Abstract
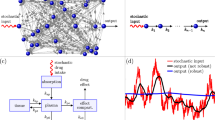
In the first part of this invited paper we review the role of both extrinsic and intrinsic stochasticity in shaping the dynamics of biomolecular networks. In particular, we stress the use of bounded stochastic processes as a model of extrinsic random perturbations. In the second part, we propose three examples of molecular circuits under the influence of external fluctuations modeled by means of bounded noises. The first two examples involve the linear decay of a protein with, respectively, large or low number of molecules, so to stress different modeling approaches. The third example concerns the spatio-temporal dynamics of proteins determining the chemotaxis-driven polarization of a cell. In these examples one can observe phenomena that are dependent on the specific class of employed bounded noise.





Similar content being viewed by others
References
Alberts B, Johnson A, Lewis J, Raff M, Roberts K, Walter P (2009) Molecular biology of the cell, 5th ed. Garland, New York
Alon U (2006) An introduction to systems biology: design principles of biological circuits. Chapman & Hall/CRC Mathematical & Computational Biology, Boca Raton, FL
Angeli D, Ferrell JE Jr, Sontag ED (2004) Detection of multistability, bifurcations, and hysteresis in a large class of biological positive-feedback systems. PNAS 101(7):1822–1827
Arnold L (1989) Random dynamical systems. Springer, Berlin
Arnold L (1998) Random dynamical systems. Springer, Berlin
Becskei A, Serrano L (2000) Engineering stability in gene networks by autoregulation. Nature 405:590–593
Becskei A, Kaufmann BB, van Oudenaarden AE (2000) Contributions of low molecule number and chromosomal positioning to stochastic gene expression. Nat Genet 37:937–944
Bobryk RV, Chrzeszczyk A (2005) Transitions induced by bounded noise. Phys A 358(2–4):263
Bobryk RV, Chrzeszczyk A (2008) Transitions in a duffing oscillator excited by random noise. Nonlinear Dyn 51:541
Borland L (1998) Ito–Langevin equations within generalized thermostatistics. Phys Lett A f245(1–2):67
Cai CQ, Lin YK (1996) Generation of non-gaussian stationary stochastic processes. Phys Rev E 54:299
Cai L, Friedman N, Xie XS (2006) Stochastic protein expression in individual cells at the single molecule level. Nature 440:358–362
Caravagna G, Mauri G, d’Onofrio A (2013) The interplay of intrinsic and extrinsic bounded noises in biomolecular networks. PLoS One 8:e51174
Caravagna G, Mauri G, d’Onofrio A (2013) Bounded extrinsic noises affecting biochemical networks with low molecule numbers. Chapter in d’Onofrio A (ed) Bounded noises in physics, biology and engineering, Birkauser, Verlag. ISBN 978-1-4614-7348-8
Caravagna G, Mauri G, d’Onofrio A (2013) NoisySIM: exact simulation of stochastic chemically reacting systems with extrinsic noises. In proceedings of the Symposium on theory of modeling and simulation vol 12, pp 1–6. Society for Computer Simulation International, San Diego, CA
Chang HH, Oh PY, Ingber DE, Huang S (2006) Stochastic approaches for systems biology. BMC Cell Biol 7:11–23
Cinquin O, Demongeot J (2005) High-dimensional switches and the modelling of cellular differentiation. J Theor Biol 233:391–411
deFranciscis S, d’Onofrio A (2012) Spatiotemporal bounded noises and transitions induced by them in solutions of the real Ginzburg–Landau model. Phys Rev E 86:021118
deFranciscis S, d’Onofrio A (2013) Cellular polarization: interaction between extrinsic bounded noises and the wave-pinning mechanism. Phys Rev E 88:032709
deFranciscis S, d’Onofrio A (2013) Spatio-temporal Sine–Wiener bounded noise and its effect on Ginzburg–Landau model. Nonlinear Dyn 74:607
Detwiler PB, Ramanathan S, Sengupta A, Shraiman BI (2000) Engineering aspects of enzymatic signal transduction: photoreceptors in the retina. Biophys J 79:2801–2817
Dimentberg M (1988) Statistical dynamics of nonlinear and time-varying systems. Research Studies Press, Baldock
d’Onofrio A (2013) Multifaceted aspects of the kinetics of immunoevasion from tumor dormancy. In: Enderling H, Almog N and Hlatky L (eds.) Systems biology of tumor dormancy, advances in experimental medicine and biology, vol 734, Springer, Berlin, p 111
d’Onofrio A (ed) (2013) Bounded noises in physics, biology and engineering, Birkauser, Basel-Boston. ISBN 978-1-4614-7348-8.
