Abstract
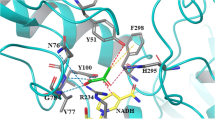
3-Phenyllactic acid (PLA) is an antimicrobial compound with broad-spectrum activity against bacteria and fungi that could be widely used in the food industry and livestock feeds. Notably, d-PLA exhibits higher antibacterial activity, which gains more attention than l-PLA. In this report, the d-lactate dehydrogenase DLDH744 from Sporolactobacillus inulinus CASD was engineered to increase the enzymatic activities toward phenylpyruvate by protein structure-guided modeling analysis. The phenylpyruvate molecule was first docked in the active center of DLDH744. The residues that might tightly pack around the benzene ring of phenylpyruvate were all selected for mutation. The single site mutant M307L showed the highest increased activity toward bulkier substrate phenylpyruvate than the wild type. By using the engineered d-lactate dehydrogenase M307L expressed in Escherichia coli strains, without coexpression of the cofactor regeneration system, 21.43 g/L d-PLA was produced from phenylpyruvate with a productivity of 1.58 g/L/h in the fed-batch biotransformation process, which ranked in the list as the highest production titer of d-PLA by d-lactate dehydrogenase. The enantiomeric excess value of produced d-PLA in the broth was higher than 99.7 %. Additionally, the structure-guided design of this enzyme will also provide referential information for further engineering other 2-hydroxyacid dehydrogenases, which are useful for a wide range of fine chemical synthesis.







Similar content being viewed by others
References
Antonyuk SV, Strange RW, Ellis MJ, Bessho Y, Kuramitsu S, Inoue Y, Yokoyama S, Hasnain SS (2009) Structure of D-lactate dehydrogenase from Aquifex aeolicus complexed with NAD+ and lactic acid (or pyruvate). Acta Crystallogr Sect F: Struct Biol Cryst Commun 65:1209–1213
Bradford MM (1976) A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem 72:248–254
Dieuleveux V, van der Pyl D, Chataud J, Gueguen M (1998a) Purification and characterization of anti-Listeria compounds produced by Geotrichum candidum. Appl Environ Microbiol 64:800–803
Dieuleveux V, Lemarinier S, Gueguen M (1998b) Antimicrobial spectrum and target site of D-3-phenyllactic acid. Int J Food Microbiol 40:177–183
Fujii T, Shimizu M, Doi Y, Fujita T, Ito T, Miura D, Wariishi H, Takaya N (2011) Novel fungal phenylpyruvate reductase belongs to d-isomer-specific 2-hydroxyacid dehydrogenase family. Biochim Biophys Acta 1814:1669–1676
Fujita T, Nguyen HD, Ito T, Zhou S, Osada L, Tateyama S, Kaneko T, Takaya N (2013) Microbial monomers custom-synthesized to build true bio-derived aromatic polymers. Appl Microbiol Biotechnol 97:8887–8894
Ishikura Y, Tsuzuki S, Takahashi O, Tokuda C, Nakanishi R, Shinoda T, Taguchi H (2005) Recognition site for the side chain of 2-ketoacid substrate in D-lactate dehydrogenase. J Biochem 138:741–749
Lavermicocca P, Valerio F, Evidente A, Lazzaroni S, Corsetti A, Gobbetti M (2000) Purification and characterization of novel antifungal compounds from the sourdough Lactobacillus plantarum strain 21B. Appl Environ Microbiol 66:4084–4090
Lavermicocca P, Valerio F, Visconti A (2003) Antifungal activity of phenyllactic acid against molds isolated from bakery products. Appl Environ Microbiol 69:634–640
Li X, Jiang B, Pan B (2007) Biotransformation of phenylpyruvic acid to phenyllactic acid by growing and resting cells of a Lactobacillus sp. Biotechnol Lett 29:593–597
Li L, Shin SY, Lee K, Han N (2014) Production of natural antimicrobial compound D-phenyllactic acid using Leuconostoc mesenteroides ATCC 8293 whole cells involving highly active D-lactate dehydrogenase. Lett Appl Microbiol 59:404–411
Morris GM, Goodsell DS, Halliday RS, Huey R, Hart WE, Belew RK, Olson AJ (1998) Automated docking using a Lamarckian genetic algorithm and an empirical binding free energy function. J Comput Chem 19:1639–1662
Mu W, Chen C, Li X, Zhang T, Jiang B (2009) Optimization of culture medium for the production of phenyllactic acid by Lactobacillus sp. SK007. Bioresour Technol 100:1366–1370
Mu W, Yu S, Zhu L, Zhang T, Jiang B (2012) Recent research on 3-phenyllactic acid, a broad-spectrum antimicrobial compound. Appl Microbiol Biotechnol 95:1155–1163
Ndagano D, Lamoureux T, Dortu C, Vandermoten S, Thonart P (2011) Antifungal activity of lactic acid bacteria of the Weissella genus isolated from food. J Food Sci 76:M305–311
