Abstract
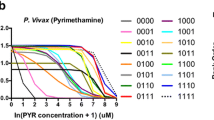
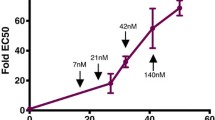
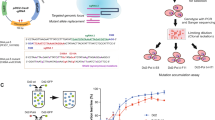
Antifolate antimalarials, such as pyrimethamine, have experienced a dramatic reduction in therapeutic efficacy as resistance has evolved in multiple malaria species. We present evidence from one such species, Plasmodium vivax, which has experienced sustained selection for pyrimethamine resistance at the dihydrofolate reductase (DHFR) locus since the 1970s. Using a transgenic Saccharomyces cerevisiae model expressing the P. vivax DHFR enzyme, we assayed growth rate and resistance of all 16 combinations of four DHFR amino acid substitutions. These substitutions were selected based on their known association with drug resistance, both in natural isolates and in laboratory settings, in the related malaria species P. falciparum. We observed a strong correlation between the resistance phenotypes for these 16 P. vivax alleles and previously observed resistance data for P. falciparum, which was surprising since nucleotide diversity levels and common polymorphic variants of DHFR differ between the two species. Similar results were observed when we expressed the P. vivax alleles in a transgenic bacterial system. This suggests common constraints on enzyme evolution in the orthologous DHFR proteins. The interplay of negative trade-offs between the evolution of novel resistance and compromised endogenous function varies at different drug dosages, and so too do the major trajectories for DHFR evolution. In simulations, it is only at very high drug dosages that the most resistant quadruple mutant DHFR allele is favored by selection. This is in agreement with common polymorphic DHFR data in P. vivax, from which this quadruple mutant is missing. We propose that clinical dosages of pyrimethamine may have historically been too low to select for the most resistant allele, or that the fitness cost of the most resistant allele was untenable without a compensatory mutation elsewhere in the genome.




Similar content being viewed by others
References
Aharoni A, Gaidukov L, Khersonsky O et al (2005) The ‘evolvability’ of promiscuous protein functions. Nat Genet 37:73–76
Ahmed A, Das MK, Dev V et al (2006) Quadruple mutations in dihydrofolate reductase of Plasmodium falciparum isolates from Car Nicobar Island, India. Antimicrob Agents Chemother 50:1546–1549
Anderson TJC, Nair S, Sudimack D et al (2005) Geographical distribution of selected and putatively neutral SNPs in Southeast Asian malaria parasites. Mol Biol Evol 22:2362–2374
Ayala FJ, Escalante AA, Rich SM (1999) Evolution of Plasmodium and the recent origin of the world populations of Plasmodium falciparum. Parasitologia 41:55–68
Berkhout B (1999) HIV-1 evolution under pressure of protease inhibitors: climbing the stairs of viral fitness. J Biomed Sci 6:298–305
Breman JG (2012) Resistance to artemisinin-based combination therapy. Lancet Infect Dis 12:820–822
Brown KM, Costanzo MS, Xu W et al (2010) Compensatory mutations restore fitness during the evolution of dihydrofolate reductase. Mol Biol Evol 27:2682–2690
Carter R (2003) Speculations on the origins of Plasmodium vivax malaria. Trends Parasitol 19:214–219
Chang H-H, Park DJ, Galinsky KJ et al (2012) Genomic sequencing of Plasmodium falciparum malaria parasites from senegal reveals the demographic history of the population. Mol Biol Evol 29:3427–3439
Choowongkomon K, Theppabutr S, Songtawee N et al (2010) Computational analysis of binding between malarial dihydrofolate reductases and anti-folates. Malar J 9:65
Chou H-H, Chiu H-C, Delaney NF et al (2011) Diminishing returns epistasis among beneficial mutations decelerates adaptation. Science 332:1190–1192
