Abstract
Vascularized composite allotransplantation (VCA) is an evolving area of transplantation. Postoperative monitoring and immunosuppression strategies draw experience from solid organ transplantation, but VCA provides unique challenges as grafts incorporate histologically heterogenous tissues with differing degrees of antigenicity. In addition, such procedures are often life-improving rather than life-saving; therefore, minimizing the risks of immunosuppression is an important clinical priority. To this end, the identification of biomarkers to monitor the health of the transplanted tissues, assess alloimmune responses under the effects of immunosuppression, and identify episodes of rejection remain key goals. In this review we look at the general considerations of alloimmune monitoring, promising biomarkers in transplantation research, and their potential application to VCA.
Similar content being viewed by others
Introduction
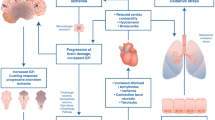
Vascularized composite allotransplantation (VCA) is an evolving field of transplantation medicine, which can improve the lives of patients with composite tissue defects including facial injuries and limb amputations. VCA involves the transplantation of a functional unit incorporating histologically heterogenous tissues with differing degrees of antigenicity, providing unique challenges to effective postoperative immunosuppression management. In addition, the procedures are often life-improving rather than life-saving and so minimizing the wide-ranging deleterious effects of immunosuppressive medication, including the increased risk of opportunistic infections, malignancies, metabolic derangement, and cardiovascular disease, is a key clinical goal in this setting. Approximately 150 different VCA operations have been completed to date including more than 85 hand and upper extremity transplants and 24 face transplant procedures. This relatively new field of transplantation has, therefore, drawn on experience of monitoring strategies from other transplant settings, particularly solid organ transplantation.
Pharmacokinetic markers remain the only routinely available tool for clinicians to monitor and adjust immunosuppressive medications within the clinic. This is despite the levels of immunosuppression not reflecting their biological effect and the absence of data from long-term clinical trials proving their validity. Over recent years, technological improvements in cellular and molecular biology have provided the techniques to identify biomarkers relevant to transplantation immunology. However, despite the potential benefits to patient care and the multiplicity of proposed candidate biomarkers, their translation into clinical practice has been very limited, and their promise still remains to be demonstrated.
The most widely used biomarker of immunological risk used to stratify patients in solid organ transplantation remains anti-HLA antibodies. This, however, requires a review paper of its own and will not be dealt with here. In this article we will first review general aspects of biomarker development and alloimmune monitoring. We will next describe a number of potential biomarkers that may aid in assessing immunological risk pre-transplantation, provide early diagnosis of rejection, and help in the identification of allograft tolerance. Finally, we will consider how these biomarkers could be relevant to the specific immunological aspects of VCA.
Biomarker Development
A biomarker is defined by the Biomarkers Definitions Working Group as “a characteristic that is objectively measured and evaluated as an indicator of normal biological processes, pathogenic processes, or pharmacological responses to a therapeutic intervention” [1]. To be effective as a clinical tool, a biomarker should ideally be capable of substituting for a clinical endpoint, i.e. to become a surrogate endpoint. This is not an easy task, as good short-term outcomes in solid organ transplantation make use of hard endpoints such as graft loss or patient survival highly impractical, due to the very large cohorts of patients that would be required to provide statistical significance. Currently, serum markers (liver function tests or serum creatinine) or pathological changes in tissue biopsy specimens are used as surrogate endpoints, but have the obvious disadvantage of identifying graft damage once it has already occurred rather than predicting potential damage [2]. In addition, they do not provide an overall indication of the individual patients’ immune response and do not always predict long-term outcomes.
In order to maximize the probability of identifying suitable biomarkers, high-throughput technologies are often used with a specific sample type such as blood, urine, or allograft tissue to test a number of contenders against a predefined gold standard. Once a biomarker has been identified, its accuracy and reproducibility in predicting a clinical outcome needs to be tested within clinical trials. Prospective studies should then be employed to show that the biomarker enables intervention to improve the clinical outcome in a real-life population of patients in which it will be used.
Employing Biomarkers to Assess Alloimmune Responses
The central role of T cells in transplant rejection has been well established. T cells recognize donor antigens by either the direct or indirect pathway. In the direct pathway, donor peptides bound to donor major histocompatibility complexes (MHCs) on the surface of donor antigen-presenting cells (APCs) are recognized by recipient T cells. In the indirect pathway recipient T cells recognize donor peptides bound to recipient APCs. In addition, a semi-direct pathway has been identified in animal models in which recipient APCs acquire donor MHC-peptide complexes and are, therefore, capable of simultaneously priming recipient T cells via both direct and indirect allorecognition pathways [3]. The direct pathway has been shown to predominate in the first few months after transplantation [4–7] in contrast to chronic immune-mediated injury in which the indirect pathway prevails [8–10].
