Abstract
Marine aquaculture and stock enhancement are major causes of the introduction of alien species. A good example of such an introduction is the Japanese shortneck clam Ruditapes philippinarum, one of the most important fishery resources in the world. To meet the domestic shortage of R. philippinarum caused by depleted catches, clams were imported to Japan from China and the Korean peninsula. The imported clam is an alien species that has a very similar morphology, and was misidentified as R. philippinarum (hereafter, Ruditapes form). We genotyped 1,186 clams of R. philippinarum and R. form at four microsatellite loci, sequenced mitochondrial DNA (COI gene fragment) of 485 clams, 34 of which were R. variegatus, and measured morphometric and meristic characters of 754 clams from 12 populations in Japan and China, including the Ariake Sea and Tokyo Bay, where large numbers of R. form were released. Our analyses confirmed that R. form was from the genus Ruditapes, and the genetic differentiation between R. philippinarum and R. form was distinct, but small, compared with five bivalve outgroups. However, R. form had distinct shell morphology, especially larger numbers of radial ribs on the shell surface, suggesting that R. form might be a new Ruditapes species or a variation of R. philippinarum that originated from southern China. A genetic affinity of the sample from the Ariake Sea to R. form was found with the intermediate shell morphology and number of radial ribs, and the hybrid proportion was estimated at 51.3 ± 4.6 % in the Ariake Sea.
Similar content being viewed by others
Introduction
Introduction of non-native species has been a method of increasing fisheries production since the 1970s. As a result, marine aquaculture and stock enhancement have led to the introduction of marine species (Bax et al. 2003). An important concern in marine introductions is hybridization between introduced alien and closely related native species. Natural hybridization has played an important role in the evolution of many plant and animal taxa, but increased anthropogenic hybridization is causing extinction of many species, subspecies and locally adapted populations by both replacement and genetic mixing (Allendorf et al. 2001). Evidence of anthropogenic hybridization in marine fishery resources has recently been observed in bivalves. Yamakawa and Imai (2012) reported hybridization between the alien Meretrix petechialis (Lamarck 1818) introduced from China and the Korean peninsula and the native Meretrix rusolia (Röding 1798) in Japan. Likewise, Hurtado et al. (2011) found hybridization between distinct Ruditapes species: the introduced Japanese shortneck clam (or Manila clam) Ruditapes philippinarum (Adams and Reeve 1850) and the native European grooved carpet shell Ruditapes decussatus (Linnaeus 1758) in Spain.
In the genus Ruditapes, R. philippinarum is the most important commercial fisheries species, which occurs in East Asia, North America and Europe. Although it is native to Sakhalin, Japan, Korea and China (Gosling 2003), it was accidentally introduced into Puget Sound, Washington, USA, with the Pacific oyster Crassostrea gigas, in the 1930s (Williams 1980), and it has been cultured since the late 1970s (Becker et al. 2008). In the early 1970s, it was introduced from the USA for aquaculture purposes to France (Flye-Sainte-Marie et al. 2007) and the UK (Utting and Spencer 1992); it was introduced to Italy in the 1980s (Chiesa et al. 2011) and to Ireland in 1984 (Drummond et al. 2006), and from the UK to Israel (Shpigel and Spencer 1996). China is the number one country in the world in terms of production of the Manila clam, producing about 1.8 million metric tons annually, which accounts for about 90 % of global production (Zhang and Yan 2006).
In Japan, R. philippinarum is one of the most important fishery resources. Until the mid-1980s, the abundant annual catch from natural populations was stable at about 110–160 thousand tons, but began to decrease from the mid-1980s. This severe decrease occurred in two major production areas: Tokyo Bay off Chiba prefecture, and the Ariake Sea off Kumamoto prefecture (Fig. 1). In contrast, in the top production area of Mikawa Bay, Aichi prefecture, the catch has remained stable since the 1970s, but has increased since the late 2000s (Fig. S1a in the Supplementary material). In the major production areas, naturally produced seed of R. philippinarum were translocated and released within and between prefectures to enhance fishery production. In 2009, 14,027 million naturally produced seed with a mean shell length (SL) of 12 mm (range 4–56), and 5,775 thousand artificially produced spat of 17 mm SL (1–30) were released for fishery stock enhancement in Japanese waters (Fig. S1b, Fisheries Research Agency and National Association for the Promotion of Productive Seas 2011). Despite stock enhancement efforts, the annual catch remained low at 31,500 tons in 2009 and close to the historical minimum of 31,022 tons in 2001 (Ministry of Agriculture, Forestry and Fisheries 1967–2011, Fig. S1a). To meet the shortage in domestic demand, R. philippinarum was imported into Japan from China and the Korean peninsula in the late 1980s. Imports increased to about 75,000 tons in 2000, but decreased to 12,000 tons in 2008 (Ministry of Finance 1991–2010, Fig. S1a). Because imported clams were shipped to consumer markets directly, this is not the problem. The issue of conservation arose from the fact that live clams were released into shallow waters for short-term aquaculture before they were sent to consumer markets. In addition, huge numbers of clams were released into natural waters, mainly in the Ariake Sea and Tokyo Bay, to enhance stocks for commercial and recreational fisheries. However, the release statistics do not include the number of clams released in Japan for recreational clam gathering, and the release information for imported clams is unknown.
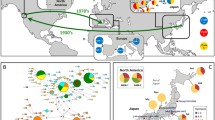
Sampling sites for Ruditapes philippinarum (green for samples from Japan and blue for samples from China), R. form (red) and R. variegatus (black).The number of the site shows the locality in Table 1. 1–3 Tokyo Bay (TKB), 4 Mikawa Bay (MKB), 5 Ariake Sea (ARS), 6 Ryugatake (RGT), 7 Ushibuka (USB), 8 Lake Notoro (LNT), 9 Tateyama (TYM), 10 Dalian (DAL), 11 Rushan (RUS), 12 Dandong (DAD), 13 Haikou (HAK). (Color figure online)
Vargas et al. (2008) examined the shell morphology and allozyme variations of R. philippinarum imported from China and the Korean peninsula at the Chiba Prefectural Fisheries Cooperative Association. They found that the imported clam was very similar to R. philippinarum, but significantly genetically different from R. philippinarum samples collected in Japan and China, which authors concluded had been misidentified as R. philippinarum. Vargas et al. (2010) confirmed that the clams collected at Dandong, Xiamen and Haikou, China were genetically the same as the above-mentioned imported clam and exhibit finer, more closely spaced radial ribs on shell surfaces compared with native R. philippinarum. The shell of the clam is thinner (data not shown), and its shell characteristics are different from R. philippinarum, as shown in the Results section. Based on shell morphology, we confirmed that large numbers of these clams were released in the recreational shell-gathering grounds in Tokyo Bay, where we took samples at TKB-2 and TKB-3 (see Table 1). The clam does not naturally inhabit Japanese waters and, in fact, we did not find this clam among our samples of native R. philippinarum. The clam is therefore an alien species. Judging from the shell morphology, the clam should be genus Ruditapes. However, the taxonomic status of the clam was not clear. Therefore, in this paper, we tentatively treat the clam as a Ruditapes form.
