Abstract
A primary challenge to managing emerging infectious disease is identifying pathways that allow pathogen transmission at the human–wildlife interface. Using Escherichia coli as a model organism, we evaluated fecal bacterial transmission between banded mongoose (Mungos mungo) and humans in northern Botswana. Fecal samples were collected from banded mongoose living in protected areas (n = 87, 3 troops) and surrounding villages (n = 92, 3 troops). Human fecal waste was collected from the same environment (n = 46). Isolates were evaluated for susceptibility to 10 antibiotics. Resistant E. coli isolates from mongoose were compared to human isolates using rep-PCR fingerprinting and MLST-PCR. Antimicrobial resistant isolates were identified in 57 % of the mongoose fecal samples tested (range 31–78% among troops). At least one individual mongoose fecal sample demonstrated resistance to each tested antibiotic, and multidrug resistance was highest in the protected areas (40.9%). E. coli isolated from mongoose and human sources in this study demonstrated an extremely high degree of genetic similarity on rep-PCR (AMOVA, F ST = 0.0027, p = 0.18) with a similar pattern identified on MLST-PCR. Human waste may be an important source of microbial exposure to wildlife. Evidence of high levels of antimicrobial resistance even within protected areas identifies an emerging health threat and highlights the need for improved waste management in these systems.
Similar content being viewed by others
Introduction
As human populations grow and transform landscapes, contact with wildlife concomitantly increases. Disease emergence has been an important consequence of this escalation in interaction, with the majority of emerging infectious diseases in humans now arising from wildlife reservoirs (Jones et al. 2008). Zoonotic disease is also an increasing threat to humanity globally, crossing cultural and socioeconomic lines with developed and developing nations equally at risk (Mayer 2000). With increased travel and global interconnectedness, emergence of zoonotic disease in any one place has the potential to spark a global pandemic, as seen with sudden acute respiratory syndrome (Dye and Gay 2003) and swine flu (H1N1 influenza) (Neumann et al. 2009). Indeed, some of the most devastating and persistent human pathogens can be traced to zoonotic origins, (i.e., human immunodeficiency virus/acquired immunodeficiency syndrome (Hahn et al. 2000) and influenza (Schoub 2012)).
Pathogen transmission from humans to wildlife is also an important emerging threat, increasing wildlife conservation and management challenges (Deem et al. 2001). For example, emergence of human metapneumovirus morbidity and mortality in mountain gorillas (Gorilla beringei beringei) in Rwanda is associated with human–gorilla contact related to ecotourism activities (Palacios 2011). Similar studies demonstrate the increasing threat of human infectious diseases to non-human primate species (Nizeyi et al. 2001; Graczyk et al. 2002), elephants (Michalak et al. 1998), sea birds (Steele et al. 2005), and reptiles (Wheeler et al. 2012). These examples illustrate the bidirectional potential for pathogen transmission at the human–wildlife interface.
Human modification of the environment is seen as a primary driver of the emergence of zoonotic disease, providing the opportunity for direct and indirect contact between humans and wildlife and increasing pathogen exposure and transmission potential (Mayer 2000; Deem et al. 2001). These changes can induce immediate as well as long-term effects on pathogen transmission dynamics, modifying genetic and biological characteristics, biophysical elements, ecological dynamics, and socioeconomics, as well as host(s)–pathogen interactions (Smolinski et al. 2003). Human fecal waste is an anthropogenic disturbance that persists across human-occupied landscapes, both in developed and developing nations, serving as an important source of environmental pathogens (Cilimburg et al. 2000; Griffin et al. 2003). Sewage sludge and refuse hosts an array of pathogenic bacteria, viruses, protozoa, and helminthes that can lead to outbreaks of disease (Hanks 1967; Burge and Marsh 1978; Arthurson 2008). Contamination of drinking water by sewage effluent is a recurring cause of gastroenteritis leading to morbidity and mortality worldwide (D’Antonio et al. 1985; Ashbolt 2004). This is especially true in developing nations where growing populations often lack regulated waste management and sanitation infrastructure. While environmental fecal waste may be an important source of pathogen exposure for both wildlife and humans, we still have a limited understanding of the complex process of pathogen spillover between wildlife and humans. The relative infrequency of pathogen spillover events limits our ability to evaluate the complexity of interacting and cascading factors driving this process.
Recent studies demonstrate the utility of Escherichia coli as a model for examining transmission of fecal microorganisms between humans and wildlife (Singh et al. 2002; Chapman et al. 2006; Skurnik et al. 2006; Goldberg et al. 2007, 2008; Rwego et al. 2008a, b; Farnleitner et al. 2010; Szekely et al. 2010). E. coli is a bacterium found in the gastrointestinal tract and feces of all mammals. While the majority of E. coli are important commensals, some strains can be pathogenic (e.g., Enterotoxigenic E. coli) in humans, domestic animals, and wildlife. For example, E. coli is one of the leading causes of diarrheal-related deaths worldwide (Gaedirelwe 2008; Drummond 2010; Stecher et al. 2012). While there are a number of limitations to the use of E. coli (Desmarais et al. 2002) as a model organism, it is still regarded as the global standard for detection of fecal contamination, indicating an increased possibility for the presence of enteric pathogens.
