Abstract
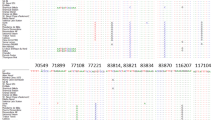
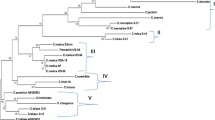
This study focused on clarifying phylogenetic relationships among Citrus accessions from Vietnam. Our phylogenetic analysis based on nucleotide sequences from the ITS of the ribosomal DNA included 69 accessions belonging to Citrus and related (sub)genera. Maximum parsimony and Bayesian analysis confirmed a clear separation of the three ‘true’ Citrus species (C. medica, C. maxima and C. reticulata). Confirming recent taxonomic revisions, Fortunella, Poncirus trifoliata and Citrus hystrix are clustered among the accessions of subgenus Citrus. C. × sinensis accessions revealed a close evolutionary relationship to either C. maxima or C. reticulata, thereby confirming their involvement in its hybrid origin. Also, some other hybrid taxa and their proposed parental species were investigated and their origin could in some cases be confirmed using the ITS sequence data.


Similar content being viewed by others
References
Barkley NA, Roose ML, Krueger RR, Federici CT (2006) Assessing genetic diversity and population structure in a Citrus germplasm collection utilizing simple sequence repeat markers (SSRs). Theor Appl Genet 112:1519–1531
Barrett HC, Rhodes AM (1976) A numerical taxonomic study of the affinity relationships in cultivated Citrus and its close relatives. Syst Bot 1:105–136
Bretó MP, Ruiz C, Pina JA, Asins MJ (2001) The diversification of Citrus clementina Hort. Ex Tan., a vegetatively propagated crop species. Mol Phyl Evol 21:285–293
Fang DQ, Roose ML (1997) Identification of closely related Citrus cultivars with inter-simple sequence repeats markers. Theor Appl Genet 95:408–417
Fang DQ, Roose ML, Krueger RR, Federici CT (1997) Fingerprinting trifoliate orange germplasm accessions with isozymes, RFLPs, and inter-simple sequence repeat markers. Theor Appl Genet 95:211–219
Federici CT, Fang DQ, Scora RW, Roose ML (1998) Phylogenetic relationships within the genus Citrus (Rutaceae) and related genera as revealed by RFLP and RAPD analysis. Theor Appl Genet 94:812–822
Gmitter FG Jr (1995) Origin, evolution and breeding of the grapefruit. Plant Breed Rev 13:345–363
Green RM, Vardi A, Galune E (1986) The plastome of Citrus. Physical map, variation among Citrus cultivars and species and comparison with related genera. Theor Appl Genet 72:170–177
Gulsen O, Roose ML (2001) Lemons: diversity and relationships with selected Citrus genotypes as measured with nuclear genome markers. J Amer Soc Hort Sci 126:309–317
Herrero R, Asins MJ, Carbonell EA, Navarro L (1996) Genetic diversity in the orange subfamily Aurantioideae. I. Intraspecies and intragenus genetic variability. Theor Appl Genet 92:599–609
Hodgson RW (1965) Taxonomy and nomenclature in the Citrus fruits. In: Krishnamurthi S (ed) Advances in agriculture sciences and their applications. Agric. Coll. Res. Inst., Coimbatore, pp 317–331
Hodgson RW (1967) Horticultural varieties of Citrus. In: Reuther W, Webber HJ, Batchelor LD (eds) The Citrus industry. Vol. I. history world distribution, botany, and varieties. Univ of California, Div of Agricultural Sci, Berkeley, pp 431–591
Hooker JD (1875) Rutaceae. In: Reeve L (ed) The flora of British India. London, pp 484–517
Jena SN, Kumar S, Nair NK (2009) Molecular phylogeny in Indian Citrus L. (Rutaceae) inferred through PCR-RFLP and trnL-trnF sequence data of chloroplast DNA. Scientia Hort 119:403–416. doi:10.1016/j.scienta.2008.08.030
Linnaeus C (1753) Species plantarum, exhibentes plantas rite cognitas, ad genera relatas, cum differentiis specificis, nominibus trivialibus, synonymis selectis, locis natalibus, secundum systema sexuale digestas. Holmiae
Mabberley DJ (2008a) 21. CITRUS Linnaeus, Sp. Pl. 2: 782. 1753. Flora China 11:90–96