Vargas et al. (2010) estimated Nei’s genetic distance (Nei 1978) from allozymic genotype frequencies between samples of R. philippinarum and R. form at 0.1069 ± 0.0258; the within-sample estimates were much smaller, being 0.0049 ± 0.0014 for R. philippinarum and 0.0105 ± 0.0044 for R. form. Using mitochondrial DNA (mtDNA) cytochrome oxidase subunit I (COI) sequences, Sekine et al. (2006) reported genetic differentiation between R. philippinarum samples collected in Japan (28 individuals from six sites) and in China (11 individuals from three sites). Mao et al. (2011) analysed the sequences of Sekine et al. (2006) combined with their own samples of R. philippinarum collected in Japan (19 individuals from two sites) and China (132 individuals from eight sites), and confirmed the distinct differentiation between R. philippinarum populations in Japan and China. However, some of their Chinese samples are not R. philippinarum but rather R. form, judging from the substantial pairwise F ST values between the Chinese and Japanese samples of 0.6417 ± 0.0470 (0.531–0.817). In this average F ST calculation, we excluded the Chinese samples from Tj (Tianjin), Qd (Qingdao) and Rs (Rushan) which might contain R. philippinarum shown as the haplotype lineage A (see Fig. 3 of Mao et al. 2011), and the Japanese sample Kag (Kagawa) which might contain R. form clams imported and released for recreational shell gathering. In fact, the samples Tj and Qd from the Bohai Sea had small F ST values of 0.0564 ± 0.0644 (−0.063–0.152) between Japanese samples, but had very large values of 0.4198 ± 0.1419 (0.199–0.576) between the Chinese samples from Dl1 (Dalian 1), Dl2 (Dalian 2), Xia (Kiaochow Bay), Lz (Laizhou), Nb (Ningbo), Pt (Putian), Xm (Xiamen), and Gz (Guangzhou). The pairwise F ST values of Mao et al. (2011) strongly suggested that the haplotype lineage A (red colour in their paper) was R. philippinarum, and the lineages B and C (green and blue colours) were R. form. Available evidence shows that Ruditapes species can hybridize if geographic isolation is removed because R. philippinarum and R. decussatus hybridize, as mentioned above. The genetic difference between R. philippinarum and R. form is much smaller than that between R. philippinarum and R. decussatus as shown in the “Results” section. Therefore, R. form can hybridize with the native Japanese R. philippinarum by anthropogenic introduction. If this is the case, backcrossing with parental types and mating among hybrids by varying numbers of generations should make the introduced population a hybrid swarm; that is, a population of individuals that are all hybrids (Allendorf et al. 2001). In this case, the genotypes of R. form should be half in the hybrid swarm and the pure R. philippinarum would become extinct in the introduced population. Therefore, the introgression—gene flow between populations whose individuals hybridize (Allendorf et al. 2001)—of R. form into R. philippinarum is a real issue of concern from a conservation viewpoint.
In this study, we first explore the phylogenetic relationships of R. form with R. philippinarum compared with five outgroups: R. bruguieri (Hanley 1845), R. decussatus, R. variegatus (synonym R. variegate, Sowerby 1852), Meretrix lusoria and M. petechialis. We then investigate genetic diversity and population structure of R. philippinarum and R. form in Japan and China, based on samples taken from natural populations using fine-scale DNA markers. Finally, we examine if the alien R. form is hybridizing with native R. philippinarum, based on genetic and morphometric measurements of clams from the Ariake Sea and Tokyo Bay, Japan, where large numbers of R. form have been released. In addition, the allozyme genotype data from Vargas et al. (2008, 2010) were reanalysed for the individuals used in this study. This study provides the first evidence for anthropogenic hybridization of R. philippinarum and R. form, based on their population structure in Japan and China. Our study also highlights a need for a taxonomic revision of species in the genus Ruditapes.
Materials and methods
Collecting samples
More than 1,200 individuals were collected from 12 populations, consisting of 10 R. philippinarum and two R. form. In addition, 50 individuals of R. variegatus were collected as an outgroup (Fig. 1; Table 1). Sampling sites in Japan for R. philippinarum were Tokyo Bay off Chiba (TKB-1 and TKB-2) and off Kanagawa prefectures (TKB-3), Mikawa Bay in Aichi prefecture (MKB), the Ariake Sea (ARS), Ryugatake (RGT) and Ushibuka (USB) in Kumamoto prefecture, and Lake Notoro (LNT) in Hokkaido. Sites for R. philippinarum in China were Dalian (DAL) in Liaoning, and Rushan (RUS) in Shandong province. Samples were collected manually from tidal flats or shallow waters, except for the ARS sample, which was taken by fishermen. Large numbers of R. form were released for fisheries stock enhancement in the Ariake Sea and Tokyo Bay, but current releases of R. form are mainly for recreational fishing. Indeed, a large number of R. form was released at the sampling sites of TKB-1 and TKB-2, which were recreational fishing areas. To take a sample of naturally born R. philippinarum in the recreational fishing areas, we collected 0+-year-old young clams, judging from their SL. From the SL, those from TKB-1 were thought to be born in autumn and TKB-2 in summer of the previous year. In contrast, there has been no release of R. form in TKB-3. The sample from ARS was taken from a fishing ground in the Ariake Sea where R. form had not been released for more than 10 years. In Mikawa Bay, naturally born spat of R. philippinarum collected in the bay were released within the bay, but there was no introduction of R. form. Samples from RGT, USB, LNT, DAL and RUS were wild clams, because stocking or introduction of R. philippinarum and R. form did not occur in these areas. The R. form sample from Haikou (HAK) was taken from a wild population from Hainan Island where there had never been translocation or stocking. The sample from Dandong (DAD) was taken from a naturally reproducing population of R. form in the coastal waters of Pyonganbuk-do, North Korea near Dandong, introduced from Fujian province, China to North Korea (personal communication, Sun Zonghmin). The R. variegatus sample from Tateyama (TYM), Chiba prefecture, Japan, was collected manually from a wild population in a shallow sandy bottom with rocks near tide pools. Thus, all samples were naturally born individuals taken from the introduced populations and/or wild individuals from pure populations without any introduction (Table 1). All specimens were stored at −80 °C before genetic analyses.
Mitochondrial DNA sequencing
In some shellfish species such as R. philippinarum, “Doubly Uniparental Inheritance” (DUI) of mitochondrial DNA (mtDNA) was reported with two different sex-related M- and F-type mtDNAs (Passamonti and Scali 2001; Passamonti et al. 2003). Plazzi and Passamonti (2010) recommended that DNA should be obtained from the foot muscle as DUI species carry very little M-type mtDNA in the foot muscle. Therefore, we only used adductor and foot muscle tissue to avoid co-extraction of male-type mtDNA. The muscle tissue from each individual was boiled in water at 96 °C for 5 min and stored in 99.5 % ethanol. Genomic DNA was extracted following the standard phenol–chloroform procedure and/or with the DNeasy Blood and Tissue Kit (Qiagen), according to the manufacturer’s instructions.
DNA fragments corresponding to COI (about 700 bp) were amplified with polymerase chain reaction (PCR) using the universal primers for invertebrates LCO1490: 5′ GGTCAACAAATCATAAAGATATTGG 3′ and HCO2198: 5′ TAAACTTCAGGGTGACCAAAAAATCA 3 ′ (Folmer et al. 1994). PCR amplifications were carried out using a GeneAmp PCR System 2700 Thermal Cycler (Applied Biosystems; www.appliedbiosystems.com) in 25 μl aliquots of a mixture containing 1.0 μl of genomic DNA as a template (~1.0 μg/μl), 0.13 μl of Takara Ex Taq polymerase (Takara Bio Inc.; www.takara-bio.com), 1.0 μl of 10 μM primers, 2.5 μl of 10× Ex PCR buffer (MgCl2 20 mM), 2.0 μl of dNTP mixture (2.5 mM of each dNTP) and 17.4 μl of sterile water with the following thermal-cycle profile: 1 min initial denaturation at 95 °C followed by 35 cycles of 1 min denaturation at 95 °C, 1 min annealing at 40 °C and 7 min extension at 72 °C. The PCR product was purified with ExoSAP-IT (USB Corp., USA). The purified product was used as the template DNA for cycle sequencing reactions performed on a GeneAmp PCR System 2700 Thermal Cycler (Applied Biosystems) in 20 μl aliquots of a mixture containing 2 μl of template DNA, 0.32 μl of 10 μM primer HCO2198, 1.5 μl of BigDye Terminator v3.1 or v1.1 Cycle Sequencing Ready Reaction Kit (Applied Biosystems) and 16.2 μl of sterile water with the following cycling profile: 1 min initial denaturation at 96 °C, followed by 25 cycles of 10 s denaturation at 96 °C, 5 s annealing at 50 °C and 4 min extension at 60 °C. Sequencing products were subjected to capillary electrophoresis in an ABI PRISM 3130xl Genetic Analyzer (Applied Biosystems). A total of 519 individuals were sequenced (Table 1).