d’Onofrio A (2010) Bounded-noise-induced transitions in a tumor-immune system interplay. Phys Rev E 81:021923
d’Onofrio A, Gandolfi A (2010) Resistance to antitumor chemotherapy due to bounded-noise-induced transitions. Phys Rev E 82:061901
Eldar A, Elowitz MB (2010) Functional role for noise in genetic circuits. Nature 467:167–173
Elowitz MB, Levine AJ, Siggia ED, Swain PS (2002) Stochastic gene expression in a single cell. Science 298:1183–1186
Gamba A, de Candia A, Di Talia S, A Coniglio A, Bussolino F, Serini G (2005) Diffusion-limited phase separation in eukaryotic chemotaxis. PNAS 102(47):16927
García-Ojalvo J, Sancho JM, Ramírez-Piscina L (1992) Generation of spatiotemporal colored noise. Phys Rev A 46:4670
García-Ojalvo J, Sancho JM, Ramírez-Piscina L (1992) A nonequilibrium phase transition with colored noise. Phys Lett A 168(1):35–39
Garcia-Ojalvo J, Sancho JM, Ramirez-Piscina I (1992) Generation of spatiotemporal colored noise. Phys Rev E 46:4670
Gardiner CW (1985) Handbook of stochastic methods, 2nd edn. Springer, Berlin
Gardner TR, Cantor CR, Collins JJ (2000) Construction of a genetic toggle switch in escherichiacoli. Nature 403:339–342
Ghaemmaghami S, Huh W, Bower K, Howson RW, Belle A, Dephoure N, O’Shea EK, Weissman JS (2003) Global analysis of protein expression in yeast. Nature 425:737–743
Gierer A, Meinhardt H (1972) A theory of biological pattern formation. Kybernetik 12:30–39
Gillespie DT (1976) A general method for numerically simulating the stochastic time evolution of coupled chemical reactions. J Comp Phys 22(4):403–434
Gillespie DT (1977) Exact stochastic simulation of coupled chemical reactions. J Phys Chem 81:2340–2361
Gillespie DT (1980) Approximating the master equation by Fokker–Planck-type equations for single-variable chemical systems. J Phys Chem 72:5363–5371
Gillespie DT (2000) The chemical Langevin equation. J Phys Chem 113:297–306
Glass L, Kauffman SA (1968) Logical analysis of systems comprising feedback loops. J Theor Biol 39:103–129
Grabert H, Hänggi P, Oppenheim I (1983) Fluctuations in reversible chemical reactions. Phys A 117:300–316
Graudenzi A, Caravagna G, De Matteis G, Antoniotti M (2014) Investigating the relation between stochastic differentiation and homeostasis in intestinal crypts via multiscale modeling. bioRxv, http://biorxiv.org/content/early/2013/11/25/000927
Griffith JS (1968) Mathematics of cellular control processes ii positive feedback to one gene. J Theor Biol 20:209–216
Guo W, Du LC, Mei DC (2012) Transitions induced by time delays and cross-correlated Sine–Wiener noises in a tumor-immune system interplay. Phys A 391:1270–1280
Hasty J, Pradines J, Dolnik M, Collins JJ (2000) Noise-based switches and amplifiers for gene expression. PNAS 97(5):2075–2080
Homburg AJ, Young TR, Gharaei M (2013) Bifurcations of random differential equations with bounded noise. In d’Onofrio A (ed) Bounded noises in physics, biology and engineering, Birkauser, Verlag. ISBN 978-1-4614-7348-8
Horsthemke W, Lefever R (2006) Noise-induced transitions: theory and applications in physics, chemistry, and biology, Series in Synergetics Springer, Berlin
Iglesias PA, Devreotes PN (2008) Navigating through models of chemotaxis. Curr Opin Cell Biol 20:35–40
Iglesias PA, Ingalls PB (2010) Control theory and systems biology. MIT Press, Cambridge
Jung P, Hänggi P (1987) Dynamical systems: a unified colored-noise approximation. Phys Rev A 35:4464
Kauffman SA (1969) Metabolic stability and epigenesis in randomly constructed genetic nets. J Theor Biol 22:437–467
Kramer BP, Fussenegger M (2005) Hysteresis in a synthetic mammalian gene network. PNAS 102:9517–9522
Lestas I, Vinnicombe G, Paulsson J (2010) Fundamental limits on the suppression of molecular fluctuations. Nature 467:174–178
Losick R, Desplan C (2008) Stochasticity and cell fate. Science 320:65–68
Macieira-Coelho A (2007) Asymmetric cell division. Springer, Berlin
Mandelbrot BB (1963) The variation of certain speculative prices. J Bus (Chicago) 36:394–419
Markevich NI, Hoek JB, Kholodenko BN (2004) Signaling switches and bistability arising from multisite phosphorylation in protein kinase cascades. J Cell Biol 164:353–359
Meinhardt H (1999) Orientation of chemotactic cells and growth cones: models and mechanisms. J Cell Sci 17:2867
Mori Y, Jilkine A, Edelstein-Keshet L (2008) Wave-pinning and cell polarity from a bistable reaction-diffusion system. Biophys J 94:3684
Murray JD (2002) Mathematical biology. Springer, New York
Onsum MD, Rao CV (2009) Calling heads from tails: the role of mathematical modeling in understanding cell polarization. Curr Opin Cell Biol 21(1):74
