Abstract
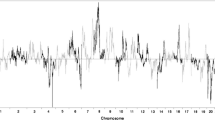
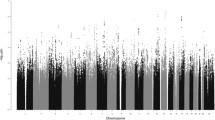
This is the first report of a full genome scan of sexual orientation in men. A sample of 456 individuals from 146 families with two or more gay brothers was genotyped with 403 microsatellite markers at 10-cM intervals. Given that previously reported evidence of maternal loading of transmission of sexual orientation could indicate epigenetic factors acting on autosomal genes, maximum likelihood estimations (mlod) scores were calculated separated for maternal, paternal, and combined transmission. The highest mlod score was 3.45 at a position near D7S798 in 7q36 with approximately equivalent maternal and paternal contributions. The second highest mlod score of 1.96 was located near D8S505 in 8p12, again with equal maternal and paternal contributions. A maternal origin effect was found near marker D10S217 in 10q26, with a mlod score of 1.81 for maternal meioses and no paternal contribution. We did not find linkage to Xq28 in the full sample, but given the previously reported evidence of linkage in this region, we conducted supplemental analyses to clarify these findings. First, we re-analyzed our previously reported data and found a mlod of 6.47. We then re-analyzed our current data, after limiting the sample to those families previously reported, and found a mlod of 1.99. These Xq28 findings are discussed in detail. The results of this first genome screen for normal variation in the behavioral trait of sexual orientation in males should encourage efforts to replicate these findings in new samples with denser linkage maps in the suggested regions.


Similar content being viewed by others
References
Adelman JP, Mason AJ, Hayflick JS, Seeburg PH (1986) Isolation of the gene and hypothalamic cDNA for the common precursor of gonadotropin-releasing hormone and prolactin release-inhibiting factor in human and rat. Proc Natl Acad Sci USA 83:179–183
Bailey JM, Pillard RC (1995) Genetics of human sexual orientation. Annu Rev Sex Res 60:126–150
Bailey JM, Zucker KJ (1995) Childhood sex-typed behavior and sexual orientation: a conceptual analysis and quantitative review. Dev Psych 31:43–55
Bailey JM, Pillard RC, Dawood K, Miller MB, Farrer LA, Trivedi S, Murphy RL (1999) A family history study of male sexual orientation using three independent samples. Behav Genet 29:79–86
Blanchard R (2004) Quantitative and theoretical analyses of the relation between older brothers and homosexuality in men. J Theor Biol 230:173–187
Bocklandt S, Hamer DH (2003) Beyond hormones: a novel hypothesis for the biological basis of male sexual orientation. J Endocrinol Invest 26:8–12
Burden S, Yarden Y (1997) Neuregulins and their receptors: a versatile signaling module in organogenesis and oncogenesis. Neuron 18:847–855
Byne W, Tobet S, Mattiace L, Lasco MS, Kemether E, Edgar MA, Morgello S, Buchsbaum MS, Jones LB (2001) The interstitial nuclei of the human anterior hypothalamus: an investigation of variation within sex, sexual orientation and HIV status. Hormones Behav 40:86–92
Diamond M (1993) Homosexuality and bisexuality in different populations. Arch Sex Behav 22:291–310
Dupree MG, Mustanski BS, Bocklandt S, Nievergelt C, Hamer DH (2004) A candidate gene study of CYP19 (aromatase) and male sexual orientation. Behav Genet 34:243–250
Hamer D (1999) Genetics and male sexual orientation. Science 285:803
Hamer DH, Hu S, Magnuson VL, Hu N, Pattatucci AM (1993) A linkage between DNA markers on the X chromosome and male sexual orientation. Science 261:321–327
Harmar AJ, Marston HM, Shen S, Spratt C, West KM, Sheward WJ, Morrison CF, Dorin JR, Piggins HD, Reubi JC, Kelly JS, Maywood ES, Hastings MH (2002) The VPAC(2) receptor is essential for circadian function in the mouse suprachiasmatic nuclei. Cell 109:497–508
Hinds D, Risch N (1996) The ASPEX package: affected sib-pair exclusion mapping [computer program]. Stanford University, Stanford
Hu S, Pattatucci A, Patterson C, Li L, Fulker D, Cherny S, Kruglyak L, Hamer D (1995) Linkage between sexual orientation and chromosome Xq28 in males but not females. Nat Genet 11:248–256
Kawakami M, Negoro H, Kimura F, Higuchi T, Asai T (1975) Neural control of LH release in anterior periventriculo-median eminence-pituitary system. Neuroendocrinology 19:137–149
Kendler KS, Thornton LM, Gilman SE, Kessler RC (2000) Sexual orientation in a U.S. national sample of twin and nontwin sibling pairs. Am J Psychiatry 157:1843–1846
Kinsey AC, Pomeroy WB, Martin CE (1948) Sexual behavior in the human male. Indiana University Press, Bloomington