Microsatellite DNA genotyping and allozyme data
Nine microsatellite markers developed for R. philippinarum by Yasuda et al. (2007) were examined, and 1,186 individuals were genotyped at four loci (Asari 16, Asari 55, Asari 62, Asari 64). All loci were scored by capillary electrophoresis using an ABI PRISM 3130xl Genetic Analyzer (Applied Biosystems) with fluorescent dye-labelled primers. The thermal cycles for PCR amplification were conducted in 13 μl aliquots of a mixture containing 1.0 μl of genomic DNA as a template, 0.05 μl of Takara Ex Taq polymerase (Takara Bio Inc.; www.takara-bio.com), 1.0 μl of 10 μM primers, 1.25 μl of 10× Ex PCR buffer (MgCl2 20 mM), 1.0 μl of dNTP mixture and 7.7 μl of sterile water with the following cycling profile, consisting of 38 cycles of denaturation at 94 °C for 10 min, a primer-specific annealing temperature for 30 s, and an extension at 51 °C for 30 s and 72 °C for 1 min. The annealing temperatures were 51 °C for all primers.
Allozyme genotypes of 627 individuals in the samples analysed for mitochondrial and microsatellite analysis that had no missing data were used for the data analysis from 722 individuals in Vargas et al. (2008, 2010). The genotypes were encoded by seven loci (IDH*, MDH-1*, MDH-2*, PGM*, 6PGD*, SOD-1* and SOD-2*), excluding the LAP* locus, which departed from Hardy–Weinberg equilibrium (HWE) in all samples. The techniques employed for electrophoretic analyses and staining protocols were described in Vargas et al. (2008).
Phylogenetic analysis
Sequences were aligned using Clustal X (Thompson et al. 1997). Alignments were also manually adjusted and no indels (insertions/deletions) were found in the sequences. Haplotypes were defined based on sequence data using DnaSP ver. 5.00 (Librado and Rozas 2009).
To explore the taxonomic status of R. form, genetic distances among the major haplotypes of R. philippinarum collected in Japan and in China (this study, Hap 2, 6, 15, 24 and 32), those of R. form (Hap 92 and 115), and all haplotypes of R. variegatus collected in Japan (Hap 1–7) were estimated with sequences of R. philippinarum F-type and M-type collected in Japan, those of R. philippinarum collected in southern China [(a) Mao et al. (2011) and (b) Chen et al. (2011)], R. variegatus collected in southern China (Chen et al. (2011), R. bruguieri, R. decussatus, M. lusoria and M. petechialis, with their GeneBank accession number. Substitution models were compared (Guindon and Gascuel 2003; Posada and Buckley 2004; Posada 2008) with the Bayesian information criterion (BIC) (Schwarz 1978), and the corrected Akaike information criterion (AICc) (Akaike 1974; Sugiura 1978; Hurvich and Tsai 1989). The maximum likelihood (ML) (Felsenstein 1981) tree of the COI haplotypes under the best-fit substitution model was constructed. The analysis was implemented using MEGA5 (Tamura et al. 2011). The same procedure was applied on our haplotypes of R. philippinarum and R. form. Genealogical relationships among haplotypes were also examined by constructing haplotype networks using a median-joining method (Bandelt et al. 1999) in Network ver. 4.6.0.0. Genetic diversity in each sample was quantified as the number of haplotypes, haplotype diversity (Nei 1987), and nucleotide diversity (Tajima 1983; Nei 1987) using Arlequin ver. 3.5.1.2 (Excoffier et al. 2005).
Population genetics analysis
The HWE was tested, and heterozygosity was calculated for each locus and population using GENEPOP version 4.0.10 (Raymond and Rousset 1995a). The exact test of population differentiation based on haplotype and allele frequencies was performed using the Markov chain procedure (1,000 dememorizations, 10,000 steps in the Markov chain) (Raymond and Rousset 1995b), as implemented in Arlequin. The initial α = 0.05 significance level was adjusted for simultaneous pairwise comparisons using the sequential Bonferroni correction (Rice 1989). The fixation index F ST is most commonly used to measure genetic divergence among subpopulations (Palsbøll et al. 2006). The global F ST over populations and posterior means of population pairwise F ST were therefore estimated based on an empirical Bayesian method (Kitada et al. 2007) using the software POPDIF1.0 (http://popgene.co-site.jp/software.html) to avoid a biased estimation of pairwise F ST, especially for species with large gene flow, such as marine fish and shellfish. Nei’s genetic distance (D, Nei 1978) and R ST (Slatkin 1995) was also calculated for all pairs of populations using GenAlEx 6.5 (Peakall and Smouse 2006) and SPAGeDi 1.3 (Hardy and Vekemans 2002), respectively. R language (R Development Core Team) was used for the cluster analysis based on pairwise D, F ST and R ST. To describe the genetic characteristics of the populations, STRUCTURE 2.3.3 (Pritchard et al. 2000) was implemented for 500,000 Markov chain Monte Carlo replicates after a burn-in period of 50,000 under the correlated allele frequency model. The number of putative ancestry populations was assumed to be two (K = 2) because we analysed two putative populations, R. philippinarum and R. form.
To infer introgression of R. form into R. philippinarum populations, the mixing proportion (here, it refers to the hybridization proportion) of R. form in the Ariake Sea and Tokyo Bay was estimated from sample multilocus allele frequencies for the mixed (= hybridized) and baseline populations by the extended method of Kishino et al. (1994) (see “Appendix” section) using POPMIX2.0 (http://popgene.co-site.jp/software.html). Although there were no baseline data before the import of R. form, in this study, the samples from USB and MKB were assumed as baseline populations for wild R. philippinarum from the Ariake Sea and Tokyo Bay, respectively before the release of R. form. ARS and TKB were treated as samples from the hybridized populations affected by releases of R. form. Our reasoning for this is that R. form has never been released in Ushibuka, so USB might not be affected by releases of R. form because of its distance from the Ariake Sea (Fig. 1). There was a long history of translocation of seed from Mikawa Bay to Tokyo Bay before the imports of R. form, and no difference in allele frequencies between MKB and TKB was found, as stated later. Therefore, the allele frequencies in USB and MKB could be regarded as similar to those in ARS and TKB at present, without releases of R. form.
Morphometric analysis
The number of radial ribs (RR) on the shell surface was counted in all samples using a magnifying glass. The SL, shell height (SH) and shell width (SW) were also measured for all samples (Table 1). The shell morphological data of RR (SH/SL, SW/SL) were analysed using discriminant canonical variate analysis (CVA). The linkage between individual numbers of RR and genotypes was examined by a principal component analysis (PCA) based on allozyme and microsatellite genotypes for 42 individuals from the ARS sample that had genotype data with the number of RR. R language was used for the multivariate analyses.