Paulsson BO (2011) Systems biology simulation of dynamic network states. Cambridge University Press, Cambridge
Rao CV, Wolf D, Arkin AP (2002) Control, exploitation and tolerance of intracellular noise. Nature 420:231–237
Ridolfi L, D’Odorico P, Laio F (2011) Noise-induced phenomena in the environmental sciences. Cambridge University Press, Cambridge
Rigney DR, Schieve WC (1977) Stochastic model of linear, continuous protein-synthesis in bacterial populations. J Theor Biol 69:761–766
Samoilov M, Plyasunov S, Arkin AP (2005) Stochastic amplification and signaling in enzymatic futile cycles through noise-induced bistability with oscillations. PNAS 102(7):2310–2315
Sanft KR, Gillespie DT, Petzold LR (2011) Legitimacy of the stochastic michaelis-menten approximation. IET Sys Bio 5(1):58–69
Semplice M, Veglio A, Naldi G, Serini G, Gamba A (2012) A bistable model of cell polarity. PLoS One 7:e30977
Shahrezaei V, Ollivier JF, Swain PS (2008) Colored extrinsic fluctuations and stochastic gene expression. Mol Sys Biol 4:196
Siegal-Gaskins D, Grotewold E, Smith GD (2009) The capacity for multistability in small gene regulatory networks. BMC Sys Biol 3:96
Simon Z (1965) Multi-steady-state model for cell differentiation. J Theor Biol 8:258–263
Sugita M (1964) Functional analysis of chemical systems in vivo using a logical circuit equivalent ii the idea of a molecular automaton. J Theor Biol 4:437–467
Thattai M, Van Oudenaarden A (2001) Attenuation of noise in ultrasensitive signaling cascades. Biophys J 82:2943–2950
Thattai M, Van Oudenaarden A (2001) Intrisic noise in gene regulatory networks. PNAS 98:8614–8619
Thomas R (1978) Logical analysis of systems comprising feedback loops. J Theor Biol 73:631–656
Tomas R, d’Ari R (1990) Biological feedbacks. Chapman & Hall/CRC Mathematical & Computational Biology, Boca Raton, FL
Turing AM (1952) The chemical basis of morphogenesis. Phil Trans R Soc Lond B 237(641):37–72
Tze-Leung T, Mahesci N (2010) Stochasticity and cell fate. Science 327:1142–1145
Ullah M, Wolkhenauer O (2011) Stochastic approaches for systems biology. Springer, Berlin
Walther GR, Marée AF, Edelstein-Keshet L, Grieneisen VA (2012) Deterministic versus stochastic cell polarisation through wave-pinning. Bull Math Biol 74:2570
Wang L, Walker BL, Iannaccone S, Bhatt D, Kennedy PJ, Tse WT (2009) Bistable switches control memory and plasticity in cellular differentiation. PNAS 106(16):6638–6643
Wilkinson DJ (2006) Stochastic modelling for systems biology. Chapman & Hall/CRC Mathematical & Computational Biology, Boca Raton, FL
Wio HS, Lindenberg K (2003) Modern challenges in statistical mechanics. In proceedings of the AIP conference vol 658(1)
Wio Hs, Deza RR (2013) Noise-induced phenomena: effects of noises based on Tsallis statistics. In d’Onofrio A (ed.) Bounded noises in physics, biology and engineering, Birkauser, Verlag. ISBN 978-1-4614-7348-8
Wio H, Toral R (2004) Effect of non-Gaussian noises in a noise induced transition. Phys D 193:161
Xiong W, Ferrell JE Jr (2003) A positive-feedback-based bistable ’memory module’ that governs a cell fate decision. Nature 426:460–465
Yamada T, Bork P (2009) Evolution of biomolecular networks: lessons from metabolic and protein interactions. Nat Rev Mol Cell Bio 10:791–803
Zhdanov VP (2011) Interplay of bistable kinetics of gene expression during cellular growth. Phys A 390:57
Zhdanov VP (2012) Periodic perturbation of genetic oscillations. Chaos, Solitons & Fractals 45:577–587
Zhu WQ, Cai GQ (2013) On Bounded stochastic processes. In d’Onofrio A (ed) bounded noises in physics, biology and engineering, Birkauser, Verlag. ISBN 978-1-4614-7348-8
Acknowledgments
The authors would like to thank the anonymous referees for invaluable comments which improved the paper. AdO wishes to thank the organizers of Wivace (Italian Workshop on Artificial Life and Evolutionary Computation) for the invitation to present these topics to the 2013 edition of the workshop and in this special issue.
Author information
Authors and Affiliations
Corresponding author
Additional information
The research of A. d’Onofrio and S. de Franciscis has been done in the framework of the Integrated Project “P-medicine—from data sharing and integration via VPH models to personalized medicine” (Project ID: 270089), which is partially funded by the European Commission under the 7th framework program.
Rights and permissions
About this article
Cite this article
de Franciscis, S., Caravagna, G. & d’Onofrio, A. Bounded noises as a natural tool to model extrinsic fluctuations in biomolecular networks. Nat Comput 13, 297–307 (2014). https://doi.org/10.1007/s11047-014-9424-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11047-014-9424-y