T cell activation requires T cell receptor complex binding to MHC-peptide complex along with co-stimulation. Over the next several days, primed, naïve T cells proliferate, synthesize cytokines, and differentiate into effector T cells or memory T cells, which can react within hours on repeat stimulation. Naïve CD4+ T Cells can commit to a number of potential cytopathic or immunoregulatory phenotypes, depending on the cytokine composition of the microenvironment in which lymphocyte activation occurs as summarized below [11–14].
-
IL-12 rich microenvironment produces Th1 IFN γ producing T cells leading to tissue destruction
-
IL-4 rich microenvironment produces Th2 IL-4 and IL-5 producing T cells leading to tissue destruction
-
In the absence of proinflammatory cytokines, TGF-β directs CD4+ T cells to commit to Foxp3+ regulatory T cells
-
IL-6 and/or IL-21 expression in the presence of TGF-β leads CD4+ T cells to a highly cytopathic Th-17 phenotype.
Current models suggest that it is the balance between cytopathic Th1 and Th17 CD4+ T cells, rather than their absolute number, that determines the clinical picture of either rejection or tolerance. Th17 and regulatory T cells have also been shown to be connected and display plasticity rather than being terminally differentiated [15]. Assessment of alloimmune responses rely on the characterization of these subpopulations of lymphocytes or measurement of the steps leading to T cell activation such as intracellular production of ATP, calcium influx, or protein phosphorylation in addition to proliferation and cytokine production.
Biomarkers of Immunological Risk Pre-transplantation
Access to donor alloantigens in the form of donor cells allows assessment of alloantigen-specific cellular immune responses. Several cellular assays have been developed to do so, such as mixed lymphocyte reaction (MLR), cytotoxic T-lymphocyte precursor (CTLp), and limiting dilution assays (LDA). In the setting of bone marrow transplantation, MLR assays have not been shown to predict graft versus host disease (GVHD) [16], whereas CTLp determination has been shown to be a good predictor. CTLp and LDAs provide a consistent measurement of effector functions such as proliferation, cytokine production, and cytotoxicity in GVHD [17, 18], but reliability in solid organ transplantation is less dependable [19–21]. Furthermore, these techniques are time-consuming, expensive, and difficult to standardize.
The development of enzyme-linked immunosorbent spot (ELISPOT) assay has allowed the quantification of cytokine producing T cells in a more reproducible and clinically applicable manner, but it is not possible to analyze simultaneously different T cell subsets and cytokines [22–24]. This limitation is overcome using multiparameter flow cytometry which can also assess cytokine production using intracellular and extracellular staining, but is not as sensitive as ELISPOT assays [25–27].
The magnitude of alloimmune responses can also be assessed employing non-alloantigen specific assays. These techniques have a number of advantages over alloantigen-specific assays in that they aim to provide a global assessment of immune competence, do not require donor tissue, and are easier to standardize, which improves the potential for translation into routine clinical care. A commercially available FDA approved assay (ImmuKnow®) quantifying whole blood CD4+ T cell intracellular adenosine triphosphate (iATP) content after phytohemagglutinin (PHA) stimulation, aims to provide a global assessment of an individual’s immune response, but remains to be validated for routine clinical use [28].
In renal transplantation, the IFN-γ ELISPOT assay has been correlated with the risk of acute cellular rejection by acting as a marker of primed donor-specific immunity even in those patients with low panel-reactive antibody (PRA) scores [24, 29, 30]. Presumed sensitization through long-term dialysis was also correlated with IFN-γ ELISPOT reactivity and the associated lower renal graft survival rates seen in these patients [31]. Using this technology, the panel-reactive T cell assay was developed which has been shown to be predict post-transplant acute rejection and 6-month renal function better than PRA by measuring responses against a panel of stimulator cells with the most common HLA antigens [32, 33].
The potential for soluble CD30 for use as a biomarker was suggested following the observation that serum sCD30 (produced following cleavage of the membrane-bound CD30 molecule on activated T cells) levels were higher in inflammatory conditions [34, 35]. However, large variations in serum levels were observed in patients awaiting renal transplantation, and further studies showed it did not identify those patients at high risk of rejection [36, 37].
Biomarkers of Immunological Risk and Graft Dysfunction
Acute cellular rejection remains a significant clinical problem in VCA, and skin rejection predominates with 85 % of patients experiencing at least one episode [38]. Lymphocyte immunophenotyping by flow cytometry has been an appealing target for identifying transplant rejection, but in other transplant settings has been unsuccessful, as increased numbers of activated and/or effector T cell subsets also occur in other causes of inflammation such as CMV infection; therefore, assessment of cell surface markers have proved unreliable as biomarkers [39–42]. Recent reports have attempted to characterize the lymphocyte subsets involved in VCA rejection. Acute skin rejection in hand transplantation is predominantly by CD3+ T cells with the majority being CD8+ in mild cases and CD4+ in severe cases. Five to ten percent of T cells are Foxp3+, but their potential role in this setting requires further clarification [43]. Immunohistochemical studies in face transplantation showed a predominantly CD4+ phenotype in facial graft, oral mucosa, and split skin grafts [44]. Antibody-mediated rejection (AMR) has not been well described in hand and face transplantation, and although approximately half of all skin biopsies show positive C4d staining, it does not correlate with cellular rejection [43, 45].