Results
Mitochondrial DNA phylogeny
The 493 bp COI sequences were amplified for R. philippinarum and R. form, and the sequences were aligned. Sequence comparisons yielded 121 haplotypes from 485 individuals (Table S1). The COI sequences of all R. philippinarum individuals amplified with the universal primers for invertebrates coincided with the F-type COI sequence (AB065375), but the M-type COI sequence of R. philippinarum (AB0653745) was very different (figure not shown). We excluded the M-type COI sequence from further analysis. The results confirmed that the COI sequences amplified with the universal primers for invertebrates were the F-type COI sequence of R. philippinarum and R. form (Fig. 2).The number of private haplotypes was 60 for R. philippinarum collected in Japan (hereafter Japanese R. philippinarum), 19 for R. philippinarum collected in China (Chinese R. philippinarum), and 31 for R. form, respectively. Hap 81, Hap 21 and Hap 72 were common for Japanese/Chinese R. philippinarum and R. form. Japanese and Chinese R. philippinarum shared four haplotypes (Hap 12, Hap 32, Hap 60, Hap 67), and Chinese R. philippinarum and R. form also shared four (Hap 22, Hap 97, Hap 106, Hap 110). For R. variegatus, 493 bp COI sequences were also amplified and aligned. Sequence comparisons yielded seven haplotypes from 50 individuals. The sequences (493 bp) were submitted to the DDBJ, the European Molecular Biology Laboratory (EMBL) and Genbank databases under accession numbers AB694757–AB694847 for R. philippinarum, AB694848–AB694884 for R. form, and AB694885–AB694891 for R. variegatus.
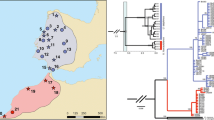
The ML tree by the best-fitted model (HYK + G) for mtDNA COI sequences (476 bp) of Ruditapes philippinarum (from Japan and China) with the F-type haplotype of R. philippinarum, R. form, R. variegatus, R. bruguieri, R. decussates, Meretrix lusoria and M. petechialis. The sequences with the accession numbers were taken from the DDBJ and those of Hap # were obtained in this study. The abbreviations in the figure refer to sampling sites in China: a Mao et al. (2011) and b Chen et al. (2011). The cluster consisting of R. form with R. philippinarum collected in southern China is marked as a box. Support values for the topology obtained from 1,000 boot strapping are listed as percentages
We aligned our sequences with those of the outgroups, and 476 bp sequences were obtained. The best substitution model selected by BIC (4829.32) and AICc (4327.24) was Hasegawa–Kishino–Yano (HKY) (Hasegawa et al. 1985) with discrete gamma rate heterogeneity among sites (+G) (Yang 1993). The best ML tree (HKY + G) showed large genetic differentiation of R. philippinarum from R. variegatus, R. decussatus, M. lusoria and M. petechialis. Our haplotypes of Japanese and Chinese R. philippinarum made a monophyletic cluster. However, R. form belonged to a different cluster of R. philippinarum collected in southern China [(a) Mao et al. (2011) and (b) Chen et al. (2011)] (marked as a box in Fig. 2). The evolutionary distance between our R. philippinarum and R. form samples was much smaller than that between R. variegatus samples collected in southern China (Chen et al. 2011), and our sample collected in Japan. The sequence of R. bruguieri (Mikkelsen et al. 2006) was homologous with those of our R. variegatus sample.
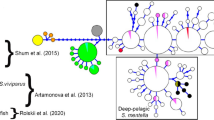
When analysing our 121 haplotype sequences (493 bp) of R. philippinarum and R. form, the best substitution model selected by BIC (5922.16) was HKY with a certain fraction of invariable sites (+I) (Fitch and Margoliash 1967; Shoemaker and Fitch 1989). The AICc (3722.30) selected a different model: Tamura–Nei (TN93) (Tamura and Nei 1993) (+I). The best BIC ML tree (HKY + I) detected clear genetic differentiation of R. form from R. philippinarum, and Chinese R. philippinarum was included in both clusters: the cluster mainly consisting of R. philippinarum and the cluster mainly consisting of R. form and some common haplotypes (Fig. 3). A network diagram of the 121 haplotypes also identified two distinct clusters of R. philippinarum and R. form (Fig. S2). Three individuals from TKB-3 and MKB had a major haplotype of R. form and Chinese R. philippinarum (Hap 81). Two individuals from ARS had minor haplotypes of R. form from Dandong (Hap 21 and Hap 72) (Table S1). However, we found no diagnostic haplotype composition between R. form and Japanese R. philippinarum samples.
The ML tree by the best-fitted model (HYK + I) for 121 mtDNA COI haplotypes (493 bp) of Ruditapes philippinarum [sampled in Japan (green circle) and in China (blue circle)] and R. form (red circle).Common haplotypes among R. philippinarum sampled in Japan and in China and R. form (blue triangle), those for among R. philippinarum sampled in Japan and in China (green triangle) and those for among R. philippinarum sampled in China and R. form (red triangle). Only topology was shown for simplicity. (Color figure online)
Genetic diversity, population structure and introgression
Numbers of haplotypes, haplotype diversity and nucleotide diversity were relatively high in all samples except for RGT and USB, which were taken from small pure populations occurring in small tidal flats (Table S1). The global F ST estimate ± SE was 0.05330 ± 0.00704. The UPGMA tree based on the posterior means of pairwise F ST described the clusters of RGT and USB, and R. form samples of DAD and HAK. ARS was in an intermediate position between R. form samples and other Japanese and Chinese R. philippinarum samples (Fig. 4a, Table S2). The network diagram suggested no mtDNA introgression of R. form into R. philippinarum populations of the Ariake Sea and Tokyo Bay (Fig. S2).
Genetic differentiation of R. philippinarum and R. form populations on the basis of an empirical Bayes (EB) \( F_{ST} \) estimate inferred from a mtDNA COI haplotype frequencies, b EB \( F_{ST} \), c Nei’s genetic distance, d \( R_{ST} \) estimates from microsatellite DNA allele frequencies and e EB \( F_{ST} \) estimates from allozymic allele frequencies
The locus Asari 55 departed significantly from HWE in all samples; therefore, we excluded the genotype data from further analysis. For the three microsatellite loci, 63 alleles were found among 807 Japanese R. philippinarum, 199 Chinese R. philippinarum, and 180 R. form. All samples were in HWE at the loci Asari 62 and Asari 64 (except LNT), although four samples departed from HWE at Asari 16, supporting the proposal that all samples were collected from naturally produced and/or wild populations. The mean number of alleles and observed heterozygosity per locus was 11.3–18.3 and 0.570–0.790. When compared with the R. philippinarum samples, there was a private allele in samples of R. form (143 at Asari 64), but it was a very minor allele. Diagnostic allele composition between R. form and Japanese R. philippinarum samples was observed in 158 alleles at the Asari 16 locus. The 158 allele was the major allele in R. form samples, and the second most frequent allele in Chinese R. philippinarum samples; it was the major allele in ARS and frequent in the adjacent RGT and USB (Fig. 5). In contrast, the 158 allele was minor in other Japanese R. philippinarum. Detailed allele frequency distributions are summarized in Table S3. For the seven allozymic loci, 58 alleles were found among 453 Japanese R. philippinarum and 174 R. form. The genetic diversity in the allozymic loci was much lower than that in the microsatellite loci. The number of alleles and the observed heterozygosity per locus were 1–14 and 0.010–0.653 (Table S4). There was no private diagnostic allele between R. form and Japanese R. philippinarum samples.
