Abstract
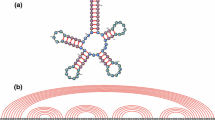
RNA secondary structure formation is a field of considerable biological interest as well as a model system for understanding generic properties of heteropolymer folding. This system is particularly attractive because the partition function and thus all thermodynamic properties of RNA secondary structure ensembles can be calculated numerically in polynomial time for arbitrary sequences and homopolymer models admit analytical solutions. Such solutions for many different aspects of the combinatorics of RNA secondary structure formation share the property that the final solution depends on differences of statistical weights rather than on the weights alone. Here, we present a unified approach to a large class of problems in the field of RNA secondary structure formation. We prove a generic theorem for the calculation of RNA folding partition functions. Then, we show that this approach can be applied to the study of the molten-native transition, denaturation of RNA molecules, as well as to studies of the glass phase of random RNA sequences.


Similar content being viewed by others
References
Alberts B, Johnson A, Lewis J, Raff M, Roberts K, Walter P (2002) Molecular biology of the cell. Garland Science, New York
Bundschuh R, Hwa T (1999) RNA structure formation: a solvable model of heteropolymer folding. Phys Rev Lett 83:1479–1482
Bundschuh R, Hwa T (2002a) Phases of the secondary structure of RNA sequences. Europhys Lett 59: 903–909
Bundschuh R, Hwa T (2002b) Statistical mechanics of secondary structures formed by random RNA sequences. Phys Rev E 65:031903
Bundschuh R, Gerland U (2006) Dynamics of intramolecular recognition: base-pairing in DNA/RNA near and far from equilibrium. Eur Phys J E 19:319–329
David F, Wiese KJ (2007) Systematic field theory of the RNA glass transition. Phys Rev Lett 98:128102
Ding Y, Chan CY, Lawrence CE (2004) Sfold web server for statistical folding and rational design of nucleic acids. Nucleic Acids Res 32:W135–W141
de Gennes PG (1968) Statistics of branching and hairpin helices for the dAT copolymer. Biopolymers 6:715–729
El Attar R (2006) Lecture notes on \(Z\)-transform. Lulu Press Inc., Morrisville
Gō N (1983) Theoretical studies of protein folding. Annu Rev Biophys Bioeng 12:183–210
Jury EI (1973) Theory and application of the \(Z\)-transform method. Robert E. Krieger Publishing Co., Huntington
Hartmann AK (2001) Comment on “Glassy transition in a disordered model for the RNA secondary structure”. Phys Rev Lett 86:1382
Higgs PG (1996) Overlaps between RNA secondary structures. Phys Rev Lett 76:704–707
Higgs PG (2000) RNA secondary structure: physical and computational aspects. Q Rev Biophys 33:199–253
Hofacker IL, Fontana W, Stadler PF, Bonhoeffer S, Tacker M, Schuster P (1994) Fast folding and comparison of RNA secondary structures. Monatshefte f Chemie 125:167–188
Hui S, Tang LH (2006) Ground state and glass transition of the RNA secondary structure. Eur Phys J B 53:77–84
Lässig M, Wiese KJ (2006) Freezing of random RNA. Phys Rev Lett 96:228101
Liu T, Bundschuh R (2005) Quantification of the differences between quenched and annealed averaging for RNA secondary structures. Phys Rev E 72:061905
Marinari E, Pagnani A, Ricci-Tersenghi F (2002) Zero-temperature properties of RNA secondary structures. Phys Rev E 65:041919
Mathews DH, Sabina J, Zuker M, Turner DH (1999) Expanded sequence dependence of thermodynamic parameters improves prediction of RNA secondary structure. J Mol Biol 288:911–940
McCaskill JS (1990) The equilibrium partition function and base pair binding probabilities for RNA secondary structure. Biopolymers 29:1105–1119
Monthus C, Garel T (2007) Freezing transition of the random bond RNA model: statistical properties of the pairing weights. Phys Rev E 75:031103
Moroz D, Hwa T (1999) Toy model of heteropolymer interactions and denaturation. In: Bulletin of the 1999 March Meeting of the American Physical Society, SC31.14
Nebel ME (2003) Combinatorial properties of RNA secondary structures. J Comput Biol 3:541–574
Pagnani A, Parisi G, Ricci-Tersenghi F (2000) Glassy transition in a disordered model for the RNA secondary structure. Phys Rev Lett 84:2026–2029
Pagnani A, Parisi G, Ricci-Tersenghi F (2001) Pagnani, Parisi, and Ricci-Tersenghi Reply. Phys Rev Lett 86:1383
Tinoco I Jr, Bustamante C (1999) How RNA folds. J Mol Biol 293:271–281
Waterman MS (1978) Secondary structure of single-stranded nucleic acids. In: Studies in foundations and combinatorics. Advances in Mathematics Supplementary Studies, vol 1. Academic Press, New York, pp 167–212
Xayaphoummine A, Bucher T, Isambert H (2005) Kinefold web server for RNA/DNA folding path and structure prediction including pseudoknots and knots. Nucleic Acids Res 33:605–610
Zuker M, Stiegler P (1981) Optimal computer folding of large RNA sequences using thermodynamics and auxiliary information. Nucleic Acids Res 9:133–148
Zuker M (2003) Mfold web server for nucleic acid folding and hybridization prediction. Nucleic Acids Res 31:3406–3415
Acknowledgments
I would like to acknowledge countless interactions on which this work is based with Terry Hwa. In addition, I would like to acknowledge discussions and collaborations with Robijn Bruinsma, Tsunglin Liu, David Moroz, Lei-Han Tang, and Michael Zuker that contributed to this work. I would like to thank the Aspen Center for Theoretical Physics where the idea for the theorem presented here was conceived. Lastly, I would like to acknowledge support by the US National Science Foundation under DMR Grants 0706002 and 1105458.
Author information
Authors and Affiliations
Corresponding author
Appendix A: Molten partition function for even length
Appendix A: Molten partition function for even length
In this appendix we calculate the \(z\)-transform of the molten phase transition function for even sequence length only. For clarity we suppress the argument \(q\) in all partition functions and understand that all partition functions depend on the Boltzmann factor \(q\).
We will actually calculate the \(z\)-transforms for even and odd sequence lengths simultaneously and start by defining
for \(N\ge 1\) (note, that odd arguments of \(G\) correspond to chains of an even length and vice versa). Here, \(G(N)\) is the molten phase partition function introduced in Sect. 2.4. These quantities have \(z\)-transforms
By their construction we have the relation
Let us additionally insert \(N=2n-1\) for \(n\ge 2\) into Eq. (5). This gives us
Multiplying this equation by \(z^{-n}\), summing over all \(n\ge 2\) and adding \(z^{-1}\) on both sides yields
which in turn can be rewritten as
If we insert this into Eq. (23) we get the equation
for \(\widehat{E}(z)\). This is a quadratic equation for \(\widehat{E}(z)\). Inserting the explicit expression (8) for \(\widehat{G}(z)\) this quadratic equation can be solved explicitly with the result
The other solution with a minus sign between the two square roots has to be excluded, since \(\widehat{O}(z)>0\) and \(\widehat{E}(z)>0\) for real and positive \(z\) and thus, according to Eq. (24), \(\widehat{E}(z)<1/(2q)\).
Rights and permissions
About this article
Cite this article
Bundschuh, R. Unified approach to partition functions of RNA secondary structures. J. Math. Biol. 69, 1129–1150 (2014). https://doi.org/10.1007/s00285-013-0737-8
Received:
Revised:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00285-013-0737-8