IFN-γ ELISPOT assay has also been used in the post-transplant period by measuring alloreactive memory/effector T cell responses primed by the indirect alloreactive pathway in renal transplant recipients, showing that indirect alloreactivity was associated with poorer graft function [46–48].
The use of mRNA and miRNA in both graft tissue and peripheral blood as a marker of rejection and inflammation has been widely studied. Assessing the quality and size of the immune response through the use of quantitative polymerase chain reaction (qPCR) to quantify the change in graft or leukocyte gene transcripts and therefore, the magnitude of the immune response, has proved interesting [49, 50]. In addition, this technique can be used on blood, graft tissue, and fluids. Gene expression markers in kidney graft tissue have been shown to identify acute cellular rejection through intragraft expression of cytotoxic T lymphocyte (CTL) transcripts such as granzyme B and perforin [51–53]. The differences in CTL transcripts have also been shown to predict the response of ACR to therapy, aid the assessment of clinical versus subclinical rejection, and even identify immunological and histological subgroups of rejection with different clinical outcomes [54–57]. mRNA expression signatures have been identified in liver, heart, and pancreas allografts [58–60]. miRNA profiling has also been used in the setting of renal and intestinal transplantation to identify ACR and to predict outcome in patients undergoing liver transplantation for hepatocellular carcinoma [61–63].
The use of peripheral blood is more advantageous due to the avoidance of invasive sampling techniques; CTL transcripts have been associated with rejection in renal transplant recipients and can be detected before clinical rejection occurred [64, 65]. Although commercial tests are available (e.g. Allomap Xdx®) to rule out rejection in heart transplant recipients without the need for endomyocardial biopsies, the use of peripheral blood has a number of limitations. There is a significant interlaboratory variability and the transcript levels tend to be much weaker than those seen in graft tissue. This has been shown in liver transplantation in which mRNA was used to identify operationally tolerant transplant recipients [66, 67••], and miRNA may be a more suitable tool as serum biomarkers given its increased stability [68].
Fluids which directly drain an allograft, such as urine in renal transplantation and bile in liver transplantation, have also been studied as a source of noninvasive biological material. Valid mRNA transcripts such as perforin and granzyme B have been identified in urine to predict rejection in renal allografts in addition to long-term outcome [49, 50]. Similarly, in lung transplantation, CTLA4, Foxp3, and granzyme B transcripts in bronchoalveolar lavage fluid were increased in ACR [69].
Another technique exploits the fact that transplanted tissue or organs have genomes that are distinct from the recipient’s genome. The genome transplant dynamics (GTD) approach requires genotyping of both the donor and recipient to establish a unique genetic fingerprint. Shotgun sequencing is then used to quantify the DNA signal by measuring the single nucleotide polymorphism (SNP) differences between individuals [70•]. GTD has been shown in heart transplant recipients to detect rejection episodes through a rise of donor-derived cell-free DNA from a baseline level of 1 % to levels of 3 – 4 % in rejection episodes [70•]. Graft-derived cell-free DNA (GcfDNA), using a droplet digital polymerase chain reaction technique in liver transplant recipients, has been used to confirm the minimally effective tacrolimus concentrations in the first 5 – 30 days following transplantation by acting as a marker of graft integrity [71].
Biomarkers of Over-Immunosuppression
An overriding challenge in transplantation medicine is to provide the correct balance of immunosuppression to avoid allograft rejection, but to prevent the development of complications associated with over-immunosuppression. To achieve this aim, individualization of immunosuppression based on biomarkers that provide a global assessment of adaptive and innate immune status would be of great benefit, but no consensus has been reached on what is considered adequate immunosuppression.
The ImmuKnow® assay, as previously described, was developed as a tool to provide an assessment of the level of immunosuppression [28]. Weak associations were identified with acute rejection in those patients with high iATP levels and post-transplant infections with low levels. However, later studies suggested that multiple samples with longitudinal patient assessment was required, to provide an accurate assessment, and the age of the patient had a significantly affect on iATP levels [72, 73]. In addition, in renal transplant recipients, iATP levels were shown to be low in the early post-transplant period, without consistently correlating with adverse events as a result of over immunosuppression [74]. The optimal immunosuppression levels are likely to vary with different allografts and at different time points from transplantation, and as such, relevant levels are yet to be established.
Mannose-binding lectin (MBL) intensifies the immune response when binding to tissues; therefore, low levels of plasma MBL prior to transplantation has been shown to predict improved renal graft outcomes [75], and high levels predict poor outcomes in lung transplantation [76]. High MBL levels in the post-transplant period are associated with protection against sepsis in kidney-pancreas transplantation [77].