The null hypothesis of homogeneity of allele frequencies of microsatellite loci between pairs of samples was rejected, except for that between TKB (1, 2 and 3) and MKB, and between MKB and USB after sequential Bonferroni correction (P < 0.05). The global F ST estimate ± SE was 0.0240 ± 0.0026, which was about half of that from the COI (0.0533 ± 0.0070), and the allozyme data (0.0469 ± 0.0108). The pairwise F ST within Japanese R. philippinarum samples, excluding ARS, was very small (0.0023–0.0073). On the other hand, ARS had much larger values of F ST between Japanese R. philippinarum (0.0168–0.0275) and Chinese R. philippinarum samples (0.0227–0.0239), but provided smaller values between R. form samples (0.0140–0.0142) (Table S5). The pairwise F ST between R. form and R. philippinarum samples, excluding ARS, was the largest at 0.0424–0.0642 (Table S5). Nei’s genetic distances were much higher than F ST, which ranged 0.015–0.095 within Japanese R. philippinarum samples, excluding ARS (Table S6). Those for ARS between Japanese R. philippinarum (0.224–0.279) and Chinese R. philippinarum samples (0.201–0.230) were similar and very large, but much smaller between R. form (0.062–0.095). Those between R. form and R. philippinarum samples, excluding ARS, was the highest at 0.317–0.567, but that within R. form was small (0.046). The UPGMA dendrogram of the pairwise F ST and Nei’s genetic distance clearly described three clusters of Japanese R. philippinarum, Chinese R. philippinarum and R. form, and revealed genetic affinity of ARS with R. form, respectively (Fig. 4b, c). The R ST dendrogram also confirmed the three clusters (Fig. 4d, Table S5). Interestingly, RGT and USB were in the same cluster with Chinese R. philippinarum from DAL and RUS, and other Japanese samples were included in another cluster. The cluster analysis of the pairwise F ST based on allozymic allele frequencies supported the significant genetic differentiation of R. form (0.0577–0.1062), excluding ARS, and the difference of ARS (0.0049–0.0154) from the Japanese R. philippinarum samples (Fig. 4e, Table S7).
The bar plots by STRUCTURE described different genetic characteristics between R. philippinarum and R. form. ARS had a genetic affinity with R. form and was quite different from the R. philippinarum samples (Fig. 6). Assuming the baseline populations of R. form as DAD + HAK (n = 180), and native R. philippinarum as USB (n = 94), the hybridization proportion ± SE of R. form in the R. philippinarum population in the Ariake Sea (ARS, n = 99) was estimated at 51.3 ± 4.6 %, based on the multilocus microsatellite allele frequencies. In contrast, the estimate in Tokyo Bay was very small at 5.8 ± 2.0 % for 2009 (TKB-1, n = 140) and 3.7 ± 1.8 % for 2010 (TKB-2, n = 97), assuming the same baseline population of R. form (n = 180) and that of R. philippinarum (MKB, n = 88). The precision was very high in the Ariake Sea (CV = 0.09, coefficient of variation), but very low in Tokyo Bay (CV = 0.35 and 0.49). Our nuclear DNA analysis revealed substantial introgression of R. form into R. philippinarum populations in the Ariake Sea, but did not detect introgression into Tokyo Bay taking account of the precisions.
The bar plots by STRUCTURE (Pritchard et al. 2000) for a microsatellite genotypes (1,186 individuals) and b allozymic (627 individuals) given the putative number of ancestral populations (K = 2). Green colour refers to the putative ancestral population of Ruditapes philippinarum and red colour to that of R. form. (Color figure online)
Morphological diversity
The mean SL was 32.5 ± 4.6 for Japanese R. philippinarum excluding MKB sample, 33.1 ± 4.5 for Chinese R. philippinarum, and 33.2 ± 3.9 for R. form, respectively. There were no differences between the adult samples (P > 0.1, Welch two sample t test with Bonferroni correction). The MKB sample was small (SL, 20.1 ± 1.23 mm), and might not have the proportions of an adult shell. R. variegatus (adult shell) had a smaller SL (27.1 ± 1.06 mm) than R. philippinarum and R. form (P < 0.01). SW/SL and SH/SL were smaller in R. form, ARS, MKB and R. variegatus samples than in R. philippinarum samples (P < 0.01, Fig. S3). The number of RR was largest in R. form samples (103.3 ± 11.5, DAD + HAK, P < 0.01). The Chinese R. philippinarum samples (70.0 ± 7.2) had slightly larger numbers of RR than those of Japanese samples (66.0 ± 9.3) when we excluded the ARS sample (P < 0.01). The ARS sample had a larger number of RR (80.3 ± 10.1) than other Japanese and Chinese R. philippinarum samples (P < 0.01). R. variegatus had a smaller number of RR (76.5 ± 8.6) than that of the R. form, but larger than R. philippinarum (P < 0.01). There was no correlation between the individual SL and the number of RR (Fig. S4); therefore, the meristic character could be used as an individual specific trait.
The sample of MKB was excluded from CVA for R. philippinarum, along with the TKB-1 (SL 21.1 ± 1.70 mm) and TKB-2 (27.0 ± 1.61 mm) samples because they were not adult. CVA revealed the morphological difference between R. philippinarum and R. form samples, and that individuals of ARS samples had intermediate canonical scores (Fig. 7). In fact, individuals of ARS samples had an intermediate proportion of the shell (Fig. 8) with intermediate numbers of RR (Fig. 9). In contrast, samples from a recreational clam gathering ground in Tokyo Bay (TKB-1 and TKB-2) had similar numbers of RR (56.2 ± 7.4 and 59.5 ± 7.6) to other Japanese R. philippinarum samples (Fig. 8). The frequency of 158 allele at the Asari16 locus, which was a diagnostic allele for introgression of R. form into Japanese R. philippinarum populations, was very high at 53.0 % in the Ariake Sea. However, the PCA scores showed that the microsatellite and allozymic marker genotypes were not linked with the number of RR (Fig. S5).
Discussion
We found considerable introgression of the alien R. form into the R. philippinarum population in the Ariake Sea based on microsatellite DNA and morphological analyses. This could be caused by the mass releases of R. form into commercial fishing grounds in the late 1980s when the catch was substantially decreasing. Considering the high hybridization proportion of R. form, the genetic affinity with R. form and the intermediate shell morphology in the Ariake Sea, F1 hybrid individuals were fertile, backcrossed with wild R. philippinarum and admixed. The generation time of R. philippinarum is 1 year, so hybrid individuals could have been produced for more than 20 generations. As our ARS sample was taken from a fishing ground free from introduction of R. form, our estimate of the hybridization proportion suggested that hybrid individuals spread over a wide area. The hybrid process might be in the equilibrium, judging from the intermediate distribution in the number of RR in the Ariake Sea, and the mixing proportion of 51.3 ± 4.6 %. The results strongly suggest that the present R. philippinarum population in the Ariake Sea is a hybrid swarm, in which essentially all individuals are of hybrid origin (Allendorf et al. 2001). The frequencies of the diagnostic 158 allele at the Asari16 locus strongly supported introgression of males, because our mitochondrial DNA analysis did not support introgression into the Ariake Sea. The result suggested that only males of R. form introduced into the fishing ground succeeded in mating with native females R. philippinarum in the Ariake Sea. Interestingly, the evidence of introgression of male R. philippinarum into native R. decussates populations (Hurtado et al. 2011) coincided with our data. R. form was imported in large hemp bags without seawater. As such, females might be more sensitive to such environmental stress than males in reproduction of Ruditapes.
In contrast, no introgression of R. form was suggested in Tokyo Bay, where R. form was released into a large tidal flat for recreational clam gathering. The difference between the Ariake Sea and Tokyo Bay was the release sites: commercial and recreational fishing grounds. As the recreational fishing intensity was very strong, especially on weekends in Tokyo Bay, successive releases were made every week. Almost all R. form released just before weekends should be caught on the weekends during the recreational clam gathering season from April to August. In addition, the recreational fishing ground spread over a large tidal flat and almost all of the tidal flat is exposed at low tide. The water temperature was therefore very high in the tidal flat. We observed many dead individuals and shells of R. form in the tidal flat. R. form clams imported without seawater and released on the tidal flats might not have a chance to reproduce in such a stressful environment in the short time before they were caught.
