Abstract
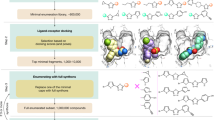
An algorithm is presented for the estimation of molecular properties over a library built around a scaffold, which has N sites for functionalization with M i moieties at the ith scaffold site, corresponding to a library of \({\prod_{i=1}^N M_i}\) molecules. The algorithm relies on a series of operations involving (i) synthesis and property measurement of a minimal number of T randomly sampled members of the library, (ii) expression of the observed property in terms of a high-dimensional model representation (HDMR) of the moiety → property map, (iii) optimization of the ordered sequence of moieties on each site to regularize the HDMR map and (iv) interpolation using the map to estimate the properties of as yet unsynthesized compounds. The set of operations is performed iteratively aiming to reach convergence of the predictive HDMR map with as few synthesized samples as possible. Through simulation, the number T of required random molecular samples is shown to scale very favorably with \({T < < \prod^N_{i=1} M_i}\) for cases up to N = 20 and M i = 20. For example, high estimation quality was attained for simulated libraries with T ~ 5,000 sampled compounds for a library of 2012 members and T ~ 12,500 sampled compounds for a library of 2020 members. The algorithm is based on the assumption that a systematic pattern exists in the moiety → property map provided that the moieties are optimally ordered on the scaffold sites within the context of HDMR. The overall procedure is referred to as the substituent reordering HDMR algorithm (SR-HDMR). The technique was also successfully tested with laboratory data for estimating C13-NMR shifts in a tri-substituted benzene library and for lac operon repression binding.
Similar content being viewed by others
References
KnowItAll Informatics System 8.0, KnowItAll U Edition. Published by the Informatics Division of Bio-Rad Laboratories, Inc
JSci—A Science API for Java (2009). http://jsci.sourceforge.net/
Bicerano J.: Prediction of Polymer Properties. Marcel Dekker, New York, NY (2002)
Chatterjee S., Hadi A.S.: Sensitivity Analysis in Linear Regression. Wiley, New York, NY (1988)
Clark M.: Generalized fragment-substructure based property prediction method. J. Chem. Inf. Model. 45(1), 30–38 (2005)
Eilers P., Marx B.D.: Flexible smoothing with b-splines and penalties. Stat. Sci. 11(2), 89–121 (1996)
Gasteiger, J., Engel, T. (eds): Chemoinformatics. Wiley-VCH, Weinheim (2003)
Hastie T., Tibshirani R., Friedman J.: The Elements of Statistical Learning. Springer, New York, NY (2009)
M. Held, R.M. Karp, A dynamic programming approach to sequencing problems. in Proceedings of the 1961 16th ACM National Meeting (ACM, New York, NY, USA, 1961), pp. 71.201–71.204. doi:10.1145/800029.808532
Jorgensen W.L.: The many roles of computation in drug discovery. Science 303(5665), 1813–1818 (2004)
P.A. DiMaggio Jr., S.R. McAllister, C.A. Floudas, X.J. Feng, J.D. Rabinowitz, H.A. Rabitz, Optimal methods for re-ordering data matrices in systems biology and drug discovery applications. in BIOMAT 2007: International Symposium on Mathematical and Computational Biology, (2008)
Lau H.T.: A Java Library of Graph Algorithms and Optimization. Discrete Mathematics and its Applications. Chapman & Hall, CRC, London (2007)
Leach A.R., Gillet V.J.: An Introduction to Chemoinformatics. Springer, The Netherlands (2007)
N. Lehming, Regeln fur protein/dna-erkennung. PhD thesis, Universitat zu Koln (1990)
Lehming N., Sartorius J., Kisters-Woike B., von Wilcken-Bergmann B., Muller-Hiller B.: Mutant lac repressors with new specificities hint at rules for protein—dna recognition. EMBO J. 9(3), 615–621 (1990)
G. Li, H. Rabitz, P.E. Yelvington, O.O. Oluwole, F. Bacon, C.E. Kolb, J. Schoendorf, Global sensitivity analysis for systems with independent and/or correlated inputs. J. Phys. Chem. A 114(19), 6022–6032 (2010). doi:10.1021/jp9096919. http://pubs.acs.org/doi/abs/10.1021/jp9096919
Li G., Rosenthal C., Rabitz H.: High dimensional model representation. J. Phys. Chem. A 105(33), 7765–7777 (2001)
Li G., Wang S.W., Rabitz H.: Practical approaches to construct RS-HDMR component functions. J. Phys. Chem. A 106, 8721–8733 (2002)
Liang F., Feng X.J., Lowry M., Rabitz H.: Maximal use of minimal libraries through the adaptive substituent reordering algorithm. J. Phys. Chem. B 109, 5842–5854 (2003)
S.R. McAllister, X.J. Feng Jr., P.A. DiMaggio, C.A. Floudas, J.D. Rabinowitz, H. Rabitz, Descriptor-free molecular discovery in large libraries by adaptive substituent reordering. Bioorg. Med. Chem. Lett. 18(22), 5967–5970 (2008)
Padberg M.W., Grotschel M.: Polyhedral computations. In: Lawler, E.L., Lenstra, J.K., Kan, A.H.G.R., Shmoys, D.B. (eds) The Traveling Salesman Problem, Wiley, Chichester (1985)
Prenter P.M.: Splines and Variational Methods. Wiley, New York, NY (1975)
Ringuest J.L.: Multiobjective Optimization: Behavioral and Computational Considerations. Kluwer, Boston, MA (1992)
Shenvi N., Geremia J.M., Rabitz H.: Substituent ordering and interpolation in moleular library optimization. J. Phys. Chem. A 107(12), 2066–2074 (2003)
Shorter J.A., Ip P.C., Rabitz H.: An efficient chemical kinetics solver using high dimensional model representation. J. Phys. Chem. A 103, 7192–7198 (1999)
Wang S., Jaffe P.R., Li G., Wang S.W., Rabitz H.A.: Simulating bioremediation of uranium-contaminated aquifers; uncertainty assessment of model parameters. J. Contam. Hydrol. 64(3–4), 283–307 (2003)
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Izmailov, S., Feng, X., Li, G. et al. A scalable algorithm for molecular property estimation in high dimensional scaffold-based libraries. J Math Chem 50, 1765–1790 (2012). https://doi.org/10.1007/s10910-012-0005-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10910-012-0005-y