The expression of HLA-DR on circulating monocytes is tightly regulated and shows an increase in expression in the presence of inflammatory mediators such as IFN-γ and a decrease in the presence of anti-inflammatory mediators such as IL-10, immunosuppressant medications, and Tregs [78–80]. Low-level HLA-DR expression on monocytes has been associated with preceding bacterial and fungal sepsis [81], which may be a useful marker of infection risk following induction immunosuppression with T cell-depleting agents.
Biomarkers of Tolerance
It has been observed that select transplant recipients maintain stable graft function in the absence of immunosuppression, and these individuals have been termed operationally tolerant. A challenge in transplantation has been to identify these patients to allow immunosuppression minimization or discontinuation.
The role of CD4+ CD25+ Foxp3+ regulatory T cells have attracted considerable attention in the setting of transplantation, having been identified as having a role in tolerance and prevention of autoimmune diseases [82, 83]. However, quantification in humans has proved problematic as activated T cells can also transiently express Foxp3 [84–86], and the recognition of a low CD127 expression to identify Tregs is also unreliable [87, 88]. Therefore, detection of the methylation status of the Treg-specific demethylated region (TSDR) has been shown to be the most dependable method of Treg identification, which can be assessed in whole blood or biopsy specimens [89]. Although variation in circulating Tregs has been found in transplant recipients showing graft tolerance or chronic graft damage, their role as biomarkers predicting clinical outcome has not been proven [90–95].
The relative frequency of monocytoid (mDC) and plasmacytoid (pDC) peripheral blood monocytic cell precursors have been used as a predictor of tolerance in the setting of liver transplantation, with a significantly higher pDC/mDC ratio being identified in tolerant individuals and those on minimal immunosuppression, compared to healthy controls or those on maintenance immunosuppression [96]. More recently, whole genome Affymetrix® microarrays have been employed to analyze the molecular patterns associated with the tolerance phenotype in liver transplant recipients, and γδT cell and NK cell transcripts were increased, as were the frequency of CD4+ CD25+ Foxp3+ T cells [97]. The results of blood and liver tissue transcriptional biomarker studies from the first prospective drug withdrawal trial in liver transplant recipients confirmed that PBMC NK transcriptional differences were already present before immunosuppression weaning was initiated between tolerant and non-tolerant individuals. In addition, this study identified functional differences in the liver tissue samples collected between tolerant and non-tolerant individuals before immunosuppression weaning was initiated. A side to side comparison of blood and liver tissue transcriptional profiles revealed that liver tissue biomarkers were substantially more accurate and reproducible than the blood signatures [67••].
Relevance to Vascularized Composite Allotransplantation
From the point of view of potential immunomonitoring strategies, vascularized composite tissue allotransplantation is an unusual transplantation setting in that the allograft comprises heterogeneous tissues that have very different immunogenicity and different ways to respond to alloimmune injury. The skin component of the allografts, in particular, is highly immunogenic and substantially contributes to the need for robust immunosuppressive regimens and to the high rate of acute cellular rejection (although it has been shown that when skin is transplanted with other components makes it less antigenic [98]).
The allogeneic skin, on the other hand, provides a unique opportunity to monitor in real time for early signs of rejection by means of direct inspection and/or easily obtainable skin biopsies. Despite these advantages, the diagnosis of rejection following VCA remains challenging for the following reasons. First, characteristic pathological changes of skin rejection are non-specific, and can be observed in other inflammatory dermatoses as well [99, 100]. Second, sub-clinical skin rejection can occur despite complete absence of clinical signs of rejection. Third, non-skin tissues are sometimes rejected without apparent skin involvement, requiring the performance of deep biopsies, which is impractical and associated with morbidity.
Thus, similarly to that which occurs in other transplantation settings, there is an unmet clinical need in VCA to identify biomarkers correlating with alloimmune responses, immunosuppression levels, and clinical outcomes, but biomarker research in this area remains in its infancy. The relatively small number of cases contributes to a major limitation in order to validate candidate biomarkers. There is little information on the use of immune-related biomarkers to manage VCA patients. The use of functional profiling techniques should be explored as a means to obtain specific signatures discriminating rejection from other inflammatory skin conditions. Cell-free DNA quantification could also be useful to identify graft damage in the absence of skin involvement to avoid the need to perform frequent deep-tissue biopsies. How much pre-existent anti-donor immune responses influence outcomes is not well defined, but will need to be taken into account to implement personalized immunosuppression approaches and establish if tolerance-inducing strategies are considered.