The genetic differentiation between R. philippinarum and R. form was distinct both in mitochondrial and nuclear DNA, with pairwise F ST values of 0.0314–0.0876 (mtDNA COI), 0.0424–0.0642 (microsatellite) and 0.0577–0.1062 (allozyme). Nei’s genetic distance between R. philippinarum and R. form (0.317–0.567) was much larger than F ST values. Our mtDNA best ML tree and the network diagram were in agreement with the haplotype network of Mao et al. (2011). Their F ST values were 0.531–0.725 (mtDNA COI) between the Japanese (excluding the Kagawa sample) and Chinese R. philippinarum samples, which were much larger than our estimates. This clearly showed that their samples did not consist of a single species but R. philippinarum and R. form, although their large F ST estimates could be biased due to the small sample sizes (2–20 individuals per sample site). Their Kagawa sample (n = 7) had haplotypes of both R. philippinarum and R. form as shown in their Fig. 3, but the latter might be individuals that were imported and released R. form for recreational clam gathering.
In our mtDNA analysis, RGT and USB formed a more distant cluster from R. form and Japanese R. philippinarum. This might be because of the lower haplotype diversities due to the small population sizes left in small tidal flats. In fact, RGT and USB were included in the same clusters of Japanese R. philippinarum populations for microsatellite and allozyme alleles, but these two samples belonged to the same cluster with Chinese R. philippinarum of DAL and RUS by the R ST values, while DAL and RUS were distinct from Japanese R. philippinarum evaluated by microsatellite allele frequencies. The allozyme dendrogram showed the largest difference between R. philippinarum and R. form, and ARS was included in the same cluster of RGT and USB. These results suggest that Japanese R. philippinarum in the Ariake (ARS) and Shiranui Sea (RGT and USB) might be a relictual species (Kanemori et al. 2006) isolated from continental (Chinese) R. philippinarum populations.
The taxonomy of R. form is somewhat confused in the literature. Three valid species were recognized in the genus Ruditapes from the Indo-West Pacific region (Fischer-Piette and Métivier 1971): R. philippinarum, R. variegatus and R. bruguieri. Genetic differentiation of R. variegatus from R. philippinarum and R. form was very large, as shown in our phylogeny tree and in the STRUCTURE barplot of Vargas et al. (2010). R. variegatus is easy to identify from its shell morphology and habitat. It is smaller, usually not exceeding 30 mm SL (Okutani 2000), and has diverse colour variation in the inner shell. The range of SL for our sample was 25.0–29.6 mm. The shells are thinner than those of R. philippinarum and they inhabit quite a different environment: sand bottoms with rocks in intertidal areas and shallow waters. R. bruguieri (DQ184829) included in our R. variegatus cluster was taken from a tide pool with rocks in Yamaguchi prefecture, Japan, suggesting it was R. variegatus. Vargas et al. (2008) tentatively treated R. form as R. bruguieri based on shell-based taxonomy and allozyme variation. However, R. form is not R. bruguieri, because R. bruguieri appears restricted to the Indian Ocean, and the shell characteristics looked different from R. form in the photographs (p. 428 in Huber 2010). Thus, R. form is not R. variegatus or R. bruguieri, and its taxonomic position remains undefined. Our analyses showed that the genetic differentiation between R. form and R. philippinarum was distinct, but the evolutionary distance was smaller than that between R. variegatus populations collected in Japan and southern China. We reject the hypothesis that R. form is a local population of R. philippinarum because of its distinct shell morphology, especially larger numbers of RR. The results suggested that R. form might be a new Ruditapes species or a variation of R. philippinarum that originated from southern China, and R. variegatus in China and Japan might be a different species or a variation of R. variegatus. Our results highlight the need for a taxonomic revision of species in the genus Ruditapes.
Conservation outcomes
Our study found that R. philippinarum in the Ariake Sea suffered from widespread hybridization with the introduced alien R. form. However, pure R. philippinarum populations remain in the adjacent Shiranui Sea and other areas in Japan. This is a case of Type 5 hybridization according to Allendorf et al. (2001). We recommend that efforts focus on maintaining and expanding remaining pure populations such as those in the Shiranui Sea. Introduction of pure R. philippinarum individuals or releases of artificially produced seed produced from pure parents may be used to recover the hybrid swarm to the original pure R. philippinarum population. However, hybrid individuals backcross with introduced pure individuals, which leads the population in the Ariake Sea to a hybrid swarm again after several generations. In addition, hybrids may expand to the adjacent Shiranui Sea through larval movement. Therefore, the only way to conserve pure R. philippinarum populations is to remove the hybrids from the Ariake Sea. Hybrid populations are of little conservation value (Allendorf et al. 2001), but those in the Ariake Sea have economic value. Therefore, the ideal management option is to completely harvest the hybrids by strong fishing pressure. Introduction of pure R. philippinarum individuals from the Shiranui Sea or releases of artificially produced seeds produced from the pure parents will help speed the recovery.
It is essential to prohibit release of alien R. form into natural waters immediately. Conservation of pure R. philippinarum populations remaining in Japan is also very important. In Japan, the Invasive Alien Species Act (established in 2004) listed 105 species and prohibits releases of the listed species into the natural environment, but no marine animal has yet been listed. Based on our data we recommend that R. form should be listed. However, it is a controversial issue with stakeholders such as fishermen, managers of recreational shell gathering and import companies. Continuous efforts should also be made to measure the introgression of R. form in other areas in Japan with additional samples from the Ariake Sea.
References
Akaike H (1974) A new look at the statistical model identification. IEEE Trans Autom Control 19:716–723
Allendorf FW, Leary RF, Spruell P, Wenburg JK (2001) The problems with hybrids: setting conservation guidelines. Trends Ecol Evol 16:613–622
Anderson EC, Waples RS, Kalinowski ST (2008) An improved method for predicting the accuracy of genetic stock identification. Can J Fish Aquat Sci 65:1475–1486
Bandelt H-J, Foster P, Röhl A (1999) Median-joining networks inferring intraspecific phylogenies. Mol Biol Evol 16:37–48
Bax N, Williamson A, Aguero M, Gonzalez E, Geeves W (2003) Marine invasive alien species: a threat to global biodiversity. Mar Policy 27:313–323
Becker P, Barringer C, Marelli DC (2008) Thirty years of sea ranching Manila clams (Venerupis philippinarum): successful techniques and lessons learned. Rev Fish Sci 16:44–50
Chen J, Qi Li Q, Kong L, Yu H (2011) How DNA barcodes complement taxonomy and explore species diversity: the case study of a poorly understood marine fauna. PLoS One 6(6):e21326. doi:10.1371/journal.pone.0021326
Chiesa S, Nonnis Marzano F, Minervini G et al (2011) The invasive Manila clam Ruditapes philippinarum (Adams and Reeve, 1850) in Northern Adriatic Sea: population genetics assessed by an integrated molecular approach. Fish Res 110:259–267
Cox DR (1975) Partial likelihood. Biometrika 62:269–276
Dempster AP, Laird NM, Rubin DB (1977) Maximum likelihood from incomplete data via the EM algorithm. J R Stat Soc B39:415–423
Drummond L, Mulcahy M, Culloty S (2006) The reproductive biology of the Manila clam, Ruditapes philippinarum, from the north-west of Ireland. Aquaculture 254:326–340
Excoffier L, Laval G, Schneider S (2005) Arlequin ver. 3.0: an integrated software package for population genetics data analysis. Evol Bioinformatics Online 1:47–50
Felsenstein J (1981) Evolutionary trees from DNA sequences: a maximum likelihood approach. J Mol Evol 17:368–376
Fischer-Piette E, Métivier B (1971) Révision de Tapetinae (mollusques bivalves). Mémoires du Muséum National D′Histoire Naturelle. Ser A (Zool) 71:1–106
Fisheries Research Agency and National Association for the Promotion of Productive Seas (2011) Annual statistics of seed production and release in 2009. National Association for the Promotion of Productive Seas, Tokyo (in Japanese)
Fitch WM, Margoliash E (1967) A method for estimating the number of invariant amino acid coding positions in a gene using cytochrome c as a model case. Biochem Genet 1:65–71
Flye-Sainte-Marie J, Jean F, Paillard C et al (2007) Ecophysiological dynamic model of individual growth of Ruditapes philippinarum. Aquaculture 266:130–143
Folmer O, Black M, Hoeh W, Lutz R, Vrijenhoek R (1994) DNA primers for amplification of mitochondrial cytochrome c oxidase subunit I from diverse metazoan invertebrates. Mol Mar Biol Biotechnol 3(5):294–299