In northern Botswana, humans live in close association with a variety of wildlife species. One species in particular is the banded mongoose (Mungos mungo). Banded mongoose occur across protected and unprotected landscapes with significant contact with humans in this system, coupled with a preference for denning and foraging in anthropogenic features (Alexander et al. 2010). Banded mongoose also utilize garbage resources and will search for insects in fecal waste (including human) found in the environment. This species is also infected with two important emerging pathogens: the globally important zoonotic pathogen, leptospirosis (Jobbins et al. 2013) and the newly discovered Mycobacterium mungi, a member of the Mycobacterium tuberculosis complex (Alexander et al. 2010). It is unclear at present if M. mungi has the potential to infect humans or other wildlife in the system. Banded mongoose’s contact with other wildlife and humans makes this species an ideal candidate for evaluating microbial exchange at the human–wildlife interface.
We determined the level of antimicrobial resistance of E. coli isolates collected from feces of banded mongoose occurring across protected and unprotected areas in northern Botswana. We then evaluated the genetic relationships between antibiotic-resistant E. coli isolates from mongoose and isolates from human fecal waste. We used these results to examine the potential for human fecal waste to act as a source gastrointestinal pathogen exposure for wildlife species living in both protected and unprotected areas.
Methods
Study Area
This study was conducted in Chobe District in northern Botswana along the Chobe River, which traverses the Chobe National Park and the associated townships of Kasane and Kazungula (Fig. 1). Chobe District is 20, 800 km2 in size with a relatively low population of ~23,387 people (Central Statistics Office BG 2011) largely restricted to villages surrounding the Chobe National Park. The rest of the land area in the District is designated as protected (e.g., national parks, wildlife management areas, and forest reserves). Chobe District has rich and diverse wildlife resources. Residential and tourism development occur near the protected area network, as does a limited amount of subsistence agriculture. A few permanent tourism facilities are located within the Chobe National Park. In this region, there are no commercial chicken or livestock production systems and agriculture is, therefore, expected to contribute minimally to the accumulation of antibiotic resistance in the region. In this same area, we have established a long-term ecological study of banded mongoose (Mungos mungo) in both protected and unprotected land areas (Alexander et al. 2010).
Map of the study site. In northern Botswana, mongoose troops were sampled (triangles) in Chobe District in protected areas (Chobe National Park and Kasane Forest Reserve, hatched outline, troops living in the protected area are denoted with "+") and towns of Kasane and Kazungula. The Chobe River flows through the Park to industrial, commercial, residential developments (gray circles) and commercial agricultural areas (light gray polygons). The sewage treatment facility is located in the Kazungula area (stripes). Tourism facilities are found within the developed and protected areas (stars). Sources of human waste such as feces and sewage (squares) found in the environment near study troops were sampled as well as the sewage treatment facility.
Banded Mongoose
The banded mongoose host is a small diurnal viverrid, group-living, communal breeder, and a natural inhabitant of wildlife areas in proximity to permanent water. They occur in groups or troops that can number between 8 and 55 individuals and frequently cohabit with humans in towns where they scavenge in human waste and feces. Banded mongoose have a low-skew reproductive strategy; multiple females breed and there is little or no observed suppression of breeding in subordinates (Gilchrist 2006). Dispersal can be infrequent and individuals may be retained within their natal group and breed with relatives upon sexual maturity (Cant et al. 2002).
In our long-term study site, at least one or two animals in each study troop are radio-collared and monitored regularly with spatially referenced habitat, behavioral, clinical, and other relevant ecological data collected on the study population (Laver et al. 2012). Home range data were used to identify mongoose troops as occurring in protected areas or in surrounding villages. This study was conducted under a permit from the Botswana Ministry of Environment, Wildlife, and Tourism and with approval of the Virginia Tech’s Institutional Animal Care and Use Committee (Protocol number 07-146-FIW and 10-154-FIW).
Fecal and Environmental Sample Collection
Fecal samples were collected from banded mongoose living in protected areas (n = 87, 3 troops named CCL+, CGL+, HP+ with “+” denoting protected area) and surrounding villages (n = 92, 3 troops, KUB, CSL, SEF, Fig. 1). With the exception of eight individuals in one troop in the protected area (HP+), all troops had some level of range overlap with human populations. The CCL+ and CGL+ troop home range included tourism facilities within the protected areas where some contact with humans and development would be experienced but at a much lower level than in the village troops. The HP+ troop occurred in a part of the protected area where there was no human infrastructural development or residences other than a dirt road. Feces from these 6 banded mongoose troops were collected following morning latrine behavior. Banded mongoose defecate individually at the same time period and in the same general location each morning upon leaving their den; it is, therefore, possible to collect fecal samples from individual mongoose in each troop without replication during that latrine event. In order to determine the required interval between troop resampling, such that each fecal sample could be considered an independent sampling point for the troop, we measured gut passage time. Colored dye was added to the diet of four captive mongoose (owned by CARACAL Biodiversity Center) and minimum gut fecal passage time was determined to be approximately three days. Based on these results, three sampling efforts were attempted at intervals of not less than 2 weeks to ensure independence of sampling events. In this study, we pooled across sampling events, considering each fecal sample an independent sample given established clearance time and latrine behavior. Fecal samples were collected in sterile tubes shortly after defecation and frozen at −20°C within 8 h of collection until processing (Roesch et al. 2009). Mongoose fecal samples are small and friable with large amounts of insect material, making sectioning of mongoose feces difficult. Mongoose also consume soil while foraging and thus sampling constraints were relaxed and the whole fecal sample was taken and used without regard to exterior or interior components.
