Abstract
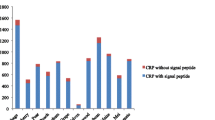
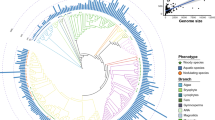
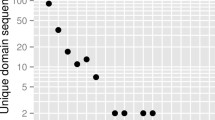
Defensive and reproductive protein genes undergo rapid evolution. Small, cysteine-rich secreted peptides (CRPs) act as antimicrobial agents and function in plant intercellular signaling and are over-represented among reproductively expressed proteins. Because of their roles in defense, reproduction and development and their presence in multigene families, CRP variation can have major consequences for plant phenotypic and functional diversification. We surveyed the CRP genes of six closely related Oryza genomes comprising Oryza sativa ssp. japonica and ssp. indica, Oryza glaberrima and three accessions of Oryza rufipogon to observe patterns of evolution in these gene families and the effects of variation on their gene expression. These Oryza genomes, like other plant genomes, have accumulated large reservoirs of CRP sequences, comprising 26 groups totaling between 676 and 843 genes, in contrast to antimicrobial CRPs in animal genomes. Despite the close evolutionary relationships between the genomes, we observed rapid changes in number and structure among CRP gene families. Many CRP sequences are in gene clusters generated by local duplications, have undergone rapid turnover and are more likely to be silent or specifically expressed. By contrast, conserved CRP genes are more likely to be highly and broadly expressed. Variable CRP genes created by repeated duplication, gene modification and inactivation can gain new functions and expression patterns in newly evolved gene copies. For the CRP proteins, the process of gain/loss by deletion or duplication at gene clusters seems to be an important mechanism in evolution of the gene families, which also contributes to their expression evolution.






Similar content being viewed by others
References
Amien S, Kliwer I, Márton ML, Debener T, Geiger D, Becker D, Dresselhaus T (2010) Defensin-like ZmES4 mediates pollen tube burst in maize via opening of the potassium channel KZM1. PLoS Biol 8:e1000388
Ammiraju JSS, Luo M, Goicoechea JL, Wang W, Kudrna D, Mueller C, Talag J, Kim H, Sisneros NB, Blackmon B, Fang E, Tomkins JB, Brar D, MacKill D, McCouch S, Kurata N, Lambert G, Galbraith DW, Arumuganathan K, Rao K, Walling JG, Gill N, Yu Y, SanMiguel P, Soderlund C, Jackson S, Wing RA (2006) The Oryza bacterial artificial chromosome library resource: construction and analysis of 12 deep-coverage large-insert BAC libraries that represent the 10 genome types of the genus Oryza. Genome Res 16:140–147
Ammiraju JS, Lu F, Sanyal A, Yu Y, Song X, Jiang N, Pontaroli AC, Rambo T, Currie J, Collura K, Talag J, Fan C, Goicoechea JL, Zuccolo A, Chen J, Bennetzen JL, Chen M, Jackson S, Wing RA (2008) Dynamic evolution of Oryza genomes is revealed by comparative genomic analysis of a genus-wide vertical data set. Plant Cell 20:3191–3209
Bedinger PA, Pearce G, Covey PA (2010) RALFs: peptide regulators of plant growth. Plant Signal Behav 5:1342–1346
Boland M (2013) Kiwifruit proteins and enzymes: actinidin and other significant proteins. Adv Food Nutr Res 68:59–80
Boutrot F, Chantret N, Gautier M-F (2008) Genome-wide analysis of the rice and arabidopsis non-specific lipid transfer protein (nsLtp) gene families and identification of wheat nsLtp genes by EST data mining. BMC Genom 9:86