Gosling E (2003) Bivalve molluscs: biology, ecology and culture. Fishing News Books, Oxford
Guindon S, Gascuel O (2003) A simple, fast, and accurate algorithm to estimate large phylogenies by maximum likelihood. Syst Biol 52:696–704
Hardy OJ, Vekemans X (2002) SPAGeDi: a versatile computer program to analyse spatialgenetic structure at the individual or population levels. Mol Ecol Notes 2:618–620
Hasegawa M, Kishino H, Yano T (1985) Dating of the human–ape splitting by a molecular clock of mitochondrial DNA. J Mol Evol 22:160–174
Huber M (2010) Compendium of bibalves. Conch Books, Hackenheim
Hurtado NS, Pérez-García C, Morán P, Pasantes JJ (2011) Genetic and cytological evidence of hybridization between native Ruditapes decussatus and introduced Ruditapes philippinarum (Mollusca, Bivalvia, Veneridae) in NW Spain. Aquaculture 311:123–128
Hurvich CM, Tsai C-L (1989) Regression and time series model selection in small samples. Biometrika 76:297–307
Kanemori Y, Takegaki T, Natsukari Y (2006) Genetic population structure of the mudskipper Boleophthalmus pectinirostris inferred from mitochondrial DNA sequences. Japan J Ichthyol 53(2): 133–141 (in Japanese with English abstract)
Kishino H, Kitada S, Hiramatsu K (1994) Sampling scheme for the estimation of the stock composition in the mixed population based on genetic data. Nippon Suisan Gakkaishi 60:359–364 (in Japanese with English abstract)
Kitada S, Kitakado T, Kishino H (2007) Empirical Bayes inference of F ST and its distribution in the genome. Genetics 177:861–873
Librado P, Rozas J (2009) DnaSP v5: a software for comprehensive analysis of DNA polymorphism data. Bioinformatics 25:1451–1452
Mao Y, Gao T, Yanagimoto T, Xiao Y (2011) Molecular phylogeography of Ruditapes philippinarum in the Northwestern Pacific Ocean based on COI gene. J Exp Mar Biol Ecol 407:171–181
Mikkelsen PM, Rudiger B, Kappner I, Rawlings TA (2006) Phylogeny of Veneroidea (Mollusca: Bivalvia) based on morphology and molecules. Zool J Linn Soc 148:439–521
Millar RB (1987) Maximum likelihood estimation of mixed stock fishery composition. Can J Fish Aquat Sci 44:583–590
Millar RB (1990) Comparison of methods for estimating mixed stock fishery composition. Can J Fish Aquat Sci 47:2235–2241
Ministry of Agriculture, Forestry and Fisheries (1967–2011) Annual statistics of fisheries and aquaculture production in 1965–2009. Association of Agriculture and Forestry Statistics, Tokyo (in Japanese)
Ministry of Finance (1991–2010) Trade Statistics of Japan in 1989–2008. Ministry of Finance, Tokyo (in Japanese)
Nei M (1978) Estimation of average heterozygosity and genetic distance from a small number of individuals. Genetics 89:583–590
Nei M (1987) Molecular evolutionary genetics. Columbia University Press, New York
Okutani T (2000) Marine mollusks in Japan. Tokai University Press, Tokyo (in Japanese)
Palsbøll PJ, Bérubé M, Allendorf FW (2006) Identification of management units using population genetic data. Trends Ecol Evol 22:11–16
Passamonti M, Scali V (2001) Gender-associated mitochondrial DNA heteroplasmy in the venerid clam Tapes philippinarum (Mollusca: Bivalvia). Curr Genet 39:117–124
Passamonti M, Boore JL, Scali V (2003) Molecular evolution and recombination in gender-associated mitochondrial DNAs of the Manila clam Tapes philippinarum. Genetics 164:603–611
Peakall R, Smouse PE (2006) GENALEX6: genetic analysis in Excel. Population genetic software for teaching and research. Mol Ecol Note 6:288–295
Pella J, Masuda M (2001) Bayesian methods for analysis of stock mixtures from genetic characters. Fish Bull 99:151–167
Plazzi F, Passamonti M (2010) Towards a molecular phylogeny of mollusks: bivalves’ early evolution as revealed by mitochondrial genes. Mol Phylogenet Evol 57:641–657
Posada D (2008) jModelTest: phylogenetic model averaging. Mol Biol Evol 25:1253–1256
Posada D, Buckley TR (2004) Model selection and model averaging in phylogenetics: advantages of Akaike information criterion and Bayesian approaches over likelihood ratio tests. Syst Biol 53:793–808
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959
Raymond M, Rousset F (1995a) GENEPOP (version 3.4): population genetics software for exact tests and ecumenicism. J Hered 86:248–249
Raymond M, Rousset F (1995b) An exact test for population differentiation. Evolution 49:1280–1283
Rice RW (1989) Analyzing tables of statistical tests. Evolution 43:223–225
Schwarz G (1978) Estimating the dimension of a model. Ann Stat 6:461–464
Sekine Y, Yamakawa H, Takazawa S, Lin Y, Toba M (2006) Geographic variation of the COX1 gene of the short-neck clam Ruditapes philippinarum in coastal regions of Japan and China. Venus 65:229–240 (in Japanese with English abstract)
Shoemaker JS, Fitch WM (1989) Evidence from nuclear sequences that invariable sites should be considered when sequence divergence is calculated. Mol Biol Evol 6:270–289
Shpigel M, Spencer B (1996) Performance of diploid and triploid Manila clams (Tapes philippinarum, Adams and Reeve) at various levels of tidal exposure in the UK and in water from fish ponds at Eilat, Israel. Aquaculture 141:159–171
Slatkin M (1995) A measure of population subdivision based on microsatellite allele frequencies. Genetics 139:457–462
Sugiura N (1978) Further analysis of the data by Akaike’s information criterion and the finite corrections. Commun Stat A 7:13–26
Tajima F (1983) Evolutionary relationship of DNA sequences in finite populations. Genetics 105:437–460
Tamura K, Nei M (1993) Estimation of the number of nucleotide substitutions in the control region of mitochondrial DNA in humans and chimpanzees. Mol Biol Evol 10:512–526
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S (2011) MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28:2731–2739
Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG (1997) The ClustalX windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res 24:4876–4882
Utting SD, Spencer BE (1992) Introductions of marine bivalve molluscs into the United Kingdom for commercial culture—case histories. ICES Mar Sci Symp 194:84–91
Vargas K, Asakura Y, Ikeda M et al (2008) Allozyme variation of littleneck clam Ruditapes philippinarum and genetic mixture analysis of foreign clams in Ariake Sea and Shiranui Sea off Kyushu Island, Japan. Fish Sci 74:533–543
Vargas K, Hamasaki K, Asakura Y et al (2010) Genetic diversity and differentiation in allozymes and shell sculpture of the clams Ruditapes spp. in natural populations of Japan and China. Fish Genet Breed Sci 40:37–46
Williams JG (1980) Growth and survival in newly settled spats of the Manila clam Tapes japonica. Fish Bull 77:891–900
Yamakawa AY, Imai H (2012) Hybridization between Meretrix lusoria and the alien congeneric species M. petechialis in Japan as demonstrated using DNA markers. Aquat Invasions 7:327–336
Yang Z (1993) Maximum-likelihood estimation of phylogeny from DNA sequences when substitution rates differ over sites. Mol Biol Evol 10:1396–1401
Yasuda N, Nagai S, Yamaguchi S, Lian C, Hamaguchi M (2007) Development of microsatellite markers for the Manila clam Ruditapes philippinarum. Mol Ecol Notes 7:43–45
Zhang G, Yan X (2006) A new three-phase culture method for Manila clam, Ruditapes philippinarum, farming in northern China. Aquaculture 258:452–461
Acknowledgments
We thank Robert Toonen and three anonymous reviewers for constructive comments on earlier versions of the manuscript. This study was funded by the Japan Society for the Promotion of Science to SK (Funding No. 18380114 and 22380110). We thank Takuma Sugaya for his advice on molecular experiments, and Mitsuharu Toba, Masaaki Kuroda, Takahiro Kudo, Toyoshige Yanagisawa, Sun Zonghmin, and staff of Dalian Ocean University and Hainan University for their kind assistance during the fieldwork in Japan and China. We also thank Jiro Tanaka and Kotaro Tsuchiya for advice on the taxonomy of Ruditapes species.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Appendix
Appendix
Maximum likelihood estimation of mixing proportion and variance formula
Estimation of stock composition has been studied for genetic stock identification (GSI) (e.g., Millar 1987, 1990); however, the conditional likelihood estimation ignores the sampling errors in the baseline data. The variances of the proportion estimates are therefore underestimated from the method. Kishino et al. (1994) explicitly derived variance formulae of stock composition estimates for GSI by a conditional likelihood approach for single-locus data. The method evaluates sampling variances separately from baseline and mixed populations, which therefore can determine sampling strategies on sample sizes to be taken from baseline and mixed populations. Pella and Masuda (2001) developed a Bayesian method for GSI, which overcomes the problem of the unevaluated sampling errors in the baseline data introducing Dirichlet prior distributions for allele frequencies of the baseline populations. Anderson et al. (2008) developed a new method yielding essentially unbiased estimates of GSI accuracy based on leave-one-out cross-validation. Here we extend the approach of Kishino et al. (1994) to multilocus data (Kishino unpublished) as follows.