Human feces were collected within mongoose home ranges from bush latrines (e.g., on ground defecation) and sewage sludge or wastewater leakage identified in the study area (Fig. 1). Only sections of human feces that did not come in contact with soil were sampled to avoid environmental contamination. Additional samples of sewage were collected from the local wastewater treatment facility. In the case of liquid samples such as wastewater, sterile gauze was swirled in the source to catch particulates along with a water sample in the same sterile tube. Soil samples were collected from areas undisturbed by apparent human or animal waste and were used to provide a limited assessment of natural levels of antimicrobial resistance in the environment.
Microbiology
Fecal and wastewater samples were homogenized in buffered peptone (0.1% BPW Sigma-Aldrich, St. Louis, MO USA), serially diluted 10-fold, and spread plated onto MacConkey Agar (Becton Dickinson Company, Franklin Lakes, NJ, USA) plates. After 24 h at 37°C, six putative E. coli colonies were picked from each mongoose fecal sample. Sewage waste represented an array of individual sources; therefore, 15 colonies were selected from human fecal samples to be used in subsequent analyses. All putative E. coli colonies were grown in tryptic soy broth (Becton Dickinson Company) for further DNA extraction and antibiotic susceptibility testing.
DNA was extracted from putative E. coli colonies using standard methods. Genera-specific primers for malB were used to confirm isolates as E. coli as previously described (Rekha et al. 2006). E. coli isolates were then tested for susceptibility to 10 antibiotics: ampicillin (10 μg), chloramphenicol (30 μg), ciprofloxacin (5 μg), doxycycline (30 μg), gentamycin (10 μg), neomycin (30 μg), streptomycin (10 μg), tetracycline (30 μg), sulfamethoxazole–trimethoprim (SXT) (25 μg), and ceftiofur (30 μg) (BBL, Becton Dickinson Company). Antimicrobials were chosen based on their local availability and on previous studies on microorganism transmission from ecotourism (Rolland et al. 1985; Skurnik et al. 2006; Rwego et al. 2008; Wheeler et al. 2012). Ceftiofur was added to the panel as it is only used in livestock and poultry production and was not available in the study area. Antibiotic susceptibility was measured using a field-modified Kirby-Bauer disk diffusion method with breakpoints indicated by the Clinical Laboratory Standards Institute (CLSI) (2006). In this modification, a McFarland standard was not used to standardize turbidity. Instead, quality results were ensured in several ways. First, E. coli ATCC 25922 culture, a quality assurance indicator strain recommended by CLSI, was treated identically to sample isolates and results remained consistently within required limits for each test batch. Second, any isolates demonstrating intermediate resistance were accepted as susceptible for conservative reporting. Third, since stage of growth can affect resistance, all cultures tested were >24 h of age (which corresponds to stationary phase) in tryptic soy broth at the time of testing. We define “multidrug resistant” isolates as those resistant to 3 or more of the antibiotics screened for susceptibility.
Human antimicrobial resistance data for the region were extracted from the local primary hospital (all data were anonymized, May 2007–July 2011). The research was conducted under permit from the Ministry of Health in Botswana and approval from the Virginia Tech Institutional Review Board (IRB# 11-573). Data were derived from routine testing of clinically isolated microorganisms from various fluids and tested on pre-established antibiotic panels in order to determine susceptibility and treatment regimes. In this clinical setting, intermediate susceptibility is reported as resistant to ensure the most appropriate drug treatment regime.
Genetic Profiling of Mongoose and Human Bacterial E. coli Isolates
While all human E. coli isolates were analyzed for genetic relatedness, only antibiotic-resistant mongoose E. coli isolates were evaluated. Genetic relationships were determined by repetitive polymerase chain reaction (rep-PCR) and multi-locus sequence-type polymerase chain reaction (MLST-PCR). In summary, rep-PCR reactions were performed in 25 μL reactions using 2 μL DNA template (of standardized concentration 100–300 ng/μL) added to 2.5 U Phire Taq (Finnzymes) and 1× of 5× Phire Taq Buffer, 1 μM of BOX AIR primer (5′-CTACGGCAAGGCGACGCTGACG-3′) and an additional 1.5 mM of MgCl2. Cycling was performed using a BioRad MyCycler (Bio-Rad) at 98°C for 3 min, followed by 35 cycles of 98°C for 1 min, 64°C for 8 min, and 71°C for 1 min. Final extension was 15 min at 71°C. PCR fragments (2 μL DNA template of standardized concentration 100–300 ng/μL) were separated by electrophoresis in a 2% agarose gel using 1× TAE buffer at 45 V for 10 h at 4–8°C and visualized. Rep-PCR fingerprint profiles were generated using densitometric (curve-based) genotype determination (Goldberg et al. 2006) with the FPQuest ver 4.5 program (BioRad, Hercules, CA). Genetic diversity of fecal E. coli isolates were assessed using analysis of molecular variance (AMOVA) in the Arlequin program version 3.0 as previously described (Excoffier et al. 1992). Phylogenetic relationships were built through Pearson’s correlation coefficient and neighbor-joining algorithms with optimization parameters identified from previously published guidelines (Goldberg et al. 2006).
To confirm the genetic relationships generated from fingerprint analysis, a subset of isolates representing each phylogenetic clade were amplified and sequenced for MLST-PCR to construct a tree based on composite genotypes at different loci in a single isolate (Archie et al. 2009). Seven housekeeping loci (aspC, clpX, fadD, icdA, lysP, mdh, and uidA) were PCR amplified and sequenced as previously described (STEC 2011). Resulting DNA sequences were aligned and phylogenetic analysis was performed using the program Geneious 5.4.2 (Biomatters) (Drummond 2010). Dendrograms were constructed using neighbor joining and pairwise similarity matrix methods. The tree was rooted with composite sequence of the E. coli K12 from the EcMLST database (www.shigatox.net).
