Abstract
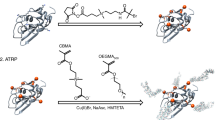
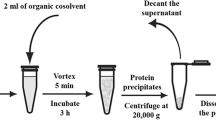
Protein precipitation using non-charged and charged polymers is a common method for protein purification, gaining broader interest among manufacturers in downstream processing. While during polymer- surface interactions, the formation of loops, tails and trains has been known for quite a long time, details of polymer conformation and chain length, interacting with the protein during protein precipitation are not fully discovered. Our research presents a more profound understanding of polymer-protein interaction, combining fluorescence and infrared spectroscopic measurements of proteins and polymer standards with well defined chain length to confirm different models of protein-polymer interaction. Lysozyme, chymotrypsinogen A, myoglobin and a monoclonal antibody, all of different molecular weight, isoelectric point and charge distribution at the protein surface, were used for protein-polymer precipitation. The use of polymers of various charge density and chain length showed that the required polymer length per precipitated protein (Ldef) is up to 25-times larger than the diameter of the corresponding protein, depending on the surface charge distribution of the protein, and its isoelectric point, as well as the charge density of the polymer. Our results support proposed mechanisms of polymer wrapping and loop formation for optimal charge neutralization during complexation and imply interaction of several polymer chains per precipitated protein molecule. Electrophoretic light scattering showed a qualitative correlation of the zeta potential of analyzed polymers with their corresponding Ldef values. Comparing protein precipitation behavior of long and short polymer chains, the latter exhibited reduced precipitation efficiency, visible as elevated Ldef.





Similar content being viewed by others
References
Muzzarelli RAA, Weckx M, Fillipini O, Lough C (1989) Carbohydr Polym 11:293–296
Bolto B, Gregory J (2007) Water Res 41:2301–2324
Shahidi F, Synowiecki J (1991) J Agric Food Chem 39:1527–1532
Sudharshan NR, Hoover DG, Knorr D (1992) Food Biotechnol 6:257–272
Papineau AM, Hoover DG, Knorr D, Farkas DF (1991) Food Biotechnol 5:45–57
Gross RA, Kalra B (2002) Science 297:803–807
Allen TM, Cullis PR (2004) Science 303:1818–1822
Gillies ER, Fréchet JM (2005) J Drug Discov Today 10:35–43
Schmaljohann D (2006) Adv Drug Deliv Rev 58:1655–1670
Kokufuta E (1992) Prog Polym Sci 17:647–697
Dubin PL, Gao J, Mattison K (1994) Sep Purif Methods 23:1–16
Ovenden C, Xiao H (2002) Colloids Surf A 197:225–234
Svec F, Fréchet JM (1992) J Anal Chem 64:820–822
Barrande M, Beurroies I, Denoyel R, Tatárová I, Gramblička M, Polakovič M, Joehnck M, Schulte M (2009) J Chromatogr A 1216:6906–6916
Thömmes J, Etzel M (2007) Biotechnol Prog 23:42–45
Low D, O’Leary R, Pujar NS (2007) J Chromatogr B 848:48–63
Capito F, Skudas R, Stanislawski B, Kolmar H (2013) Colloid Polym Sci 1-11
Cooper CL, Goulding A, Kayitmazer AB, Ulrich S, Stoll S, Turksen S, Dubin PL (2006) Biomacromolecules 7:1025–1035
Cooper CL, Dubin PL, Kayitmazer AB, Turksen S (2005) Curr Opin Colloid Interface Sci 10:52–78
Benmansour K, Medjahed K, Tennouga L, Mansri A (2003) Eur Polym J 39:1443–1449
Mattison KW, Dubin PL, Brittain IJ (1998) J Phys Chem B 102:3830–3836
Mattison KW, Brittain IJ, Dubin PL (1995) Biotechnol Prog 11:632–637
Park J, Muhoberac BB, Dubin PL, Xia J (1992) Macromolecules 25:290–295
Rawat K, Pathak J, Bohidar HB (2013) Phys Chem Chem Phys 15:12262–12273
Hattori T, Bat-Alder S, Kato R, Bohidar HB, Dubin PL (2005) Anal Chem 342:229–236
Stoll S, Chodanowski P (2002) Macromolecules 35:9556–9562
Xia J, Dubin PL, Kim Y, Muhoberac BB, Klimkowski VJ (1993) J Phys Chem 97:4528–4534
Sato T, Mattison KW, Dubin PL, Kamachi M, Morishima Y (1998) Langmuir 14:5430–5437
Yamaguchi K, Hachiya K, Moriyama Y, Takeda K (1996) J Coll Interf Sci 179:249–254