Single-locus data
Consider a mixed population composed of k baseline populations, (j = 1,…,k), and j th stock has m genotypes on a locus with probabilities \( {\mathbf{p}}^{(j)} = (p_{1}^{(j)} , \ldots ,p_{m - 1}^{(j)} )^{\prime},\;p_{m}^{(j)} \; = \;1 - p_{1}^{(j)} - \ldots - p_{m - 1}^{(j)} \). Here the prime refers to the transpose of the matrix. Let the proportions of the baseline populations in the mixed population be \( {\varvec{\theta}} = (\theta_{1} , \ldots ,\theta_{k - 1} )^{\prime},\quad \theta_{k} = 1 - \theta_{1} - \ldots - \theta_{k - 1} \) ith genotype frequency of the mixed population is expressed by the sum of the i th genotype frequency of the baseline populations as \( {p}_{i}^{(0)} = \theta_{1}{p}_{i}^{(1)}+..+ \theta_{k}{p}_{i}^{(k)}\) (i = 1,···, m).
The likelihood function for the samples from baseline and mixed populations is:
where \( {\mathbf{n}}^{(0)} = (n_{1}^{(0)} , \ldots ,n_{m}^{(0)} )^{\prime} \) is the number of individuals that have each genotype taken from the mixed population and \( {\mathbf{n}}^{(j)} = (n_{1}^{(j)} , \ldots ,n_{m}^{(j)} )^{\prime}(j = 1, \ldots ,k) \) is that from the baseline population j. The first term of Eq. (1) refers to the likelihood for a mixed population, where \( n_{i}^{(0)} \) is the number of individuals that have allele i in the sample drawn from the mixed population. The second term in Eq. (1) refers to the likelihood of the baseline populations, where \( n_{i}^{(j)} \) is the number of individuals that have allele i in the sample taken from the baseline population j. In place of maximizing Eq. (1) simultaneously with respect to \( {\varvec{\theta}},{\mathbf{p}}^{(1)} , \ldots ,{\mathbf{p}}^{(k)} \), we first maximize the second term and estimate the baseline genotype frequencies by \( \hat{p}_{i}^{(j)} = {{n_{i}^{(j)} } \mathord{\left/ {\vphantom {{n_{i}^{(j)} } {n_{{_{i.} }}^{(j)} }}} \right. \kern-0pt} {n_{{_{i.} }}^{(j)} }} \), where \( n_{{_{i.} }}^{(j)} = \sum\nolimits_{i = 1}^{m} {n_{{_{i} }}^{(j)} } \). Substitute these estimates to the first term of Eq. (1) and obtain \( \widehat{{\varvec{\theta}}} \) by maximizing Eq. (1) with respect to \( {\varvec{\theta}} \). This is a conditional likelihood approach (Cox 1975). Estimates are obtained by an EM algorithm to renew \( \widehat{{\varvec{\theta}}} \) (Dempster et al. 1977) in each step. Given \( \theta_{j}^{h} \) in step h, \( \theta \) in step (h + 1) is calculated by:
where \( \widehat{p}_{i}^{(0)} = n_{i}^{(0)} /n_{.}^{(0)} \). This iterative procedure is continued to the convergence.
The variance matrix of \( \widehat{{\varvec{\theta}}} \) is evaluated by:
where \( \nabla_{\theta } \nabla^{\prime}_{{p^{(j)} }} l_{1} = {{\partial^{2} l_{1} } \mathord{\left/ {\vphantom {{\partial^{2} l_{1} } {\partial \theta_{j} }}} \right. \kern-0pt} {\partial \theta_{j^2} }} \) and \( \nabla_{\theta } \nabla^{\prime}_{{p^{(j)} }} l_{1} = {{\partial^{2} l_{1} } \mathord{\left/ {\vphantom {{\partial^{2} l_{1} } {\partial \theta_{j} }}} \right. \kern-0pt} {\partial \theta_{j} }}\partial_{{p_{i} }}^{(j)} \). Each component of the variance matrix can be obtained explicitly. The first term in {} in Eq. (2) provides the variance for sampling from the mixed stock, and the second term is the variance for sampling from the baselines. In the ordinal full likelihood approach, the variances of the parameters are numerically estimated from the inverse of the Hessian matrix. These variances include sampling errors from both baseline and mixed populations, and it is not possible to evaluate these errors separately. In our approach, the variances from mixed and baseline stocks can be clearly evaluated by explicitly writing down the variances given in Eq. (2). If we neglect the sampling variances in estimating allele frequencies of the baseline populations, the second term in {} equals 0 and the variance matrix becomes \( \widehat{V}(\widehat{{\varvec{\theta}}}) = \left( { - \nabla_{{\varvec{\theta}}} \nabla^{\prime}_{{\varvec{\theta}}} l_{1} } \right)^{ - 1} \).
Multi-locus data
Assuming linkage equilibriums between loci, the likelihood function across–over loci is the product of the likelihood function across loci:
where v is the number of loci (u = 1, …, v).Similarly, for the single-locus model, maximum likelihood estimates of the mixing proportion \( \widehat{{\varvec{\theta}}} \) are obtained by an EM algorithm. Given \( \theta_{j}^{h} \) in step h, \( \theta \) in step (h + 1) is calculated by:
where \( \widehat{p}_{i}^{(0,u)} = n_{i}^{(0,u)} /n_{.}^{(0,u)} \). This iterative procedure is continued to the convergence.
The variance matrix of \( {\widehat{\varvec{\theta }}} \) is evaluated by:
Each component of the variance matrix can be explicitly given similarly to the single-locus model.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 2.0 International License ( https://creativecommons.org/licenses/by/2.0 ), which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
About this article
Cite this article
Kitada, S., Fujikake, C., Asakura, Y. et al. Molecular and morphological evidence of hybridization between native Ruditapes philippinarum and the introduced Ruditapes form in Japan. Conserv Genet 14, 717–733 (2013). https://doi.org/10.1007/s10592-013-0467-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10592-013-0467-x