Data Analysis
Fisher’s exact and χ2 tests were used to compare antibiotic resistance levels among groups using the statistical software JMP, version 8.0 (SAS Institute). Exact binomial 95% confidence intervals were calculated using the Epitools package in R (Aragon and Enanoria 2007).
Results
E. coli Isolation and Antibiotic Susceptibility
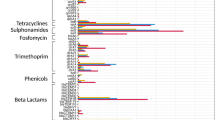
E. coli were isolated from 49% (n = 179) of sampled mongoose feces. No E. coli isolates were obtained from soil samples collected from environments expected to be free of human or animal fecal contamination (n = 13). In mongoose fecal samples where E. coli were isolated, 57% demonstrated antimicrobial resistance. Resistance was identified among individuals in all sampled troops (31–78%, Table 1). No significant differences were observed in the prevalence of antimicrobial resistant E. coli isolates across fecal sampling events for each respective troop (p > 0.05) and data were pooled by troop for analysis. At least one or more mongoose fecal samples demonstrated resistance to each antibiotic tested in this study. Mongoose fecal isolates were most commonly resistant to ampicillin (AM10), followed by doxycyline (D30), tetracycline (Te30), streptomycin (S10), SXT, and chloramphenicol (C30) (Fig. 3; Table 2). A low number of isolates (n = 5) demonstrated resistance to ceftiofur (XNL30), an antibiotic used in livestock and poultry production. Multidrug resistance (7.7–50%) was identified among a number of samples (Table 1). Prevalence of multidrug resistance among sampled mongoose feces was significantly different between sampled troops (p = 0.01, χ2 test). Troop KUB had the lowest level of multidrug resistance among individual fecal samples (7.7% with 95% CI 0.2–36%) while troops SEF (50% with 95% CI 1.3–98.7% n = 2) and CGL+ (40.9% with 95% CI 20.7–63.6% n = 22) had the highest levels. There were no significant differences in resistance to two or fewer antimicrobials (p = 0.11) among fecal samples from study troops. Troop CGL+, located within the protected area, did not have the highest prevalence of resistant isolates overall (Fig. 2), but was the only troop where at least one fecal isolate was resistant to each of the antibiotics screened (Fig. 3; Table 2). Troop HP+ had the highest prevalence of antibiotic-resistant isolates although these were resistant to ampicillin only (Fig. 2; Table 2).
Prevalence of antibiotic resistance among total E. coli isolates collected from banded mongoose (n = 253), and human fecal waste (n = 75) in Chobe District Botswana. Resistance data were also extracted from routine laboratory microorganism susceptibility assessments conducted at the Kasane Primary Hosptial within the study site (2007–2011), on pre-established panels of antibiotics. Isolates would originate from patients from all villages in the District. Not every isolate was subjected to the same panel of antibiotics at the hospital and so the number of isolates screened for each drug varied. Antibiotics are listed as abbreviations followed by dosage (μg) in numbers, the number of human isolates tested at the hospital are noted in brackets (e.g., AM10 = ampicillin [n = 420], Te30 = tetracycline [n = 315], D30 = doxycycline, SXT = sulfamethoxazole-trimethaprim [n = 375], S10 = streptomycin, C30 = chloramphenicol [n = 256], XNL30 = ceftiofur, GM10 = gentamycin [n = 521], N30 = neomycin, and CIP5 = ciprofloxacin). Black bars denote antibiotics that were not evaluated at health facilities in Chobe District during the assessment period.
E. coli were isolated from 41.3% (n = 46, feces and sewage) of human fecal waste samples assessed [human feces found directly in the environment: (n = 5), sewage spill (n = 8), and sewage treatment pond samples (n = 6)]. From these samples, 77 E. coli isolates were screened for antibiotic susceptibility and 80.3% (95% CI 69.5–88.5%) were found to be resistant to at least one antibiotic. Of human clinical samples screened at the local hospital, 89.9% (95% CI 87–92% for all years) of isolated microorganisms (various sources and bacteria species) were resistant to at least one antibiotic. Human and banded mongoose E. coli isolates collected in this study demonstrated resistance to the same drugs as microorganisms isolated from patients at the Kasane Primary Hospital (Fig. 3). Overall, resistance in banded mongoose was lower than that found in the human population.
Repetitive BOX-PCR
Using AMOVA, significant variation was identified among sampled mongoose feces (F ST = 0.02, p = 0.01, Table 3), with the majority of the variation identified at the individual fecal sample level (98% of observed variation) rather than at the troop level (2% of the observed variation). However, E. coli strains isolated from the mongoose troop living in an unmodified habitat (HP+) were indistinguishable genetically from those troops living in association with humans in villages or at lodges in protected areas (CSL, CGL+, SEF, KUB, CCL+, F ST = 0, p = 0.64, Table 2). When comparing humans and mongoose isolates, a greater level of genetic diversity of E. coli was found within each species (99.75% of the observed variation) than among species (0.25% of the observed variation). Mongoose and human associated E. coli demonstrated an extremely high degree of genetic similarity (F ST = 0.0027, p = 0.18, Table 3).
Antibiotic-resistant isolates from all troops of mongoose consistently clustered with those of humans and human fecal waste in rep-PCR analysis (Fig. 4). Clades containing exclusively mongoose or human isolates were not observed. Mongoose isolates did not cluster significantly by troop, but instead were highly intermixed with isolates from human sources.
