Abstract
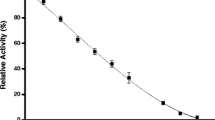
In this study, we investigated structural changes in alpha-glucosidase during urea denaturation. Alpha-glucosidase was inactivated by urea in a dose-dependent manner. The inactivation was a first-order reaction with a monophase process. Urea inhibited alpha-glucosidase in a mixed-type reaction. We found that an increase in the hydrophobic surface of this enzyme induced by urea resulted in aggregation caused by unstable folding intermediates. We also simulated the docking between alpha-glucosidase and urea. The docking simulation suggested that several residues, namely THR9, TRP14, LYS15, THR287, ALA289, ASP338, SER339, and TRP340, interact with urea. Our study provides insights into the alpha-glucosidase unfolding pathway and 3D structure of alpha-glucosidase.










Similar content being viewed by others
Abbreviations
- pNPG:
-
p-Nitrophenyl α-d-glucopyranoside
- pNP:
-
4-Nitrophenol
- SDS:
-
Sodium dodecyl sulfate
- ANS:
-
1-Anilino-8-naphthalenesulfonate
References
Charron, M. J., Dubin, R. A., & Michels, C. A. (1986). Molecular and Cellular Biology, 6, 3891–3899.
Kuriki, T., & Imanaka, T. (1999). Journal of Bioscience and Bioengineering, 87, 557–565. doi:10.1016/S1389-1723(99)80114-5.
Noguchi, A., Nakayama, T., Hemmi, H., & Nishino, T. (2003). Biochemical and Biophysical Research Communications, 304, 684–690. doi:10.1016/S0006-291X(03)00647-8.
Jespersen, H. M., MacGregor, E. A., Henrissat, B., Sierks, M. R., & Svensson, B. (1993). Journal of Protein Chemistry, 12, 791–805. doi:10.1007/BF01024938.
Godbout, A., & Chiasson, J. L. (2007). Current Diabetes Reports, 7, 333–339. doi:10.1007/s11892-007-0055-x.
Scheen, A. J. (2003). Drugs, 63, 933–951. doi:10.2165/00003495-200363100-00002.
Wehmeier, U. F., & Piepersberg, W. (2004). Applied Microbiology and Biotechnology, 63, 613–625. doi:10.1007/s00253-003-1477-2.
Moosavi-Movahedi, A. A., & Nazari, K. (1995). International Journal of Biological Macromolecules, 17, 43–47. doi:10.1016/0141-8130(95)93517-2.
Zou, Q., Habermann-Rottinghaus, S. M., & Murphy, K. P. (1998). Proteins, 31, 107–115. doi:10.1002/(SICI)1097-0134(19980501)31:2<107::AID-PROT1>3.0.CO;2-J.
Shemer, A., Nathansohn, N., Kaplan, B., Weiss, G., Newman, N., & Trau, H. (2000). International Journal of Dermatology, 39, 532–534. doi:10.1046/j.1365-4362.2000.00986-3.x.
Hagemann, I., & Proksch, E. (1996). Acta Dermato-Venereologica, 76, 353–356.
Savica, S., Tamburic, S., Savic, M., Cekic, N., Milic, J., & Vuleta, G. (2004). International Journal of Pharmaceutics, 271, 269–280. doi:10.1016/j.ijpharm.2003.11.033.
do Couto, S. G., Oliveira Mde, S., & Alonso, A. (2005). Biophysical Chemistry, 116, 23–31. doi:10.1016/j.bpc.2005.01.009.
Lodén, M., Andersson, A. C., Andersson, C., Frödin, T., Oman, H., & Lindberg, M. (2001). Skin Research and Technology, 7, 209–213. doi:10.1034/j.1600-0846.2001.070401.x.
Peck, K. D., Ghanem, A. H., & Higuchi, W. I. (1995). Journal of Pharmaceutical Sciences, 84, 975–982. doi:10.1002/jps.2600840813.
Hahn, H. S., Park, Y. D., Lee, J. R., Park, K. H., Kim, T. J., Yang, J. M., et al. (2003). Journal of Protein Chemistry, 22, 563–570. doi:10.1023/B:JOPC.0000005506.98513.43.
Zou, H. C., Yu, Z. H., Wang, Y. J., Yang, J. M., Zhou, H. M., Meng, F. G., et al. (2007). Journal of Biomolecular Structure & Dynamics, 24, 359–368.
Zou, H. C., Lü, Z. R., Wang, Y. J., Zhang, Y. M., Zou, F., & Park, Y. D. (2009). Applied Biochemistry and Biotechnology, 152, 15–28. doi:10.1007/s12010-008-8282-4.
John, B., & Sali, A. (2003). Nucleic Acids Research, 31, 3982–3992. doi:10.1093/nar/gkg460.
Rodriguez, R., Chinea, G., Lopez, N., Pons, T., & Vriend, G. (1998). Bioinformatics (Oxford, England), 14, 523–528. doi:10.1093/bioinformatics/14.6.523.
Watanabe, K., Hata, Y., Kizaki, H., Katsube, Y., & Suzuki, Y. (1997). Journal of Molecular Biology, 269, 142–153. doi:10.1006/jmbi.1997.1018.
Bryson, K., McGuffin, L. J., Marsden, R. L., Ward, J. J., Sodhi, J. S., & Jones, D. T. (2005). Nucleic Acids Research, 33, W36–W38. doi:10.1093/nar/gki410.
Morris, G. M., Goodsell, D. S., Halliday, R. S., Huey, R., Hart, W. E., Belew, R. K., et al. (1998). Journal of Computational Chemistry, 19, 1639–1662. doi:10.1002/(SICI)1096-987X(19981115)19:14<1639::AID-JCC10>3.0.CO;2-B.
Moustakas, D. T., Lang, P. T., Pegg, S., Pettersen, E., Kuntz, I. D., Brooijmans, N., et al. (2006). Journal of Computer-Aided Molecular Design, 20, 601–619. doi:10.1007/s10822-006-9060-4.
Rarey, M., Kramer, B., Lengauer, T., & Klebe, G. (1996). Journal of Molecular Biology, 261, 470–489. doi:10.1006/jmbi.1996.0477.
Moitessier, N., Englebienne, P., Lee, D., Lawandi, J., & Corbeil, C. R. (2008). British Journal of Pharmacology, 153, S7–S26. doi:10.1038/sj.bjp.0707515.
Acknowledgements
Dr. Fei Zou was supported by a grant from the National Basic Research Program of China (no. 2006CB504100). Dr. Jong Bhak was supported by a grant from the KRIBB Research Initiative Program of Korea. Dr. Yong-Doo Park was supported by fund from the Science and Technology Planning Project of Jiaxing (no. 2008AZ1024), Zhejiang.
Author information
Authors and Affiliations
Corresponding author
Additional information
Xue-Qiang Wu and Jun Wang have contributed equally to this study.
Rights and permissions
About this article
Cite this article
Wu, XQ., Wang, J., Lü, ZR. et al. Alpha-Glucosidase Folding During Urea Denaturation: Enzyme Kinetics and Computational Prediction. Appl Biochem Biotechnol 160, 1341–1355 (2010). https://doi.org/10.1007/s12010-009-8636-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12010-009-8636-6