Chae K, Zhang K, Zhang L, Morikis D, Kim ST, Mollet J-C, de la Rosa N, Tan K, Lord EM (2007) Two SCA (stigma/style cysteine-rich Adhesin) isoforms show structural differences that correlate with their levels of in vitro pollen tube adhesion activity. J Biol Chem 282:33845–33858
Cosgrove DJ (2000) Loosening of plant cell walls by expansins. Nature 407:321–326
Costa LM, Marshall E, Tesfaye M, Silverstein KAT, Mori M, Umetsu Y, Otterback SL, Papareddy R, Dickinson HG, Boutiller K, VandenBosch KA, Ohki S, Gutierrez-Marcos JS (2014) Central cell-derived peptides regulate early embryo patterning in flowering plants. Science 344:168–172
Davidson RM, Gowda M, Moghe G, Lin H, Vaillancourt B, Shiu S-H, Jiang N, Buell RC (2012) Comparative transcriptomics of three Poaceae species reveals patterns of gene expression evolution. Plant J. 71:492–502
de Carvalho A, Gomes VM (2011) Plant defensins and defensin-like peptides—biological activities and biotechnological applications. Curr Pharm Des 17:4270–4293
De Graaf BHJ, Knuiman BA, Derksen J, Mariani C (2003) Characterization and localization of the transmitting tissue-specific PELPIII proteins of Nicotiana tabacum. J Exp Bot 54:55–63
Dean RA, Talbot NJ, Ebbole DJ, Farman ML, Mitchell TK, Orbach MJ, Thon M, Kulkarni R, Xu J-R, Pan H, Read NR, Lee Y-H, Carbone I, Brown D, Oh YY, Donofrio N, Jeong JS, Soanes DM, Djonovic S, Kolomlets E, Rehmeyer C, Li W, Harding M, Kim S, Lebrun M-H, Bohnert H, Coughlan S, Butler J, Calvo S, Ma L-J, Nicol R, Purcell S, Nusbaum C, Galagan JE, Birren BW (2005) The genome sequence of the rice blast fungus Magnaporthe grisea. Nature 434:980–986
Eddy SR (1998) Profile hidden Markov models. Bioinformatics 14:755–763
Eddy SR (2011) Accelerated profile HMMsearches. PLoS Comput Biol 7:e1002195
Edstam MM, Viitanen L, Salminen TA, Edqvist J (2011) Evolutionary history of the non-specific lipid transfer proteins. Mol Plant 4:947–964
Finn RD, Mistry J, Coggill P, Heger A, Pollington JE, Gavin OL, Gunasekaran P, Ceric G, Forslund K, Holm L, Sonnhammer EL, Eddy SR, Bateman A (2009) The Pfam protein families database. Nucleic Acids Res 38:D211–D222
Gan X, Stegle O, Behr J, Steffen JG, Drewe P, Hildebrand KL, Lyngsoe R, Schultheiss SJ, Osborne EJ, Sreedharan VT, Kahles A, Bohnert R, Jean G, Derwent P, Kersey P, Belfield EJ, Harberd NP, Kemen E, Toomajian C, Kover PX, Clark RM, Rätsch G, Mott R (2011) Multiple reference genomes and transcriptomes for Arabidopsis thaliana. Nature 477:419–423
Gibbs GM, Roelants K, O’Bryan MK (2008) The CAP superfamily: cysteine-rich secretory proteins, antigen 5, and pathogenesis-related 1 proteins—roles in reproduction, cancer, and immune defense. Endocr Rev 29:865–897
Gutiérrez-Marcos JF, Costa LM, Biderre-Petit C, Khbaya B, O’Sullivan DM, Wormald M, Perez P, Dickinson HG (2004) Maternally expressed gene1 is a novel maize endosperm transfer cell-specific gene with a maternal parent-of-origin pattern of expression. Plant Cell 16:1288–1301
Hara K, Yokoo T, Kajita R, Onishi T, Yahata S, Peterson KM, Torii KU, Kakimoto T (2009) Epidermal cell density is autoregulated via a secretory peptide, EPIDERMAL PATTERNING FACTOR 2 in Arabidopsis leaves. Plant Cell Physiol 50:1019–1031
Jacquemin J, Ammiraju JSS, Haberer G, Billheimer DD, Yu Y, Liu LC, Rivera LF, Mayer K, Chen M, Wing RA (2013) 15 MYA of evolution in the Oryza genus shows extensive gene family expansion. Mol Plant 7:642–656
José-Estanyol M, Gomis-Rüth FX, Puigdomènech P (2004) The eight-cysteine motif, a versatile structure in plant proteins. Plant Physiol Biochem 42:355–365