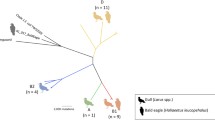
Phylogram of rep-PCR fingerprint analysis of E. coli isolates obtained from human and antibiotic-resistant banded mongoose fecal samples (n = 74 mongoose, n = 87 human). Comparisons were constructed using the Pearson’s correlation and neighbor-joining algorithms. Values alongside branches correspond to % confidence generated from 1,000 bootstraps. Isolates that did not visibly differ were not included. Tree is rooted with the fingerprint profile of E. coli K12 from the EcMLST database.
MLST-PCR
MLST-PCR performed on a subset of E. coli isolates (n = 23) representing each of the clades observed in rep-PCR analysis revealed very slight genetic divergence (maximum < 0.10 base substitutions per site) between isolates from banded mongoose and human fecal waste (Fig. 5). E. coli isolated from banded mongoose and sources of human fecal waste consistently clustered together, forming one monophyletic tree. Genetic relationships between isolates interpreted through branching patterns were not identical between fingerprint- and DNA sequence-based analyses in this study, even when using optimization parameters published by previous studies (Goldberg et al. 2006).
Phylogenetic tree based on E. coli composite sequences from seven housekeeping genes (uidA, mdh, lysP, icdA, clpX, aspC, and fadD) (n = 23). Values alongside branches correspond to % confidence generated from 1,000 bootstraps. Phylogenetic relationships were built from seven consensus trees using Tamura-Nei distance model and neighbor-joining algorithms, and 70% support threshold. The tree is rooted with composite sequence of E. coli K12 from the EcMLST database (www.shigatox.net). Scale is based on the number of base substitutions per site.
Discussion
Using molecular strain comparisons, E. coli isolated from feces of banded mongoose troops demonstrated an extremely high degree of genetic similarity to isolates obtained from human fecal waste in these environments. This was identified among troops with home ranges in both protected and unprotected land areas. Other studies have applied similar molecular techniques demonstrating the potential for microorganism exchange to occur between humans, wildlife, and livestock sharing the same land area. Comparisons between populations focused on the direct collection of fecal material from human volunteers living in these environments (Goldberg et al. 2006; Rwego et al. 2008). While these examinations have provided substantial insight into the occurrence of microbial exchange at the human–animal interface, the specific pathways where gastrointestinal microbial transmission might occur remain elusive. In this study, we focused our sampling strategy on environmental sources of human fecal waste and compared isolated E. coli to that of isolates collected from mongoose living in these same areas.
Recent studies identify a high degree of genetic similarity among E. coli strains globally, regardless of human or animal origin (Clermont et al. 2011) and suggest that microbial exchange events implied by molecular genetic techniques should be viewed conservatively. However, in this study, we did find significant variation in E. coli by species (human and mongoose, respectively). Our results provide direct support for the possibility that direct human fecal contamination of the environment is an important potential source of microbial exposure and transmission to wildlife living in these areas. This transmission appears to occur even in protected wildlife areas.
Antimicrobial resistance is an important marker for human-source bacteria (Kabler et al. 1964) and previous studies identified antimicrobial resistance in wild populations living in association with human populations (Cole et al. 2005; Skurnik et al. 2006; Goldberg et al. 2007; Rwego et al. 2008; Wheeler et al. 2012). In this study, we found high levels of antimicrobial resistance including multidrug resistance among sampled mongoose across all sample troops irrespective of land use (protected and unprotected).
Important in evaluating the significance of the antimicrobial resistance identified in wildlife is evaluating the occurrence in the resident human population. In this study, resistance to the same antimicrobials were identified among mongoose isolates (although lower levels) as that identified in study-derived human E. coli isolates and patients presenting to the Kasane Primary Hospital (2007–2011, Fig. 3). Most isolates obtained from the mongoose troop living in a natural area of Chobe National Park (HP+) were resistant to only ampicillin, with one isolate demonstrating additional drug resistance. This result was in contrast to the other sample troops living in contact with humans where a greater diversity of drug resistance was identified and a higher prevalence of multidrug resistance occurred (Fig. 2; Table 2). Ampicillin resistance can be naturally acquired from environmental sources (Martinez 2009). Patient isolates from the local hospital were also frequently resistant to ampicillin (Fig. 3). Other studies have observed a gradient effect of antimicrobial resistance among environments and animals in relation to human proximity (Skurnik et al. 2006; Allen et al. 2010; Wheeler et al. 2012). Our study design did not allow for this type of evaluation; however, our limited data suggest that this might be an important area of future work.