Conclusion
Advances in surgical techniques and modern immunosuppression medications have provided the basis for successful transplantation of vascular composite allografts and improved the lives of patients with composite tissue defects, including facial injuries and limb amputations. However, the risks of immunosuppression remain significant, and current postoperative monitoring relies on pharmacokinetic markers to assess the adequacy of immune suppression and non-specific serum markers as signs of rejection with the use of invasive tissue biopsies for confirmation. Modern biological molecular techniques have provided potential new biomarkers to enable monitoring of the health of tissue grafts, provide an assessment of alloimmunity, and personalized immunosuppression monitoring. Reliable, accurate, and reproducible biomarkers relevant to VCA remain an unmet clinical need, but the absence of a gold standard against which to measure their effectiveness makes this particularly challenging and larger validative prospective clinical trials will be required.
References
Papers of particular interest, published recently, have been highlighted as • Of importance •• Of major importance
Biomarkers Definitions Working, G. Biomarkers and surrogate endpoints: preferred definitions and conceptual framework. Clin Pharmacol Ther. 2001;69(3):89–95.
Hartono C, Muthukumar T, Suthanthiran M. Noninvasive diagnosis of acute rejection of renal allografts. Curr Opin Organ Transplant. 2010;15(1):35–41.
Domenig C, Sanchez-Fueyo A, Tian Y, et al. The role of immunoregulatory networks in tolerant mixed chimeras induced by a non-myeloablative irradiation free protocol. Am J Transplant. 2003;3(Supplement s5):156.
Lechler RI, Batchelor JR. Immunogenicity of retransplanted rat kidney allografts. Effect of inducing chimerism in the first recipient and quantitative studies on immunosuppression of the second recipient. J Exp Med. 1982;156(6):1835–41.
Batchelor JR, Lechler RI. Why MHC incompatible grafts induce strong primary alloimmunity. Transplant Proc. 1982;14(3):535–7.
Lechler RI, Batchelor JR. Restoration of immunogenicity to passenger cell-depleted kidney allografts by the addition of donor strain dendritic cells. J Exp Med. 1982;155(1):31–41.
van Besouw NM et al. The direct and indirect allogeneic presentation pathway during acute rejection after human cardiac transplantation. Clin Exp Immunol. 2005;141(3):534–40.
Baker RJ et al. Loss of direct and maintenance of indirect alloresponses in renal allograft recipients: implications for the pathogenesis of chronic allograft nephropathy. J Immunol. 2001;167(12):7199–206.
Vella JP et al. Indirect allorecognition of major histocompatibility complex allopeptides in human renal transplant recipients with chronic graft dysfunction. Transplantation. 1997;64(6):795–800.
Sayegh MH, Carpenter CB. Role of indirect allorecognition in allograft rejection. Int Rev Immunol. 1996;13(3):221–9.
Strom TB, Koulmanda M. Recently discovered T cell subsets cannot keep their commitments. J Am Soc Nephrol. 2009;20(8):1677–80.
Korn T et al. IL-21 initiates an alternative pathway to induce proinflammatory T(H)17 cells. Nature. 2007;448(7152):484–7.
Bettelli E et al. Reciprocal developmental pathways for the generation of pathogenic effector TH17 and regulatory T cells. Nature. 2006;441(7090):235–8.
Weaver CT, Hatton RD. Interplay between the TH17 and TReg cell lineages: a (co-)evolutionary perspective. Nat Rev Immunol. 2009;9(12):883–9.
Mitchell P et al. The T helper 17-regulatory T cell axis in transplant rejection and tolerance. Curr Opin Organ Transplant. 2009;14(4):326–31.
Segall M et al. Lack of correlation of MLC reactivity with acute graft-versus-host disease and mortality in unrelated donor bone marrow transplantation. Hum Immunol. 1996;49(1):49–55.
Hernandez-Fuentes MP, Salama A. In vitro assays for immune monitoring in transplantation. Methods Mol Biol. 2006;333:269–90.
van Besouw NM et al. The granzyme B and interferon-gamma enzyme-linked immunospot assay as alternatives for cytotoxic T-lymphocyte precursor frequency after renal transplantation. Transplantation. 2005;79(9):1062–6.
Kreijveld E et al. Immunological monitoring of renal transplant recipients to predict acute allograft rejection following the discontinuation of tacrolimus. PLoS One. 2008;3(7):e2711.
van der Mast BJ et al. Calcineurin inhibitor withdrawal in stable kidney transplant patients decreases the donor-specific cytotoxic T lymphocyte precursor frequency. Transplantation. 2005;80(9):1220–5.
Steinmann J et al. Failure of in vitro T-cell assays to predict clinical outcome after human kidney transplantation. J Clin Lab Anal. 1994;8(3):157–62.
van Besouw NM et al. After discontinuation of calcineurin inhibitors, tapering of mycophenolate mofetil further impairs donor-directed cytotoxicity. Clin Transplant. 2008;22(2):129–35.
Gebauer BS et al. Evolution of the enzyme-linked immunosorbent spot assay for post-transplant alloreactivity as a potentially useful immune monitoring tool. Am J Transplant. 2002;2(9):857–66.