Kirk KM, Bailey JM, Dunne MP, Martin NG (2000) Measurement models for sexual orientation in a community twin sample. Behav Genet 30:345–356
Lalumiere ML, Blanchard R, Zucker KJ (2000) Sexual orientation and handedness in men and women: a meta-analysis. Psychol Bull 126:575–592
Lander E, Kruglyak L (1995) Genetic dissection of complex traits: guidelines for interpreting and reporting linkage results. Nat Genet 11:241–247
Laumann EO, Gagnon JH, Michael RT, Michaels S (1994) The social organization of sexuality: sexual practices in the United States. University of Chicago Press, Chicago
LeVay S (1991) A difference in hypothalamic structure between heterosexual and homosexual men. Science 253:1034–1037
Macke JP, Hu N, Hu S, Bailey M, King VL, Brown T, Hamer D, Nathans J (1993) Sequence variation in the androgen receptor gene is not a common determinant of male sexual orientation. Am J Hum Genet 53:844–852
Marshall TC, Slate J, Kruuk L, Pemberton JM (1998) Statistical confidence for likelihood-based paternity inference in natural populations. Mol Ecol 7:639–655
McKnight J, Malcolm J (2000) Is male homosexuality maternally linked? Psych Evol Gender 2:229–239
Metwali A, Elliott D, Blum AM, Li J, Sandor M, Weinstock JV (1996) T cell vasoactive intestinal peptide receptor subtype expression differs between granulomas and spleen of schistosome-infected mice. J Immunol 157:265–270
Mustanski BS, Chivers ML, Bailey JM (2002) A critical review of recent biological research on human sexual orientation. Annu Rev Sex Res 12:89–140
Nievergelt CM, Smith DW, Kohlenberg JB, Schork NJ (2004) Large-scale integration of human genetic and physical maps. Genome Res 14:1199–1205
Rahman Q, Wilson GD (2003) Sexual orientation and the 2nd to 4th finger length ratio: evidence for organising effects of sex hormones or developmental instability? Psychoneuroendocrinology 28:288–303
Rice G, Risch N, Ebers G (1999a) Genetics and male sexual orientation. Science 28:803
Rice G, Anderson C, Risch N, Ebers G (1999b) Male homosexuality: absence of linkage to microsatellite markers at Xq28. Science 284:665–667
Roessler E, Belloni E, Gaudenz K, Jay P, Berta P, Scherer SW, Tsui LC, Muenke M (1996) Mutations in the human Sonic Hedgehog gene cause holoprosencephaly. Nat Genet 14:357–360
Sanders AR, Dawood K (2003) Sexual orientation. In: Nature Encyclopedia of Life Sciences. Nature Publishing Group, London (http://www.els.net/els/public/search/search_public.asp)
Strichman-Almashanu LZ, Lee RS, Onyango PO, Perlman E, Flam F, Frieman MB, Feinberg AP (2002) A genome-wide screen for normally methylated human CpG islands that can identify novel imprinted genes. Genome Res 12:543–554
Sugawara T, Holt JA, Driscoll D, Strauss JF III, Lin D, Miller WL, Patterson D, Clancy KP, Hart IM, Clark BJ, et al (1995) Human steroidogenic acute regulatory protein: functional activity in COS-1 cells, tissue-specific expression, and mapping of the structural gene to 8p11.2 and a pseudogene to chromosome 13. Proc Natl Acad Sci USA 92:4778–4782
Swaab DF, Hofman MA (1990) An enlarged suprachiasmatic nucleus in homosexual men. Brain Res 24:141–148
Tsukui T, Capdevila J, Tamura K, Ruiz-Lozano P, Rodriguez-Esteban C, Yonei-Tamura S, Magallon J, Chandraratna RA, Chien K, Blumberg B, Evans RM, Belmonte JC (1999) Multiple left-right asymmetry defects in Shh(−/−) mutant mice unveil a convergence of the shh and retinoic acid pathways in the control of Lefty-1. Proc Natl Acad Sci USA 96:11376–11381
Wellings K, Field J, Johnson A, Wadsworth J (1994) Sexual behaviour in Britain. Penguin Books, New York
Wen D, Suggs SV, Karunagaran D, Liu N, Cupples RL, Luo Y, Janssen AM, Ben-Baruch N, Trollinger DB, Jacobsen VL, et al (1994) Structural and functional aspects of the multiplicity of Neu differentiation factors. Mol Cell Biol 14:1909–1919
Acknowledgements
We thank all the individuals who participated in the project for their time and openness and Lynn Goldin and Danielle Dick for comments on the manuscript. B.S.M. was supported by a NSF Graduate Research Fellowship and an NIH Summer Research Fellowship. N.J.S. and C.M.N. were supported in part by the NHLBI Family Blood Pressure Program (FBPP; HL64777-01).
Author information
Authors and Affiliations
Corresponding author
Additional information
Brian S. Mustanski and Michael G. DuPree contributed equally to this work.
Rights and permissions
About this article
Cite this article
Mustanski, B.S., DuPree, M.G., Nievergelt, C.M. et al. A genomewide scan of male sexual orientation. Hum Genet 116, 272–278 (2005). https://doi.org/10.1007/s00439-004-1241-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00439-004-1241-4