Multidrug resistance is increasingly identified as an important global health threat, involving all major microbial pathogens and antimicrobial drugs. Multidrug resistance was identified among isolates collected from troops living in proximity to humans, both in the National Park and in village areas. Of these, CGL+ (Park troop) had the highest level, with half of all sampled mongoose demonstrating multidrug resistance among isolates. In some troops, multidrug-resistant isolates were aggregated to a few individual mongoose fecal samples (e.g., KUB, Kazungula Village), but in the CGL+ troop, multidrug resistance was widespread across individuals and isolates (40.9%, 95% CI 20.7–63.6%). This troop commonly denned in the soak away of the septic tank servicing the employee accommodations and foraged around employee living quarters, including eating food remains from dishes left outside. Humans have also been observed feeding raw meat waste from commercially produced chickens to mongoose at this lodge, potentially contributing to the levels of drug resistance in this troop. Antibiotic resistance to ampicillin and tetracycline has been identified in strains of Salmonella at commercial chicken farms in Botswana (Gaedirelwe 2008) and may be an important source of resistant microbiota to species fed raw animal byproducts. Alternatively, this resistance could be present in the human population because of consumption of meat with antibiotic residues. Wildlife could then be subsequently exposed through human fecal waste. Either or both of these mechanisms may have contributed to the low occurrence of resistance to ceftiofur (XNL30, Table 2), a broad-spectrum veterinary antibiotic not available in the study area. All of the study troops, with the exception of HP + , utilized garbage resources and would have access to such waste albeit at varying levels depending on the facility. Uncontrolled and uncontained waste disposal provides a ubiquitous non-seasonal food resource in both peri-urban environments and protected areas, the latter in association with ecotourism enterprises (Rolland et al. 1985). While ecotourism developments are important for conservation and economic growth, associated human waste (both garbage and feces) remains a primary negative consequence. It is unclear how garbage utilization may contribute to microbial exchange and accumulation of antimicrobial resistance. Given the increasing nature of this anthropogenic landscape modification, it is a crucial area for continued research.
There was a high level of resistance among human isolates collected in this study and those tested at the primary hospital. This may appear surprising at first given poverty and expectations of diminished access to health care and pharmaceuticals that is assumed for much of Africa. However, antibiotics are widely available to patients visiting government hospitals in Botswana, where health care is largely free, as well as local pharmacies where few controls on the dispensing of antibiotics are identified. Decreased regulation of antibiotic use and access is an important contributor to the epidemic spread of resistance (Okeke et al. 1999, 2005a, b) and has likely contributed to the occurrence of increased microbial resistance in this human population.
A number of study limitations could have influenced our results. For example, the potential for contamination of sewage samples with non-human fecal material and soil is a possibility. It is likely, however, that such contamination, if it occurred, would be present at exceptionally low levels relative to the volume of human sewage. Sewage and direct fecal contamination of the only surface water in the region creates another vehicle for microorganism transmission between humans and wildlife and could be an important mechanism of E. coli exchange between humans and mongoose. Antibiotics from local human use may accumulate in both the soil (Tolls 2001) and contaminated surface water (Kummerer 2009), potentially increasing the spread of antibiotic resistance.
Conclusions
Our study suggests that environmentally associated human fecal waste may serve as an important source of microbial transmission to mongoose. Furthermore, widespread accumulation of antimicrobial resistance is identified among mongoose even in protected areas considered to be relatively free from human impacts. The impact of microbial exchange and antibiotic resistance accumulation in mongoose may extend through food webs. Mongoose are eaten by a large number of avian, reptile, and mammalian predators including domestic dogs in this system. Thus, the cascading effects of exposure of wildlife species to human waste-associated microbes can impact an array of susceptible species across an ecosystem and in turn increase human exposure, coupling humans and natural systems in complicated ways.
Human fecal waste and garbage are widespread across many environments and are an increasingly global threat to animal and public health (Rushton 2003; Cantalupo et al. 2011). This phenomenon extends to protected areas, where increasing pressure to develop economic enterprises based on tourism brings people and their waste even to remote areas of a protected area network. In order to control the threat of potential pathogen transmission, closed sewage systems should be required. Feeding of poultry and livestock products from kitchen waste should be actively prohibited to decrease the potential for transmission of resistant bacteria. Waste receptacles need to be completely enclosed and “wildlife-proofed” to discourage wildlife access and exposure.
Transmission of pathogens through fecal waste is likely to be amplified in places such as Africa, where sanitation and water treatment facilities are often poor, and fecal contamination of the environment and limited surface water resources represents a growing problem. Common water resources may represent an important exposure mechanism, increasing the potential for bidirectional microbial traffic and increased health risks to both humans and wildlife living in these systems. The intimate connections between human and animal populations are increasingly evident, as is the need to better understand these system coupling points where human and animal health are interlinked.
References
Alexander, K. A., P. N. Laver, et al. (2010). “Novel Mycobacterium tuberculosis complex pathogen, M. mungi.” Emerging infectious diseases 16(8): 1296.
Allen, H. K., J. Donato, et al. (2010). “Call of the wild: antibiotic resistance genes in natural environments.” Nature Reviews Microbiology 8(4): 251-259.
Aragon TJ, Enanoria WT (2007) Applied Epidemiology Using R, MedEpi Publishing. http://www.medepi.net/epir/index.html. Calendar Time. Accessed 10 June 2010
Archie, E. A., G. Luikart, et al. (2009). “Infecting epidemiology with genetics: a new frontier in disease ecology.” Trends in Ecology and Evolution 24(1): 21-30.
Arthurson, V. (2008). “Proper Sanitization of Sewage Sludge: a Critical Issue for a Sustainable Society.” Applied Environmental Microbiology 74(17): 5267-5275.
Ashbolt, N. J. (2004). “Microbial contamination of drinking water and disease outcomes in developing regions.” Toxicology 198(1): 229-238.
Burge, W. and P. Marsh (1978). “Infectious disease hazards of landspreading sewage wastes.” Journal of Environmental Quality 7(1): 1-9.
Cant, M., E. Otali, et al. (2002). “Fighting and mating between groups in a cooperatively breeding mammal, the banded mongoose.” Ethology 108(6): 541-555.
Cantalupo, P. G., B. Calgua (2011). “Raw sewage harbors diverse viral populations.” mBio 51-11.