Cortese JF, Plowe CV (1998) Antifolate resistance due to new and known Plasmodium falciparum dihydrofolate reductase mutations expressed in yeast. Mol Biochem Parasitol 94:205–214
Cowman AF, Morry MJ, Biggs BA et al (1988) Amino acid changes linked to pyrimethamine resistance in the dihydrofolate reductase–thymidylate synthase gene of Plasmodium falciparum. Proc Natl Acad Sci USA 85:9109–9113
de Pécoulas PE, Tahar R, Ouatas T et al (1998) Sequence variations in the Plasmodium vivax dihydrofolate reductase–thymidylate synthase gene and their relationship with pyrimethamine resistance. Mol Biochem Parasitol 92:265–273
Depristo MA, Zilversmit MM, Hartl DL (2006) On the abundance, amino acid composition, and evolutionary dynamics of low-complexity regions in proteins. Gene 378:19–30
Gregson A, Plowe CV (2005) Mechanisms of resistance of malaria parasites to antifolates. Pharmacol Rev 57:117–145
Hastings MD, Porter KM, Maguire JD et al (2004) Dihydrofolate reductase mutations in Plasmodium vivax from Indonesia and therapeutic response to sulfadoxine plus pyrimethamine. J Infect Dis 189:744–750
Hawkins VN, Joshi H, Rungsihirunrat K et al (2007) Antifolates can have a role in the treatment of Plasmodium vivax. Trends Parasitol 23:213–222
Hawkins VN, Auliff A, Prajapati SK et al (2008) Multiple origins of resistance-conferring mutations in Plasmodium vivax dihydrofolate reductase. Malar J 7:72
Howell EE, Foster PG, Foster LM (1988) Construction of a dihydrofolate reductase-deficient mutant of Escherichia coli by gene replacement. J Bacteriol 170:3040–3045
Hyde JE (2005) Drug-resistant malaria. Trends Parasitol 21:494–498
Joseph SB, Hall DW (2004) Spontaneous mutations in diploid Saccharomyces cerevisiae: more beneficial than expected. Genetics 168:1817–1825
Kepler TB, Perelson AS (1998) Drug concentration heterogeneity facilitates the evolution of drug resistance. Proc Natl Acad Sci USA 95:11514–11519
Khim N, Kim S, Bouchier C et al (2012) Reduced impact of pyrimethamine drug pressure on Plasmodium malariae dihydrofolate reductase gene. Antimicrob Agents Chemother 56:863–868
Kompis IM, Islam K, Then RL (2005) DNA and RNA synthesis: antifolates. Chem Rev 105:593–620
Kondrashov FA (2005) In search of the limits of evolution. Nat Genet 37:9–10
Kongsaeree P, Khongsuk P, Leartsakulpanich U et al (2005) Crystal structure of dihydrofolate reductase from Plasmodium vivax: pyrimethamine displacement linked with mutation-induced resistance. Proc Natl Acad Sci USA 102:13046–13051
Leartsakulpanich U, Imwong M, Pukrittayakamee S et al (2002) Molecular characterization of dihydrofolate reductase in relation to antifolate resistance in Plasmodium vivax. Mol Biochem Parasitol 119:63–73
Looareesuwan S, Viravan C, Webster HK et al (1996) Clinical studies of atovaquone, alone or in combination with other antimalarial drugs, for treatment of acute uncomplicated malaria in Thailand. Am J Trop Med Hyg 54:62–66
Lozovsky ER, Chookajorn T, Brown KM et al (2009) Stepwise acquisition of pyrimethamine resistance in the malaria parasite. Proc Natl Acad Sci USA 106:12025–12030
Mendis K, Sina BJ, Marchesini P, Carter R (2001) The neglected burden of Plasmodium vivax malaria. Am J Trop Med Hyg 64:97–106
Nair S, Miller B, Barends M et al (2008) Adaptive copy number evolution in malaria parasites. PLoS Genet 4:e1000243
Neafsey DE, Galinsky K, Jiang RHY et al (2012) The malaria parasite Plasmodium vivax exhibits greater genetic diversity than Plasmodium falciparum. Nat Genet 44:1046–1050
Parekh F, Moorthy V (2011) Plasmodium vivax vaccine research: insights from Colombian studies. Am J Trop Med Hyg 84:1
Peng B, Kimmel M (2005) simuPOP: a forward-time population genetics simulation environment. Bioinformatics 21:3686–3687
Peterson DS, Walliker D, Wellems TE (1988) Evidence that a point mutation in dihydrofolate reductase–thymidylate synthase confers resistance to pyrimethamine in falciparum malaria. Proc Natl Acad Sci USA 85:9114–9118