Nather BJ et al. Modified ELISPOT technique--highly significant inverse correlation of post-Tx donor-reactive IFNgamma-producing cell frequencies with 6 and 12 months graft function in kidney transplant recipients. Transpl Immunol. 2006;16(3–4):232–7.
Heeger PS et al. Pretransplant frequency of donor-specific, IFN-gamma-producing lymphocytes is a manifestation of immunologic memory and correlates with the risk of posttransplant rejection episodes. J Immunol. 1999;163(4):2267–75.
Volk HD, Kern F. Insights into the specificity and function of (allo)antigen-reactive T cells. Am J Transplant. 2001;1(2):109–14.
Couzi L et al. Immunological monitoring of calcineurin inhibitors for predicting cytomegalovirus infection in kidney transplant recipients. Transplantation. 2008;86(8):1060–7.
Kowalski RJ et al. Assessing relative risks of infection and rejection: a meta-analysis using an immune function assay. Transplantation. 2006;82(5):663–8.
Hricik DE et al. Enzyme linked immunosorbent spot (ELISPOT) assay for interferon-gamma independently predicts renal function in kidney transplant recipients. Am J Transplant. 2003;3(7):878–84.
Nickel P et al. Enzyme-linked immunosorbent spot assay for donor-reactive interferon-gamma-producing cells identifies T-cell presensitization and correlates with graft function at 6 and 12 months in renal-transplant recipients. Transplantation. 2004;78(11):1640–6.
Augustine JJ et al. Hemodialysis vintage, black ethnicity, and pretransplantation antidonor cellular immunity in kidney transplant recipients. J Am Soc Nephrol. 2007;18(5):1602–6.
Andree H et al. Identification of dialysis patients with panel-reactive memory T cells before kidney transplantation using an allogeneic cell bank. J Am Soc Nephrol. 2006;17(2):573–80.
Poggio ED et al. Panel of reactive T cells as a measurement of primed cellular alloimmunity in kidney transplant candidates. J Am Soc Nephrol. 2006;17(2):564–72.
Schlaf G et al. Soluble CD30 serum level–an adequate marker for allograft rejection of solid organs? Histol Histopathol. 2007;22(11):1269–79.
Saini D et al. Activated effector and memory T cells contribute to circulating sCD30: potential marker for islet allograft rejection. Am J Transplant. 2008;8(9):1798–808.
Platt RE et al. Soluble CD30 as a prognostic factor for outcome following renal transplantation. J Clin Pathol. 2009;62(7):662–3.
Slavcev A et al. Soluble CD30 in patients with antibody-mediated rejection of the kidney allograft. Transpl Immunol. 2007;18(1):22–7.
Petruzzo P, Dubernard JM. The international registry on hand and composite tissue allotransplantation. Clin Transpl. 2011;247–53.
Creemers P et al. Evaluation of peripheral blood CD4 and CD8 lymphocyte subsets, CD69 expression and histologic rejection grade as diagnostic markers for the presence of cardiac allograft rejection. Transpl Immunol. 2002;10(4):285–92.
Trzonkowski P et al. Homeostatic repopulation by CD28-CD8+ T cells in alemtuzumab-depleted kidney transplant recipients treated with reduced immunosuppression. Am J Transplant. 2008;8(2):338–47.
Pearl JP et al. Immunocompetent T-cells with a memory-like phenotype are the dominant cell type following antibody-mediated T-cell depletion. Am J Transplant. 2005;5(3):465–74.
Ticha O et al. Monitoring of CD38high expression in peripheral blood CD8+ lymphocytes in patients after kidney transplantation as a marker of cytomegalovirus infection. Transpl Immunol. 2010;24(1):50–6.
Hautz T et al. Histopathologic characterization of mild rejection (grade I) in skin biopsies of human hand allografts. Transpl Int. 2012;25(1):56–63.
Kanitakis J et al. Clinicopathologic monitoring of the skin and oral mucosa of the first human face allograft: report on the first eight months. Transplantation. 2006;82(12):1610–5.
Landin L et al. CD3+-mediated rejection and C4d deposition in two composite tissue (bilateral hand) allograft recipients after induction with alemtuzumab. Transplantation. 2009;87(5):776–81.
van Besouw NM et al. The frequency of interferon-gproducing cells reflects alloreactivity against minor histocompatibility antigens. Transplantation. 2003;75(8):1400–4.
Poggio ED et al. Alloreactivity in renal transplant recipients with and without chronic allograft nephropathy. J Am Soc Nephrol. 2004;15(7):1952–60.
Bestard O et al. Circulating alloreactive T cells correlate with graft function in longstanding renal transplant recipients. J Am Soc Nephrol. 2008;19(7):1419–29.
Li B et al. Noninvasive diagnosis of renal-allograft rejection by measurement of messenger RNA for perforin and granzyme B in urine. N Engl J Med. 2001;344(13):947–54.