Central Statistics Office BG (2011) 2011 Botswana Population and Housing Census, Alphabetical Index of Districts. http://www.cso.gov.bw/media/2011Census_Alphabetical.Index.of.Districts.pdf. Accessed 31 March 2013
Chapman, C. A., M. L. Speirs, et al. (2006). “Life on the edge: Gastrointestinal parasites from the forest edge and interior primate groups.” American Journal of Primatology 68(4): 397-409.
Cilimburg, A., C. Monz, et al. (2000). “Wildland recreation and human waste: A review of problems, practices, and concerns.” Environmental Management 25(6): 587-598.
Clermont, O., M. Olier, et al. (2011). “Animal and human pathogenic Escherichia coli strains share common genetic backgrounds.” Infection, genetics and evolution : journal of molecular epidemiology and evolutionary genetics in infectious diseases 11(3): 654-662.
Clinical and Laboratory Standards Institute (2006). Performance standards for antimicrobial disk susceptibility. Wayne, PA, Clinical Laboratory Standards Institute.
Cole, D., D. J. Drum, et al. (2005). “Free-living Canada geese and antimicrobial resistance.” Emerging Infectious Diseases 11(6): 935-938.
D’Antonio, R. G., R. E. Winn, et al. (1985). “A Waterborne Outbreak of Cryptosporidiosis in Normal Hosts.” Annals of Internal Medicine 103(6): 886.
Deem, S. L., W. B. Karesh, et al. (2001). “Putting theory into practice: Wildlife health in conservation.” Conservation Biology 15(5): 1224-1233.
Desmarais, T. R., H. M. Solo-Gabriele, et al. (2002). “Influence of soil on fecal indicator organisms in a tidally influenced subtropical environment.” Applied and Environmental Microbiology 68(3): 1165-1172.
Drummond AJ, A. B., Buxton S, Cheung M, Cooper A, Duran C, Field M, Heled J, Kearse M, Markowitz S, Moir R, Stones-Havas S, Sturrock S, Thierer T, Wilson A (2010). Geneious ver. 5.1.
Dye, C. and N. Gay (2003). “Epidemiology. Modeling the SARS epidemic.” Science 300(5627): 1884-1885.
Excoffier, L., P. Smouse, et al. (1992). “Analysis of molecular variance inferred from metric distances among DNA haplotypes: application to human mitochondrial DNA restriction data.” Genetics 131(2): 479.
Farnleitner, A. H., G. Ryzinska-Paier, et al. (2010). “Escherichia coli and enterococci are sensitive and reliable indicators for human, livestock and wildlife faecal pollution in alpine mountainous water resources.” Journal of Applied Microbiology 109(5): 1599-1608.
Gaedirelwe, O. G. a. T. K. S. (2008). “The Prevalence and Antibiotic Susceptibility of Salmonella sp in Poultry and Ostrich Samples from Slaughter Houses in Gaborone, Botswana.” Journal of Animal and Veterinary Advances 7(9): 1151-1154.
Gilchrist, J. S. (2006). “Reproductive success in a low skew, communal breeding mammal: the banded mongoose, Mungos mungo.” Behavioral Ecology and Sociobiology 60(6): 854-863.
Goldberg, T. L., T. R. Gillespie, et al. (2008). “Forest fragmentation and bacterial transmission among nonhuman primates, humans, and livestock, Uganda.” Emerging Infectious Diseases 14(9): 1375-1382.
Goldberg, T. L., T. R. Gillespie, et al. (2007). “Patterns of gastrointestinal bacterial exchange between chimpanzees and humans involved in research and tourism in western Uganda.” Biological Conservation 135(4): 511-517.
Goldberg, T. L., T. R. Gillespie, et al. (2006). “Optimization of analytical parameters for inferring relationships among Escherichia coli isolates from repetitive-element PCR by maximizing correspondence with multilocus sequence typing data.” Applied and Environmental Microbiology 72(9): 6049-6052.
Graczyk, T. K., J. Bosco-Nizeyi, et al. (2002). “A single genotype of Encephalitozoon intestinalis infects free-ranging gorillas and people sharing their habitats in Uganda.” Parasitology Research 88(10): 926-931.
Griffin, D. W., K. A. Donaldson, et al. (2003). “Pathogenic human viruses in coastal waters.” Clinical Microbiology Reviews 16(1): 129-143.
Hahn, B. H., G. M. Shaw, et al. (2000). “AIDS as a zoonosis: scientific and public health implications.” Science 287(5453): 607-614.
Hanks, T. (1967). Solid waste/disease relationships, A literature survey. U.S. National Center for Urban and Industrial Health. Cincinnati
Jobbins SE, Sanderson CE, et al. (2013) Leptospirosis in banded mongoose at the human—wildlife interface in northern Botswana. Zoonoses and Public Health (accepted)
Jones, K., N. Patel, et al. (2008). “Global trends in emerging infectious diseases.” Nature 451(7181): 990-993.
Kabler, P. W., H. F. Clark, et al. (1964). “Sanitary Significance of Coliform and Fecal Coliform Organisms in Surface Water.” Public Health Reports 79: 58-60.
Kummerer, K. (2009). “Antibiotics in the aquatic environment–a review–part I.” Chemosphere 75(4): 417-434.
Laver, P., A. Ganswindt, et al. (2012). Non-invasive monitoring of glucocorticoid metabolites in banded mongooses (Mungos mungo) in response to physiological and biological challenges. General and Comparative Endocrinology. 179: 178–183.
Martinez, J. L. (2009). “The role of natural environments in the evolution of resistance traits in pathogenic bacteria.” Proceedings of the Royal Society: Biological sciences 276(1667): 2521-2530.