Mabberley DJ (2008b) Mabberley’s plant book: a portable dictionary of plants, 3rd edn. Cambridge University Press, Avon
Moore GF (2001) Orange and lemons: clues to the taxonomy of Citrus from molecular markers. Trends Genet 17:536–540
Nicolosi E, Deng ZN, Gentile A, La Malfa S, Continella G, Tribulato E (2000) Citrus phylogeny and genetic origin of important species as investigated by molecular markers. Theor Appl Genet 100:1155–1166
Pang XM, Hu DG, Deng XX (2003) Phylogenetic relationships among Citrus and its relatives as revealed by SSR markers. Acta Genet Sin 30:81–87
Pang XM, Hu CG, Deng XX (2007) Phylogenetic relationships within Citrus and its related genera as inferred from AFLP markers. Genet Resour Crop Evol 54:429–436
Rogers SO, Bendich AJB (1988) Extraction of DNA from plant tissues Plant molecular Biology Manual, vol A6. Kluwer Academic Publishers, Dordrecht, pp 1–10 printed in Belgium
Ronquist F, Huelsenbeck JP (2003) MrBayes 3: Bayesian phylogenetic inference under mixed models. Bioinformatics 12:1572–1574
Scora RW (1975) On the history and origin of Citrus. Bull Torrey Bot Club 102:369–375
Singh R, Nath N (1969) Practical approach to the classification of Citrus. Proceedings of 1st international Citrus symposium, pp 435–440
Swingle WT, Reece PC (1967) The botany of Citrus and its wild relatives. In: Reuther W, Webber HJ, Batchelor LD (eds) The Citrus industry. Vol. I. History, world distribution, botany, and varieties. Univ of California, Div of Agricultural Sci, Berkeley, pp 190–430
Swofford DL (2002) PAUP. Phylogenetic analysis using parsimony (* and other methods). Version 4.0. Sinauer Associates, Boston
Tanaka T (1954) Species problem in Citrus. A critical study of wild and cultivated units of Citrus, based upon field studies in their native homes. Japanese Society for the Promotion of Science, Ueno
Tanaka T (1977) Fundamental discussion of Citrus classification. Stud Citrol 14:1–6
Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG (1997) The CLUSTAL_X Windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res 25:4876–4882
Torres AM, Soost RK, Diedenhofen U (1978) Leaf isozymes as genetic markers in Citrus. Am J Bot 65:869–881
Webber HJ (1967) History and development of the Citrus industry. In: Reuther W, Webber HJ, Batchelor LD (eds) The Citrus industry. Vol. I. history world distribution, botany, and varieties. Univ of California, Div of Agricultural Sci, Berkeley, pp 1–39
White TJ, Bruns T, Lee S, Taylor J (1990) Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In: Innis MA, Gelfand DH, Shinsky JJ, White TJ (eds) PCR protocols: a guide to methods and applications. Academic Press, San Diego, pp 315–322
Acknowledgments
Special thanks are given to colleagues and institutions for providing Citrus accessions and technical assistance: Dr. Do Dinh Ca-Research Institute for Fruits and Vegetables (RIFAV) (Ha Noi), Ms Vo Thi Tuyet-Phu Qui’s Fruit Tree Center, (PQFC), Dr. Pham Van Chuong-Agricultural Science Institute of North-Central Vietnam (ASINC), MSc Doan Nhan Ai- Fruit Tree Research & Development Center of ThuaThien-Hue (FRDC-Hue), Dr. Nguyen Minh Chau, Dr. Nguyen Thi Thu Hong, Ir Pham Ngoc Lieu-Institute of Southern Fruit Research Institute (SOFRI).
Author information
Authors and Affiliations
Corresponding author
Additional information
Tina Kyndt and Tran Nhan Dung equally contributed to this work.
Rights and permissions
About this article
Cite this article
Kyndt, T., Dung, T.N., Goetghebeur, P. et al. Analysis of ITS of the rDNA to infer phylogenetic relationships among Vietnamese Citrus accessions. Genet Resour Crop Evol 57, 183–192 (2010). https://doi.org/10.1007/s10722-009-9460-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10722-009-9460-0