Randrianasolo L, Randriamanantena A, Ranarivelo L et al (2004) Monitoring susceptibility to sulfadoxine–pyrimethamine among cases of uncomplicated, Plasmodium falciparum malaria in Saharevo, Madagascar. Ann Trop Med Parasitol 98:551–554
Rastelli G, Sirawaraporn W, Sompornpisut P et al (2000) Interaction of pyrimethamine, cycloguanil, WR99210 and their analogues with Plasmodium falciparum dihydrofolate reductase: structural basis of antifolate resistance. Bioorg Med Chem 8:1117–1128
Sibley CH, Hyde JE, Sims PF et al (2001) Pyrimethamine–sulfadoxine resistance in Plasmodium falciparum: what next? Trends Parasitol 17:582–588
Sikorski RS, Hieter P (1989) A system of shuttle vectors and yeast host strains designed for efficient manipulation of DNA in Saccharomyces cerevisiae. Genetics 122:19–27
Simpson JA, Watkins E, Price RN et al (2000) Mefloquine pharmacokinetic–pharmacodynamic models: implications for dosing and resistance. Antimicrob Agents Chemother 44:3414–3424
Sina B (2002) Focus on Plasmodium vivax. Trends Parasitol 18:287–289
Sirawaraporn W, Sathitkul T, Sirawaraporn R et al (1997) Antifolate-resistant mutants of Plasmodium falciparum dihydrofolate reductase. Proc Natl Acad Sci USA 94:1124–1129
Snewin VA, England SM, Sims PF, Hyde JE (1989) Characterisation of the dihydrofolate reductase–thymidylate synthetase gene from human malaria parasites highly resistant to pyrimethamine. Gene 76:41–52
Tjitra E, Baker J, Suprianto S et al (2002) Therapeutic efficacies of artesunate–sulfadoxine–pyrimethamine and chloroquine–sulfadoxine–pyrimethamine in vivax malaria pilot studies: relationship to Plasmodium vivax dhfr mutations. Antimicrob Agents Chemother 46:3947–3953
Tokuriki N, Jackson CJ, Afriat-Jurnou L, Wyganowski KT, Tang R, Tawfik DS (2012) Diminishing returns and tradeoffs constrain the laboratory optimization of an enzyme. Nat Commun 3:1257
van den Broek IVF, van der Wardt S, Talukder L et al (2004) Drug resistance in Plasmodium falciparum from the Chittagong Hill Tracts, Bangladesh. Trop Med Int Health 9:680–687
Wang X, Minasov G, Shoichet BK (2002) Evolution of an antibiotic resistance enzyme constrained by stability and activity trade-offs. J Mol Biol 320:85–95
Weinreich DM, Delaney NF, Depristo MA, Hartl DL (2006) Darwinian evolution can follow only very few mutational paths to fitter proteins. Science 312:111–114
Wellems TE (2002) Plasmodium chloroquine resistance and the search for a replacement antimalarial drug. Science 298:124–126
White NJ, Pongtavornpinyo W, Maude RJ et al (2009) Hyperparasitaemia and low dosing are an important source of anti-malarial drug resistance. Malar J 8:253–271
Wooden JM, Hartwell LH, Vasquez B, Sibley CH (1997) Analysis in yeast of antimalaria drugs that target the dihydrofolate reductase of Plasmodium falciparum. Mol Biochem Parasitol 85:25–40
Yuvaniyama J, Chitnumsub P, Kamchonwongpaisan S et al (2003) Insights into antifolate resistance from malarial DHFR-TS structures. Nat Struct Biol 10:357–365
Acknowledgments
This research was supported by FQEB Grant #RFP-12-09 to DLH from the John Templeton Foundation program on Foundational Questions in Evolutionary Biology (FQEB). We’d also like to thank Hsiao-Han Chang for her helpful insights during data analysis and write-up.
Conflict of interest
The authors declare that they have no conflict of interest.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Jiang, PP., Corbett-Detig, R.B., Hartl, D.L. et al. Accessible Mutational Trajectories for the Evolution of Pyrimethamine Resistance in the Malaria Parasite Plasmodium vivax . J Mol Evol 77, 81–91 (2013). https://doi.org/10.1007/s00239-013-9582-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00239-013-9582-z