Kasprzewska A (2003) Plant chitinases—regulation and function. Cell Mol Biol Lett 8:809–824
Kawahara Y, Oono Y, Kanamori H, Matsumoto T, Itoh T, Minami E (2012) Simultaneous RNA-seq analysis of a mixed transcriptome of rice and blast fungus interaction. PLoS ONE 7:e49423
Kim D, Pertea G, Trapnell C, Pimentel H, Kelley R, Salzberg SL (2013) TopHat2: accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol 14:R36
Langmead B, Trapnell C, Pop M, Salzberg SL (2009) Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol 10:R25
Larkin MA, Blackshields G, Brown NP, Chenna R, McGettigan PA, McWilliam H, Valentin F, Wallace IM, Wilm A, Lopez R, Thompson JD, Gibson TJ, Higgins DG (2007) Clustal W and Clustal X version 2.0. Bioinformatics 23:2947–2948
Li H, Durbin R (2009) Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25:1754–1760
Light S, Sagit R, Sachenkova O, Ekman D, Elofsson A (2013) Protein expansion is primarily due to indels in intrinsically disordered regions. Mol Biol Evol 30:2645–2653
Marshall E, Costa LM, Gutierrez-Marcos J (2011) Cysteine-rich peptides (CRPs) mediate diverse aspects of cell-cell communication in plant reproduction and development. J Exp Bot 62:1677–1686
Matsuyama T, Yasumura N, Funakoshi M, Yamada Y, Hashimoto T (1999) Maize genes specifically expressed in the outermost cells of root cap. Plant Cell Physiol 40:469–476
Mergaert P, Nikovics K, Kelemen Z, Maunoury N, Vaubert D, Kondorosi A, Kondorosi E (2003) A novel family in Medicago truncatula consisting of more than 300 nodule-specific genes coding for small, secreted polypeptides with conserved cysteine motifs. Plant Physiol 132:161–173
Mizuno H, Kawahara Y, Sakai H, Kanamori H, Wakimoto H, Yamagata H, Oono Y, Wu J, Ikawa H, Itoh T, Matsumoto T (2010) Massive parallel sequencing of mRNA in identification of unannotated salinity stress-inducible transcripts in rice (Oryza sativa L.). BMC Genom 11:683
Naito K, Zhang F, Tsukiyama T, Saito H, Hancock N, Richardson AO, Okumoto Y, Tanisaka T, Wessler SR (2009) Unexpected consequences of a sudden and massive transposon amplification on rice gene expression. Nature 461:1130–1134
Nallu S, Silverstein KAT, Samac DA, Bucciarelli B, Vance CP, Vanden Bosch KA (2013) Regulatory patterns of a large family of defensin-like genes expressed in nodules of Medicago truncatula. PLoS ONE 8:e60355
Okuda S, Tsutsui H, Shiina K, Sprunck S, Takeuchi H, Yui R, Kasahara RD, Hamamura Y, Mizukami A, Susaki D, Kawano N, Sakakibara T, Namiki S, Itoh K, Otsuka K, Matsuzaki M, Nozaki H, Kuroiwa T, Nakano A, Kanaoka MM, Dresselhaus T, Sasaki N, Higashiyama T (2009) Defensin-like polypeptide LUREs are pollen tube attractants secreted from synergid cells. Nature 458:357–361
Rice P, Longden I, Bleasby A (2000) EMBOSS: the European molecular biology open software suite. Trends Genet 16:276–277
Schmidt A, Teeter M, Weckert E, Lamzin VS (2011) Crystal structure of small protein crambin at 0.48 Å resolution. Acta Crystallograph Sect F Struct Biol Cryst Commun 67: 424
Schopfer CR, Nasrallah ME, Nasrallah JB (1999) The male determinant of self-incompatibility in Brassica. Science 286:1697–1700
Silverstein KAT, Moskal WA Jr, Wu HC, Underwood BA, Graham MA, Town CD, Vanden Bosch KA (2007) Small cysteine-rich peptides resembling antimicrobial peptides have been under-predicted in plants. Plant J 51:262–280
Stec B (2006) Plant thionins—the structural perspective. CMLS Cell Mol Life S 63:1370–1385
