Abstract
Objective
Dental caries is caused by acidogenic plaque microbiota formed on saliva-bathed tooth surfaces, in which multiple organisms act collectively to initiate and expand a cavity. We explored bacterial species associated with the salivary microbiome of individuals with low susceptibility to dental caries.
Materials and methods
The bacterial composition of saliva from 19 young adults was analyzed using barcoded pyrosequencing of the 16S rRNA gene; we compared 10 caries-experienced (CE) and nine caries-free (CF) individuals. A quantitative PCR assay of saliva from 139 orally healthy adults aged 40–59 years was carried out to confirm the result obtained by pyrosequencing analysis.
Results
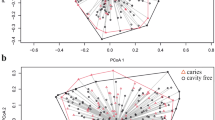
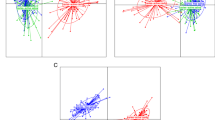
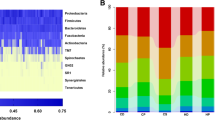
The microbiomes of CF individuals showed more diverse communities with a significantly greater proportion of the genus Porphyromonas. Among operational taxonomic units (OTUs) corresponding to the genus Porphyromonas, the OTU corresponding to P. pasteri was the most predominant and its relative abundance in CF individuals was significantly greater than in CE individuals (P < 0.001, Wilcoxon rank sum test). A quantitative PCR assay of saliva confirmed that the amounts of P. pasteri were significantly higher in individuals with lower caries experience (filled teeth <15, n = 67) than in those with higher caries experience (filled teeth ≥15, n = 72) (P < 0.001, Student’s t test).
Conclusion
These results revealed an association between a greater abundance of P. pasteri and lower susceptibility to dental caries.
Clinical relevance
P. pasteri may be a bacterial species that could potentially be used as a marker for maintaining a healthy oral microbiome against dental caries.




Similar content being viewed by others
References
Belstrom D, Fiehn NE, Nielsen CH, Holmstrup P, Kirkby N, Klepac-Ceraj V, Paster BJ et al (2014) Altered bacterial profiles in saliva from adults with caries lesions: a case-cohort study. Caries Res 48:368–375
Burne RA, Marquis RE (2000) Alkali production by oral bacteria and protection against dental caries. FEMS Microbiol Lett 193:1–6
Caporaso JG, Kuczynski J, Stombaugh J, Bittinger K, Bushman FD, Costello EK, Fierer N et al (2010) QIIME allows analysis of high-throughput community sequencing data. Nat Methods 7:335–336
Chen T, Yu WH, Izard J, Baranova OV, Lakshmanan A and Dewhirst FE (2010) The Human Oral Microbiome Database: a web accessible resource for investigating oral microbe taxonomic and genomic information. Database (Oxford):baq013
Chhour KL, Nadkarni MA, Byun R, Martin FE, Jacques NA, Hunter N (2005) Molecular analysis of microbial diversity in advanced caries. J Clin Microbiol 43:843–849
Crielaard W, Zaura E, Schuller AA, Huse SM, Montijn RC, Keijser BJ (2011) Exploring the oral microbiota of children at various developmental stages of their dentition in the relation to their oral health. BMC Med Genet 4:22
Edgar RC (2013) UPARSE: highly accurate OTU sequences from microbial amplicon reads. Nat Methods 10:996–998
Eren AM, Borisy GG, Huse SM, Mark Welch JL (2014) Oligotyping analysis of the human oral microbiome. Proc Natl Acad Sci U S A 111:E2875–E2884
Gomar-Vercher S, Cabrera-Rubio R, Mira A, Montiel-Company JM, Almerich-Silla JM (2014) Relationship of children's salivary microbiota with their caries status: a pyrosequencing study. Clin Oral Investig 18:2087–2094
Gross EL, Beall CJ, Kutsch SR, Firestone ND, Leys EJ, Griffen AL (2012) Beyond Streptococcus Mutans: dental caries onset linked to multiple species by 16S rRNA community analysis. PLoS One 7:e47722
Gross EL, Leys EJ, Gasparovich SR, Firestone ND, Schwartzbaum JA, Janies DA, Asnani K et al (2010) Bacterial 16S sequence analysis of severe caries in young permanent teeth. J Clin Microbiol 48:4121–4128
Haas BJ, Gevers D, Earl AM, Feldgarden M, Ward DV, Giannoukos G, Ciulla D et al (2011) Chimeric 16S rRNA sequence formation and detection in sanger and 454-pyrosequenced PCR amplicons. Genome Res 21:494–504
Huang X, Exterkate RA, ten Cate JM (2012) Factors associated with alkali production from arginine in dental biofilms. J Dent Res 91:1130–1134
Kassebaum NJ, Bernabe E, Dahiya M, Bhandari B, Murray CJ, Marcenes W (2015) Global burden of untreated caries: a systematic review and metaregression. J Dent Res 94:650–658
Kolenbrander PE, Palmer RJ Jr, Periasamy S, Jakubovics NS (2010) Oral multispecies biofilm development and the key role of cell-cell distance. Nat Rev Microbiol 8:471–480
Ling Z, Kong J, Jia P, Wei C, Wang Y, Pan Z, Huang W et al (2010) Analysis of oral microbiota in children with dental caries by PCR-DGGE and barcoded pyrosequencing. Microb Ecol 60:677–690
Loesche WJ (1986) Role of Streptococcus mutans in human dental decay. Microbiol Rev 50:353–380
Lozupone C, Knight R (2005) UniFrac: a new phylogenetic method for comparing microbial communities. Appl Environ Microbiol 71:8228–8235
