LA, Farmer LM, Vierstra RD, Callis J (2006) KEEP ON GOING, a RING E3 ligase essential for Arabidopsis growth and development, is involved in abscisic acid signaling. Plant Cell 18:3415–3428
Tardif G, Kane NA, Adam H, Labrie L, Major G, Gulick P, Sarhan F, Laliberté JF (2007) Interaction network of proteins associated with abiotic stress response and development in wheat. Plant Mol Biol 63:703–718
Tusnády GE, Simon I (2001) The HMMTOP transmembrane topology prediction server. Bioinformatics 17:849–850
Yang G, Zou H, Wu Y, Liu H, Yuan Y (2011) Identification and characterization of candidate genes involved in chilling responses in maize (Zea mays L.). Plant Cell Tissue Organ Cult 106:127–141
Zhang X, Garreton V, Chua NH (2005) The AIP2 E3 ligase acts as a novel negative regulator of ABA signaling by promoting ABI3 degradation. Genes Dev 19:1532–1543
Zhang Y, Yan C, Li Y, Zheng N, Chen H, Zhao Q, Gao T, Guo H, Xie Q (2007) SDIR1 is a RING finger E3 ligase that positively regulates stress responsive abscisic acid signaling in arabidopsis. Plant Cell 19:1912–1929
Zhu X, Gong H, Chen G, Wang S, Zhang C (2005) Different solute levels in two spring wheat cultivars induced by progressive field water stress at different developmental stages. J Arid Environ 62:1–14
Acknowledgments
We are thankful to Vice-Chancellor of the Banasthali University and Head, Department of Biotechnology, Jamia Hamdard, New Delhi, India for providing necessary facilities. Ms Swati Puranik acknowledges the award of Senior Research Fellowship from the Council of Scientific and Industrial Research, New Delhi.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
About this article
Cite this article
Garg, B., Puranik, S., Misra, S. et al. RETRACTED ARTICLE: Transcript profiling identifies novel transcripts with unknown functions as primary response components to osmotic stress in wheat (Triticum aestivum L.). Plant Cell Tiss Organ Cult 113, 91–101 (2013). https://doi.org/10.1007/s11240-012-0254-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11240-012-0254-2