Abstract
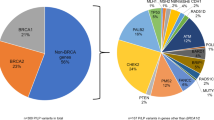
Since the 2013 Supreme Court ruling on BRCA1/BRCA2 patenting, hereditary cancer gene panels now include BRCA1 and BRCA2, making these panels an option for first-tier testing. However, questions remain about the clinical utility and implications of these panels for medical management with inclusion of genes of unknown to moderate penetrance. To better understand how use of these panels affected our practice, we reviewed patients who underwent testing in our clinic from July 1, 2013 through May 23, 2014. Indications for testing included personal and/or family history of breast and/or ovarian cancer. A total of 136 patients underwent panel testing via a single commercial laboratory; 12 (8.8 %) patients were positive for a pathogenic or likely pathogenic mutation (four BRCA2 mutations, two TP53 mutations, one CDH1 mutation, two ATM mutations, and one patient each with a CHEK2, NBN, or PALB2 mutation). Of these positive patients, 100 % met the National Comprehensive Cancer Network (NCCN) guidelines for Hereditary Breast and Ovarian Cancer genetic testing (2.2014). Mutations in seven of twelve (58 %) patients led to changes in medical management; three of seven (43 %) had a non-BRCA1 or BRCA2 gene mutation. Our findings suggest that there is clinical utility of panels that include genes of unknown to moderate penetrance.
Similar content being viewed by others
References
Breast Cancer Linkage, C. (1999). Cancer risks in BRCA2 mutation carriers. Journal of the National Cancer Institute, 91(15), 1310–1316.
Chen, S., & Parmigiani, G. (2007). Meta-analysis of BRCA1 and BRCA2 penetrance. Journal of Clinical Oncology, 25(11), 1329–1333.
Couch, F. J., Nathanson, K. L., & Offit, K. (2014). Two decades after BRCA: setting paradigms in personalized cancer care and prevention. Science, 343(6178), 1466–1470.
Daly, M. B., Pilarski, R., Axilbund, J. E., Buys, S. S., Crawford, B., Friedman, S., et al. (2014). Genetic/familial high-risk assessment: breast and ovarian, version 1.2014. Journal of the National Comprehensive Cancer Network, 12(9), 1326–1338.
Debniak, T., Gorski, B., Cybulski, C., Jakubowska, A., Kurzawski, G., Lener, M., et al. (2003). Germline 657del5 mutation in the NBS1 gene in patients with malignant melanoma of the skin. Melanoma Research, 13(4), 365–370.
Doherty, J., Bonadies, D. C., & Matloff, E. T. (2015). Testing for hereditary breast cancer: panel or targeted testing? Experience from a clinical cancer genetics practice. Journal of Genetic Counseling, 24(4), 683–687.
Domchek, S. M., Bradbury, A., Garber, J. E., Offit, K., & Robson, M. E. (2013). Multiplex genetic testing for cancer susceptibility: out on the high wire without a net? Journal of Clinical Oncology, 31(10), 1267–1270.
Easton, D. F., Pharoah, P. D., Antoniou, A. C., Tischkowitz, M., Tavtigian, S. V., Nathanson, K. L., et al. (2015). Gene-panel sequencing and the prediction of breast-cancer risk. The New England Journal of Medicine, 372(23), 2243–2257.
Friedman, L. S., Ostermeyer, E. A., Szabo, C. I., Dowd, P., Lynch, E. D., Rowell, S. E., et al. (1994). Confirmation of BRCA1 by analysis of germline mutations linked to breast and ovarian cancer in ten families. Nature Genetics, 8(4), 399–404.
Gabai-Kapara, E., Lahad, A., Kaufman, B., Friedman, E., Segev, S., Renbaum, P., et al. (2014). Population-based screening for breast and ovarian cancer risk due to BRCA1 and BRCA2. Proceedings of the National Academy of Sciences of the United States of America, 111(39), 14205–14210.
Genetic/Familial High-Risk Assessment: Breast and Ovarian Version 2.2014. (2014). National Comprehensive Cancer Network. Retrieved from www.nccn.org
Gumaste, P. V., Penn, L. A., Cymerman, R. M., Kirchhoff, T., Polsky, D., & McLellan, B. (2015). Skin cancer risk in BRCA1/2 mutation carriers. The British Journal of Dermatology, 172(6), 1498–1506.