Luo AH, Yang DQ, Xin BC, Paster BJ, Qin J (2012) Microbial profiles in saliva from children with and without caries in mixed dentition. Oral Dis 18:595–601
Marcenes W, Kassebaum NJ, Bernabe E, Flaxman A, Naghavi M, Lopez A, Murray CJ (2013) Global burden of oral conditions in 1990-2010: a systematic analysis. J Dent Res 92:592–597
Moritani K, Takeshita T, Shibata Y, Ninomiya T, Kiyohara Y, Yamashita Y (2015) Acetaldehyde production by major oral microbes. Oral Dis 21:748–754
Obata J, Takeshita T, Shibata Y, Yamanaka W, Unemori M, Akamine A, Yamashita Y (2014) Identification of the microbiota in carious dentin lesions using 16S rRNA gene sequencing. PLoS One 9:e103712
Price MN, Dehal PS, Arkin AP (2009) FastTree: computing large minimum evolution trees with profiles instead of a distance matrix. Mol Biol Evol 26:1641–1650
Rasiah IA, Wong L, Anderson SA, Sissons CH (2005) Variation in bacterial DGGE patterns from human saliva: over time, between individuals and in corresponding dental plaque microcosms. Arch Oral Biol 50:779–787
Sakamoto M, Li D, Shibata Y, Takeshita T, Yamashita Y and Ohkuma M (2015) Porphyromonas pasteri sp. nov., isolated from human saliva. Int J Syst Evol Microbiol
Simon-Soro A, Mira A (2015) Solving the etiology of dental caries. Trends Microbiol 23:76–82
Stahringer SS, Clemente JC, Corley RP, Hewitt J, Knights D, Walters WA, Knight R et al (2012) Nurture trumps nature in a longitudinal survey of salivary bacterial communities in twins from early adolescence to early adulthood. Genome Res 22:2146–2152
Takeshita T, Matsuo K, Furuta M, Shibata Y, Fukami K, Shimazaki Y, Akifusa S et al (2014) Distinct composition of the oral indigenous microbiota in South Korean and Japanese adults. Sci Rep 4:6990
Takeshita T, Nakano Y, Kumagai T, Yasui M, Kamio N, Shibata Y, Shiota S et al (2009) The ecological proportion of indigenous bacterial populations in saliva is correlated with oral health status. ISME J 3:65–78
Takeshita T, Yasui M, Shibata Y, Furuta M, Saeki Y, Eshima N, Yamashita Y (2015) Dental plaque development on a hydroxyapatite disk in young adults observed by using a barcoded pyrosequencing approach. Sci Rep 5:8136
Tong H, Chen W, Merritt J, Qi F, Shi W, Dong X (2007) Streptococcus oligofermentans inhibits Streptococcus mutans through conversion of lactic acid into inhibitory H2O2: a possible counteroffensive strategy for interspecies competition. Mol Microbiol 63:872–880
Yamanaka W, Takeshita T, Shibata Y, Matsuo K, Eshima N, Yokoyama T, Yamashita Y (2012) Compositional stability of a salivary bacterial population against supragingival microbiota shift following periodontal therapy. PLoS One 7:e42806
Yang F, Zeng X, Ning K, Liu KL, Lo CC, Wang W, Chen J et al (2012) Saliva microbiomes distinguish caries—active from healthy human populations. ISME J 6:1–10
Ye J, Coulouris G, Zaretskaya I, Cutcutache I, Rozen S, Madden TL (2012) Primer-BLAST: a tool to design target-specific primers for polymerase chain reaction. BMC Bioinformatics 13:134
Yoshida A, Suzuki N, Nakano Y, Kawada M, Oho T, Koga T (2003) Development of a 5' nuclease-based real-time PCR assay for quantitative detection of cariogenic dental pathogens Streptococcus mutans and Streptococcus sobrinus. J Clin Microbiol 41:4438–4441
Zaura E, Brandt BW, Teixeira de Mattos MJ, Buijs MJ, Caspers MP, Rashid MU, Weintraub A et al (2015) Same exposure but two radically different responses to antibiotics: resilience of the salivary microbiome versus long-term microbial shifts in feces. MBio 6
Zhou Y, Gao H, Mihindukulasuriya KA, La Rosa PS, Wylie KM, Vishnivetskaya T, Podar M et al (2013) Biogeography of the ecosystems of the healthy human body. Genome Biol 14:R1
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Funding
This study was supported in part by Grants-in Aid for Scientific Research 25463249 (T. T.), 25293428 (Y. Y.), and 15K15774 (Y. Y.) from the Ministry of Education, Culture, Sports, Science and Technology of Japan.
Ethical approval
All procedures performed in studies involving human participants were in accordance with the ethical standards of the institutional and/or national research committee and with the 1964 Helsinki declaration and its later amendments or comparable ethical standards. The Ethics Committee of Kyushu University approved this study design (reference numbers: 26–130 for the barcoded pyrosequencing analysis and 19B-1 for the quantitative PCR analysis) and the procedure for obtaining informed consent.
Informed consent
Informed consent was obtained from all individual participants included in this study.
Rights and permissions
About this article
Cite this article
Yasunaga, H., Takeshita, T., Shibata, Y. et al. Exploration of bacterial species associated with the salivary microbiome of individuals with a low susceptibility to dental caries. Clin Oral Invest 21, 2399–2406 (2017). https://doi.org/10.1007/s00784-016-2035-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00784-016-2035-5