Svensson B, Svendsen I, Højrup P, Roepstorff P, Ludvigsen S, Poulsen FM (1992) Primary structure of barwin: a barley seed protein closely related to the C-terminal domain of proteins encoded by wound-induced plant genes. Biochem Mosc 31:8767–8770
Takeuchi H, Higashiyama T (2012) A species-specific cluster of defensin-like genes encodes diffusible pollen tube attractants in Arabidopsis. PLoS Biol 10:e1001449
Tang L, Zou X, Achoundong G, Potgieter C, Second G, Zhang D, Ge S (2010) Phylogeny and biogeography of the rice tribe (Oryzeae): evidence from combined analysis of 20 chloroplast fragments. Mol Phylogenet Evol 54:266–277
Trapnell C, Roberts A, Goff L, Pertea G, Kim D, Kelley DR, Pimentel H, Salzberg SL, Rinn JL, Pachter L (2012) Differential gene and transcript expression analysis of RNA-seq experiments with TopHat and Cufflinks. Nat Protoc 7:562–578
Tsukamoto T, Qin Y, Huang Y, Dunatunga D, Palanivelu R (2010) A role for LORELEI, a putative glycosylphosphatidylinositol-anchored protein, in Arabidopsis thaliana double fertilization and early seed development. Plant J 62:571–588
Wang M, Yu Y, Haberer G, Marri PR, Fan C, Goicoechea JL, Zuccolo A, Song X, Kudrna D, Ammiraju JSS, Cossu RM, Maldonado C, Chen J, Lee S, Sisneros N, de Baynast K, Golser W, Wissotski M, Kim W, Sanchez P, Ndjiondjop M-N, Sanni K, Long M, Carney J, Panaud O, Wicker T, Machado CA, Chen M, Mayer KFX, Rounsley S, Wing RA (2014) The genome sequence of African rice (Oryza glaberrima) and evidence for independent domestication. Nat Genet 46:982–988
Wheeler MJ, de Graaf BHJ, Hadjiosif N, Perry RM, Poulter NS, Osman K, Vatovec S, Harper A, Franklin FCH, Franklin-Tong VE (2009) Identification of the pollen self-incompatibility determinant in Papaver rhoeas. Nature 459:992–995
Xu H, Swoboda I, Bhalla PL, Singh MB (1999) Male gametic cell-specific gene expression in flowering plants. Proc Natl Acad Sci 96:2554–2558
Yang Z (2007) PAML 4: phylogenetic analysis by maximum likelihood. Mol Biol Evol 24:1586–1591
Zimmermann R, Sakai H, Hochholdinger F (2010) The gibberellic acid stimulated-like gene family in maize and its role in lateral root development. Plant Physiol 152:356–365
Acknowledgements
This work was supported partly from a grant by the Transdisciplinary Research Integration Center (TRIC) of the Research Organization of Information and Systems, and partly from the National BioResource Project, Japan (NBRP). We would like to thank Drs. Masahiro Fujita (NIG) and Qiang Zhao (CAS) for their help to register genome sequences. This work was funded by a grant from the Transdisciplinary Research Integration Center (TRIC) of the Research Organization of Information and Systems for the project “Genetic Function Systems”, and by the National BioResource Project Japan (NBRP) for the project “Rice Bioresources”.
Conflict of interest
The authors declare that they have no conflict of interest.
Research involving human participants and/or animals
This article does not contain any studies with human participants or animals performed by any of the authors.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by B. Yang.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Shenton, M.R., Ohyanagi, H., Wang, ZX. et al. Rapid turnover of antimicrobial-type cysteine-rich protein genes in closely related Oryza genomes. Mol Genet Genomics 290, 1753–1770 (2015). https://doi.org/10.1007/s00438-015-1028-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00438-015-1028-4