Muthukumar T et al. Messenger RNA for FOXP3 in the urine of renal-allograft recipients. N Engl J Med. 2005;353(22):2342–51.
Strehlau J et al. Quantitative detection of immune activation transcripts as a diagnostic tool in kidney transplantation. Proc Natl Acad Sci U S A. 1997;94(2):695–700.
Pavlakis M et al. Intragraft IL-15 transcripts are increased in human renal allograft rejection. Transplantation. 1996;62(4):543–5.
Lipman ML, Stevens AC, Strom TB. Heightened intragraft CTL gene expression in acutely rejecting renal allografts. J Immunol. 1994;152(10):5120–7.
Nickel P et al. Cytotoxic effector molecule gene expression in acute renal allograft rejection: correlation with clinical outcome; histopathology and function of the allograft. Transplantation. 2001;72(6):1158–60.
Hoffmann SC et al. Functionally significant renal allograft rejection is defined by transcriptional criteria. Am J Transplant. 2005;5(3):573–81.
Mengel M et al. Scoring total inflammation is superior to the current Banff inflammation score in predicting outcome and the degree of molecular disturbance in renal allografts. Am J Transplant. 2009;9(8):1859–67.
Reeve J et al. Diagnosing rejection in renal transplants: a comparison of molecular- and histopathology-based approaches. Am J Transplant. 2009;9(8):1802–10.
Barker AK et al. Combined analysis of allograft inflammatory factor-1, interleukin-18, and Toll-like receptor expression and association with allograft rejection and coronary vasculopathy. Am Surg. 2010;76(8):872–8.
Luan FL et al. A pilot study of gene expression-based categorization of pancreas transplant biopsies. Transplantation. 2009;87(2):222–6.
Minisini R et al. Early activation of interferon-stimulated genes in human liver allografts: relationship with acute rejection and histological outcome. J Gastroenterol. 2011;46(11):1307–15.
Asaoka T et al. MicroRNA signature of intestinal acute cellular rejection in formalin-fixed paraffin-embedded mucosal biopsies. Am J Transplant. 2012;12(2):458–68.
Sui W et al. Microarray analysis of MicroRNA expression in acute rejection after renal transplantation. Transpl Immunol. 2008;19(1):81–5.
Chen HY et al. miR-203 expression predicts outcome after liver transplantation for hepatocellular carcinoma in cirrhotic liver. Med Oncol. 2012;29(3):1859–65.
Sabek O et al. Quantitative detection of T-cell activation markers by real-time PCR in renal transplant rejection and correlation with histopathologic evaluation. Transplantation. 2002;74(5):701–7.
Simon T et al. Serial peripheral blood perforin and granzyme B gene expression measurements for prediction of acute rejection in kidney graft recipients. Am J Transplant. 2003;3(9):1121–7.
Sawitzki B et al. Identification of gene markers for the prediction of allograft rejection or permanent acceptance. Am J Transplant. 2007;7(5):1091–102.
Bohne F et al. Intra-graft expression of genes involved in iron homeostasis predicts the development of operational tolerance in human liver transplantation. J Clin Invest. 2012;122(1):368–82. First prospective immunosuppression withdrawal trial in which tissue gene expression measurements accurately predicted the outcome of immunosuppresion withdrawal.
Farid WR et al. Hepatocyte-derived microRNAs as serum biomarkers of hepatic injury and rejection after liver transplantation. Liver Transpl. 2012;18(3):290–7.
Madsen CB et al. Elevated mRNA levels of CTLA-4, FoxP3, and granzyme B in BAL, but not in blood, during acute rejection of lung allografts. Transpl Immunol. 2010;24(1):26–32.
Snyder TM et al. Universal noninvasive detection of solid organ transplant rejection. Proc Natl Acad Sci U S A. 2011;108(15):6229–34. A promising noninvasive technique to diagnose rejection in difficult to access transplanted tissues & organs.
Oellerich M et al. Use of graft-derived cell-free DNA as an organ integrity biomarker to reexamine effective tacrolimus trough concentrations after liver transplantation. Ther Drug Monit. 2014;36(2):136–40.
Sommerer C et al. Pharmacodynamic monitoring of calcineurin inhibitor therapy: is there a clinical benefit? Nephrol Dial Transplant. 2009;24(1):21–7.
Hooper E et al. Establishing pediatric immune response zones using the Cylex ImmuKnow assay. Clin Transplant. 2005;19(6):834–9.
Gralla J, Huskey J, Wiseman AC. Trends in immune function assay (ImmuKnow; Cylex) results in the first year post-transplant and relationship to BK virus infection. Nephrol Dial Transplant. 2012;27(6):2565–70.
Berger SP et al. Association between mannose-binding lectin levels and graft survival in kidney transplantation. Am J Transplant. 2005;5(6):1361–6.
Carroll KE et al. High levels of mannose-binding lectin are associated with poor outcomes after lung transplantation. Transplantation. 2011;91(9):1044–9.