Mayer, J. (2000). “Geography, ecology and emerging infectious diseases.” Social Science and Medicine 50(7-8): 937-952.
Michalak, K., C. Austin, et al. (1998). “Mycobacterium tuberculosis infection as a zoonotic disease: Transmission between humans and elephants.” Emerging Infectious Diseases 4(2): 283-287.
Neumann, G., T. Noda, et al. (2009). “Emergence and pandemic potential of swine-origin H1N1 influenza virus.” Nature 459(7249): 931-939.
Nizeyi, J. B., R. B. Innocent, et al. (2001). “Campylobacteriosis, salmonellosis, and shigellosis in free-ranging human-habituated mountain gorillas of Uganda.” Journal of Wildlife Diseases 37(2): 239-244.
Okeke, I. N., K. P. Klugman, et al. (2005). “Antimicrobial resistance in developing countries. Part II: strategies for containment.” The Lancet Infectious Diseases 5(9): 568-580.
Okeke, I. N., A. Lamikanra, et al. (1999). “Socioeconomic and behavioral factors leading to acquired bacterial resistance to antibiotics in developing countries.” Emerging Infectious Diseases 5(1): 18-27.
Okeke, I. N., R. Laxminarayan, et al. (2005). “Antimicrobial resistance in developing countries. Part I: recent trends and current status.” The Lancet Infectious Diseases 5(8): 481-493.
Palacios, G. (2011). “Human metapneumovirus infection in wild mountain gorillas, Rwanda.” Emerging Infectious Diseases 17(4): 711-713.
Rekha, R., M. Alam Rizvi, et al. (2006). Designing and validation of genus-specific primers for human gut flora study. Electronic Journal of Biotechnology. 9:505–511.
Roesch, L. F., G. Casella, et al. (2009). “Influence of fecal sample storage on bacterial community diversity.” Open Microbiology Journal 3: 40-46.
Rolland, R. M., G. Hausfater, et al. (1985). “Antibiotic-Resistant Bacteria in Wild Primates - Increased Prevalence in Baboons Feeding on Human Refuse.” Applied and Environmental Microbiology 49(4): 791-794.
Rushton, L. (2003). “Health hazards and waste management.” British Medical Bulletin 68: 183-197.
Rwego, I. B., T. R. Gillespie, et al. (2008). “High rates of Escherichia coli transmission between livestock and humans in rural Uganda.” Journal of Clinical Microbiology 46(10): 3187-3191.
Rwego, I. B., G. Isabirye-Basuta, et al. (2008). “Gastrointestinal bacterial transmission among humans, mountain gorillas, and livestock in Bwindi Impenetrable National Park, Uganda.” Conservation Biology 22(6): 1600-1607.
Schoub, B. D. (2012). “Zoonotic diseases and human health: The influenza example.” Onderstepoort Journal of Veterinary Research 79(2): 4.
Singh, D. K., C. McCormick, et al. (2002). “Pathogenic and nef-interrupted simian-human immunodeficiency viruses traffic to the macaque CNS and cause astrocytosis early after inoculation.” Virology 296(1): 39-51.
Skurnik, D., R. Ruimy, et al. (2006). “Effect of human vicinity on antimicrobial resistance and integrons in animal faecal Escherichia coli.” Journal of Antimicrobial Chemotherapy 57(6): 1215-1219.
Smolinski, M., M. Hamburg, et al. (2003). Microbial threats to health: emergence, detection, and response. Washington, D.C., The National Academies Press.
STEC (2011) http://shigatox.net. Accessed 1 June 2010
Stecher, B., R. Denzler, et al. (2012). “Gut inflammation can boost horizontal gene transfer between pathogenic and commensal Enterobacteriaceae.” Proceedings of the National Academy of Sciences of the United States of America 109(4): 1269-1274.
Steele, C. M., R. N. Brown, et al. (2005). “Prevalences of zoonotic bacteria among seabirds in rehabilitation centers along the Pacific Coast of California and Washington, USA.” Journal of Wildlife Diseases 41(4): 735-744.
Szekely, B. A., J. Singh, et al. (2010). “Fecal bacterial diversity of human-habituated wild chimpanzees (Pan troglodytes schweinfurthii) at Mahale Mountains National Park, Western Tanzania.” American Journal of Primatology 72 (7): 566-574.
Tolls, J. (2001). “Sorption of veterinary pharmaceuticals in soils: a review.” Environmental Science and Technology 35(17): 3397-3406.
Wheeler, E., P. Y. Hong, et al. (2012). “Carriage of antibiotic-resistant enteric bacteria varies among sites in Galapagos reptiles.” Journal of Wildlife Diseases 48(1): 56-67.
Acknowledgments
We would like to thank the Botswana Ministry of Health and the Ministry of Environment Wildlife, and Tourism for permission to conduct this study. We thank M. Vandewalle and P. Laver for their invaluable assistance in this study and E. Hallerman and J. Fox for comments and assistance on this manuscript. This project was funded under the National Science Foundation Coupled Human Environmental Systems Award CNH#1114953, Morris Animal Foundation Award # D10Z0-828A, and the WildiZe Foundation. The National Science Foundation S-STEM Program (DUE-0850198) provided partial financial support for R. Pesapane.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Pesapane, R., Ponder, M. & Alexander, K.A. Tracking Pathogen Transmission at the Human–Wildlife Interface: Banded Mongoose and Escherichia coli . EcoHealth 10, 115–128 (2013). https://doi.org/10.1007/s10393-013-0838-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10393-013-0838-2