Katz, S. J., Kurian, A. W., & Morrow, M. (2015). Treatment decision making and genetic testing for breast cancer: mainstreaming mutations. JAMA, 314(10), 997–998.
Kean, S. (2014). Breast cancer. The 'other' breast cancer genes. Science, 343(6178), 1457–1459.
King, M. C. (2014). “the race” to clone BRCA1. Science, 343(6178), 1462–1465.
Kurian, A. W., Hare, E. E., Mills, M. A., Kingham, K. E., McPherson, L., Whittemore, A. S., et al. (2014). Clinical evaluation of a multiple-gene sequencing panel for hereditary cancer risk assessment. Journal of Clinical Oncology, 32(19), 2001–2009.
Rainville, I. R., & Rana, H. Q. (2014). Next-generation sequencing for inherited breast cancer risk: counseling through the complexity. Current Oncology Reports, 16(3), 371.
Siegel, R. L., Miller, K. D., & Jemal, A. (2015). Cancer statistics, 2015. CA: a Cancer Journal for Clinicians, 65(1), 5–29.
Steffen, J., Varon, R., Mosor, M., Maneva, G., Maurer, M., Stumm, M., et al. (2004). Increased cancer risk of heterozygotes with NBS1 germline mutations in Poland. International Journal of Cancer, 111(1), 67–71.
Susswein, L. R., Marshall, M. L., Nusbaum, R., Vogel Postula, K. J., Weissman, S. M., Yackowski, L., et al. (2015). Pathogenic and likely pathogenic variant prevalence among the first 10,000 patients referred for next-generation cancer panel testing. Genetics in Medicine. doi:10.1038/gim.2015.166.
Tedaldi, G., Danesi, R., Zampiga, V., Tebaldi, M., Bedei, L., Zoli, W., et al. (2014). First evidence of a large CHEK2 duplication involved in cancer predisposition in an Italian family with hereditary breast cancer. BMC Cancer, 14, 478.
Turner, J., Kelly, B., Clarke, D., Yates, P., Aranda, S., Jolley, D., et al. (2011). A randomised trial of a psychosocial intervention for cancer patients integrated into routine care: the PROMPT study (promoting optimal outcomes in mood through tailored psychosocial therapies). BMC Cancer, 11, 48.
Walsh, T., Casadei, S., Coats, K. H., Swisher, E., Stray, S. M., Higgins, J., et al. (2006). Spectrum of mutations in BRCA1, BRCA2, CHEK2, and TP53 in families at high risk of breast cancer. JAMA, 295(12), 1379–1388.
Walsh, T., Lee, M. K., Casadei, S., Thornton, A. M., Stray, S. M., Pennil, C., et al. (2010). Detection of inherited mutations for breast and ovarian cancer using genomic capture and massively parallel sequencing. Proceedings of the National Academy of Sciences of the United States of America, 107(28), 12629–12633.
Walsh, T., Casadei, S., Lee, M. K., Pennil, C. C., Nord, A. S., Thornton, A. M., et al. (2011). Mutations in 12 genes for inherited ovarian, fallopian tube, and peritoneal carcinoma identified by massively parallel sequencing. Proceedings of the National Academy of Sciences of the United States of America, 108(44), 18032–18037.
Wooster, R., Bignell, G., Lancaster, J., Swift, S., Seal, S., Mangion, J., et al. (1995). Identification of the breast cancer susceptibility gene BRCA2. Nature, 378(6559), 789–792.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
A. E. Bunnell, C. A. Garby, E. J. Pearson, S. A. Walker, L. E. Panos and Joanne L. Blum state that they have no conflict of interest. Laura Panos, who was employed by Baylor Dallas, took a job at Ambry Genetic Laboratories, but it was well after the start of this study.
Human Studies and Informed Consent
All procedures followed were in accordance with the ethical standards of the responsible committee on human experimentation (institutional and national) and with the Helsinki Declaration of 1975, as revised in 2000. Informed consent was obtained from all patients for being included in the study.
Animal Studies
No animal studies were carried out by the authors of this article.
Rights and permissions
About this article
Cite this article
Bunnell, A.E., Garby, C.A., Pearson, E.J. et al. The Clinical Utility of Next Generation Sequencing Results in a Community-Based Hereditary Cancer Risk Program. J Genet Counsel 26, 105–112 (2017). https://doi.org/10.1007/s10897-016-9985-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10897-016-9985-2