Ohhira I, Kuwaki S, Morita H, Suzuki T, Tomita S, Hisamatsu S, Sonoki S, Shinoda S (2004) Identification of 3-phenyllactic acid as a possible antibacterial substance produced by Enterococcus faecalis TH10. Biocontrol Sci 9:77–81
Razeto A, Kochhar S, Hottinger H, Dauter M, Wilson KS, Lamzin VS (2002) Domain closure, substrate specificity and catalysis of D-lactate dehydrogenase from Lactobacillus bulgaricus. J Mol Biol 318:109–119
Schwenninger SM, Lacroix C, Truttmann S, Jans C, Spörndli C, Bigler L, Meile L (2008) Characterization of low-molecular-weight antiyeast metabolites produced by a food-protective Lactobacillus-Propionibacterium coculture. J Food Protect 71:2481–2487
Tokuda C, Ishikura Y, Shigematsu M, Mutoh H, Tsuzuki S, Nakahira Y, Tamura Y, Shinoda T, Arai K, Takahashi O, Taguchi H (2003) Conversion of Lactobacillus pentosus D-lactate dehydrogenase to a D-hydroxyisocaproate dehydrogenase through a single amino acid replacement. J Bacteriol 185:5023–5026
Urban A, Neukirchen S, Jaeger KE (1997) A rapid and efficient method for site-directed mutagenesis using one-step overlap extension PCR. Nucleic Acids Res 25:2227–2228
Valerio F, Lavermicocca P, Pascale M, Visconti A (2004) Production of phenyllactic acid by lactic acid bacteria: an approach to the selection of strains contributing to food quality and preservation. FEMS Microbiol Lett 233:289–295
Wang LM, Cai YM, Zhu LF, Guo HL, Yu B (2014) Major role of NAD-dependent lactate dehydrogenases in high optically pure L-lactic acid production by thermophilic Bacillus coagulans. Appl Environ Microbiol 80:7134–7141
Xu GC, Zhang LL, Ni Y (2015) Enzymatic preparation of D-phenyllactic acid at high space-time yield with a novel phenylpyruvate reductase identified from Lactobacillus sp. CGMCC 9967. J Biotechnol S0168–1656:30210–30218
Yu B, Su F, Wang L, Xu K, Zhao B, Xu P (2011) Draft genome sequence of Sporolactobacillus inulinus strain CASD, an efficient D-lactic acid-producing bacterium with high-concentration lactate tolerance capability. J Bacteriol 193:5864–5865
Yu S, Zhu L, Zhou C, An T, Jiang B, Mu W (2014) Enzymatic production of D-3-phenyllactic acid by Pediococcus pentosaceus D-lactate dehydrogenase with NADH regeneration by Ogataea parapolymorpha formate dehydrogenase. Biotechnol Lett 36:627–631
Yu S, Zhou C, Zhang T, Jiang B, Mu W (2015) Short communication: 3-phenyllactic acid production in milk by Pediococcus pentosaceus SK25 during laboratory fermentation process. J Dairy Sci 98:813–817
Zheng Z, Ma C, Gao C, Li F, Qin J, Zhang H, Wang K, Xu P (2011) Efficient conversion of phenylpyruvic acid to phenyllactic acid by using whole cells of Bacillus coagulans SDM. PLoS One 6:e19030
Zheng Z, Sheng B, Gao C, Zhang H, Qin T, Ma C, Xu P (2013) Highly stereoselective biosynthesis of (R)-α-hydroxy carboxylic acids through rationally re-designed mutation of D-lactate dehydrogenase. Sci Rep 3:3401
Zhu LF, Xu XL, Wang LM, Dong H, Yu B, Ma YH (2015a) NADP+-preferring D-lactate dehydrogenase from Sporolactobacillus inulinus. Appl Environ Microbiol 81:6294–6301
Zhu LF, Xu XL, Wang LM, Dong H, Yu B (2015b) The D-lactate dehydrogenase from Sporolactobacillus inulinus also possessing reversible deamination activity. PLoS One 10:e0139066
Zhu Y, Hu F, Zhu Y, Wang L, Qi B (2015c) Enhancement of phenyllactic acid biosynthesis by recognition site replacement of D-lactate dehydrogenase from Lactobacillus pentosus. Biotechnol Lett 37:1233–1241
Acknowledgments
The work was supported by grants from the National Natural Science Foundation of China (31270108). BY is supported by the Youth Innovation Promotion Association, Chinese Academy of Sciences.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no competing interests.
Ethical statement
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Min Wang and Lingfeng Zhu contributed equally to this work.
Rights and permissions
About this article
Cite this article
Wang, M., Zhu, L., Xu, X. et al. Efficient production of enantiomerically pure d-phenyllactate from phenylpyruvate by structure-guided design of an engineered d-lactate dehydrogenase. Appl Microbiol Biotechnol 100, 7471–7478 (2016). https://doi.org/10.1007/s00253-016-7456-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-016-7456-1