Gummel J, Cousin F, Boué F (2008) Macromolecules 41:2898–2907
Bohidar H, Dubin PL, Majhi PR, Tribet C, Jaeger W (2005) Biomacromolecules 6:1573–1585
Borrega R, Tribet C, Audebert R (1999) Macromolecules 32:7798–7806
Nguyen TT, Shklovskii BI (2001) J Chem Phys 114:5905–5916
Ulrich S, Laguecir A, Stoll S (2005) Macromolecules 38:8939–8949
Brynda M, Chodanowski P, Stoll S (2002) Colloid Polym Sci 280:789–797
Schiessel H, Rudnick J, Bruinsma R, Gelbart WM (2000) Europhys Lett 51:237
Akinchina A, Linse P (2002) Macromolecules 35:5183–5193
Laguecir A, Stoll S, Kirton G, Dubin PL (2003) J Phys Chem B 107:8056–8065
Chodanowski P, Stoll S (2001) J Chem Phys 115:4951–4960
Chodanowski P, Stoll S (2001) Macromolecules 34:2320–2328
Carlsson F, Malmsten M, Linse P (2003) J Am Chem Soc 125:3140–3149
Skepö M, Linse P (2003) Macromolecules 36:508–519
Jy S (1994) Glatz CE In Macromolecular Complexes in Chemistry and Biology; Dupin PL et al., Eds. Springer-Verlag, Berlin
Houska M, Brynda E, Bohata K (2004) J Coll Interf Sci 273:140–147
Pawar N, Bohidar HB (2010) Phys Rev E 82:36107
von Goeler F, Muthukumar M (1994) J Chem Phys 100:7796–7803
Kong CY, Muthukumar M (1998) J Chem Phys 109:1522–1527
Hattori T, Hallberg R, Dubin PL (2000) Langmuir 16:9738–9743
Singh J, Dutta PK (2009) J Polym Res 16:231–238
Kiefer F, Arnold K, Künzli M, Bordoli L, Schwede T (2009) Nucleic Acids Res 37:387–392
Arnold K, Bordoli L, Kopp J, Schwede T (2006) Bioinformatics 22:195–201
Peitsch MC (1995) Bio/Technology 13:658–660
Olsson MH, Søndergaard CR, Rostkowski M, Jensen JH (2011) J Chem Theory Comput 7:525–537
Bas DC, Rogers DM, Jensen JH (2008) Proteins Struct Funct Bioinforma 73:765–783
Dolinsky TJ, Czodrowski P, Li H, Nielsen JE, Jensen JH, Klebe G, Baker NA (2007) Nucleic Acids Res 35:522–525
Li H, Robertson AD, Jensen JH (2005) Proteins Struct Funct Bioinforma 61:704–721
Dolinsky TJ, Nielsen JE, McCammon JA, Baker NA (2004) Nucleic Acids Res 32:665–667
Baker NA, Sept D, Joseph S, Holst MJ, McCammon JA (2001) Proc Natl Acad Sci U S A 98:10037–10041
Yang Z, Lasker K, Schneidman-Duhovny D, Webb B, Huang CC, Pettersen EF, Goddard TD, Meng EC, Sali A, Ferrin TE (2012) J Struct Biol 179:269–278
Pettersen EF, Goddard TD, Huang CC, Couch GS, Greenblatt DM, Meng EC, Ferrin TE (2004) J Comput Chem 25:1605–1612
Roth CM, Lenhoff AM (1995) Langmuir 11:3500–3509
Kisler JM, Stevens GW O, Connor AJ (2001) Mat Phy Mech 4:89–93
Papadopoulos S, Jürgens KD, Gros G (2000) Biophys J 79:2084–2094
Striemer CC, Gaborski TR, McGrath JL, Fauchet PM (2007) Nature 445:749–753
McDonald P, Victa C, Carter‐Franklin JN, Fahrner R (2009) Biotechnol Bioeng 102:1141–1151
Izumi T, Hirata M, Kokufuta E, Cha HJ, Frank CW (1994) J Macromol Sci Pure Appl Chem 31:31–37
Adamczyk Z, Zembala M, Warszýnski P, Jachimska B (2004) Langmuir 20:10517–10525
Donath E, Walther D, Shilov VN, Knippel E, Budde A, Lowack K, Helm CA, Möhwald H (1997) Langmuir 13:5294–5305
Tricot M (1984) Macromolecules 17:1698–1704
Le Bret M (1982) J Chem Phys 76:6243–6255
Messina R, Holm C, Kremer K (2003) Langmuir 19:4473
Acknowledgments
The authors thank Merck KGaA for financial and material support. Thanks to Johann Bauer and Stephan von der Au, both Merck KGaA for helpful advice and supplying the Zetasizer Nano. Part of this work was performed within the frame of the project BIOPUR and IOLIPRO, funded by the German Federal Ministry of Education and Research (BMBF). We thank Charly Hohlschuh, Merck KGaA, for language assistance and proof reading.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(PDF 354 kb)
Rights and permissions
About this article
Cite this article
Capito, F., Kolmar, H., Stanislawski, B. et al. Required polymer lengths per precipitated protein molecule in protein-polymer interaction. J Polym Res 21, 346 (2014). https://doi.org/10.1007/s10965-013-0346-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10965-013-0346-7