Manuel O et al. Meningococcal disease in a kidney transplant recipient with mannose-binding lectin deficiency. Transpl Infect Dis. 2007;9(3):214–8.
Docke WD et al. Monocyte deactivation in septic patients: restoration by IFN-gamma treatment. Nat Med. 1997;3(6):678–81.
Crockard AD et al. Methylprednisolone attenuates interferon-beta induced expression of HLA-DR on monocytes. J Neuroimmunol. 1996;70(1):29–35.
Koppelman B et al. Interleukin-10 down-regulates MHC class II alphabeta peptide complexes at the plasma membrane of monocytes by affecting arrival and recycling. Immunity. 1997;7(6):861–71.
Haveman JW et al. Low HLA-DR expression on peripheral blood monocytes predicts bacterial sepsis after liver transplantation: relation with prednisolone intake. Transpl Infect Dis. 1999;1(3):146–52.
Wood KJ, Sakaguchi S. Regulatory T cells in transplantation tolerance. Nat Rev Immunol. 2003;3(3):199–210.
Bennett CL et al. The immune dysregulation, polyendocrinopathy, enteropathy, X-linked syndrome (IPEX) is caused by mutations of FOXP3. Nat Genet. 2001;27(1):20–1.
Walker MR et al. Induction of FoxP3 and acquisition of T regulatory activity by stimulated human CD4+CD25- T cells. J Clin Invest. 2003;112(9):1437–43.
Roncador G et al. Analysis of FOXP3 protein expression in human CD4+CD25+ regulatory T cells at the single-cell level. Eur J Immunol. 2005;35(6):1681–91.
Ziegler SF. FOXP3: not just for regulatory T cells anymore. Eur J Immunol. 2007;37(1):21–3.
Seddiki N et al. Expression of interleukin (IL)-2 and IL-7 receptors discriminates between human regulatory and activated T cells. J Exp Med. 2006;203(7):1693–700.
Liu W et al. CD127 expression inversely correlates with FoxP3 and suppressive function of human CD4+ T reg cells. J Exp Med. 2006;203(7):1701–11.
Bestard O et al. Intragraft regulatory T cells in protocol biopsies retain foxp3 demethylation and are protective biomarkers for kidney graft outcome. Am J Transplant. 2011;11(10):2162–72.
Akl A et al. An investigation to assess the potential of CD25highCD4+ T cells to regulate responses to donor alloantigens in clinically stable renal transplant recipients. Transpl Int. 2008;21(1):65–73.
Louis S et al. Contrasting CD25hiCD4+T cells/FOXP3 patterns in chronic rejection and operational drug-free tolerance. Transplantation. 2006;81(3):398–407.
Braudeau C et al. Variation in numbers of CD4+CD25highFOXP3+ T cells with normal immuno-regulatory properties in long-term graft outcome. Transpl Int. 2007;20(10):845–55.
Li Y et al. Analyses of peripheral blood mononuclear cells in operational tolerance after pediatric living donor liver transplantation. Am J Transplant. 2004;4(12):2118–25.
Martinez-Llordella M et al. Multiparameter immune profiling of operational tolerance in liver transplantation. Am J Transplant. 2007;7(2):309–19.
van de Wetering J et al. Discontinuation of calcineurin inhibitors treatment allows the development of FOXP3+ regulatory T-cells in patients after kidney transplantation. Clin Transplant. 2011;25(1):40–6.
Mazariegos GV et al. Dendritic cell subset ratio in peripheral blood correlates with successful withdrawal of immunosuppression in liver transplant patients. Am J Transplant. 2003;3(6):689–96.
Martinez-Llordella M et al. Using transcriptional profiling to develop a diagnostic test of operational tolerance in liver transplant recipients. J Clin Invest. 2008;118(8):2845–57.
Lee WP et al. Relative antigenicity of components of a vascularized limb allograft. Plast Reconstr Surg. 1991;87(3):401–11.
Cendales LC et al. The Banff 2007 working classification of skin-containing composite tissue allograft pathology. Am J Transplant. 2008;8(7):1396–400.
Kanitakis J. The challenge of dermatopathological diagnosis of composite tissue allograft rejection: a review. J Cutan Pathol. 2008;35(8):738–44.
Compliance with Ethics Guidelines
Conflict of Interest
Gavin Whitehouse and Alberto Sanchez-Fueyo declare that they have no conflict of interest.
Human and Animal Rights and Informed Consent
This article does not contain any studies with human or animal subjects performed by any of the authors.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Whitehouse, G., Sanchez-Fueyo, A. Postoperative Monitoring: Biomarkers and Alloimmune Responses and Their Relevance to Vascularized Composite Allotransplantation. Curr Transpl Rep 1, 203–210 (2014). https://doi.org/10.1007/s40472-014-0022-9
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40472-014-0022-9