Abstract
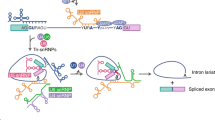
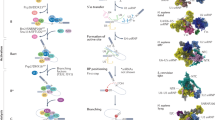
Alternative splicing is a post-transcriptional mechanism that can substantially change the pattern of gene expression. Proper regulation of alternative splicing is important for cell physiology, and aberrant splicing may lead to clinical manifestations. Cellular signals or environmental stimuli can determine the outcome of alternative splicing through trans-acting splicing regulatory factors. Networks of signaling cascades may post-translationally modify these splicing factors, thereby altering their subcellular localization or activity and hence impacting pre-mRNA splicing. Moreover, some extracellular signals, mostly steroid hormones, may regulate alternative splicing through a transcription-coupled splicing mechanism. Nevertheless, further intensive investigation will be needed to fully understand the intricacies of signal-mediated alternative splicing control.
Similar content being viewed by others
References
Black D.L. (2003) Mechanisms of alternative pre-messenger RNA splicing. Annu. Rev. Biochem. 72:291–336
Stamm S., Ben-Ari S., Rafalska I., Tang Y., Zhang Z., Toiber D., Thanaraj T.A., Soreq H. (2005) Function of alternative splicing. Gene 344:1–20
Smith C.W., Valcarcel J. (2000) Alternative pre-mRNA splicing: the logic of combinatorial control. Trends Biochem. Sci. 25: 381–388
Wilson K.F., Cerione R.A. (2000) Signal transduction and post-transcriptional gene expression. Biol. Chem. 381: 357–365
Shifrin V.I., Neel B.G. (1993) Growth factor-inducible alternative splicing of nontransmembrane phosphotyrosine phosphatase PTP-1B pre-mRNA. J. Biol. Chem. 268: 25376–25384
Stamm S. (2002) Signals and their transduction pathways regulating alternative splicing: a new dimension of the human genome. Hum. Mol. Genet. 11: 2409–2416
Shin C., Manley J.L. (2004) Cell signalling and the control of pre-mRNA splicing. Nat. Rev. Mol. Cell Biol. 5: 727–738
Srebrow A., Kornblihtt A.R. (2006) The connection between splicing and cancer. J. Cell Sci. 119:2635–2641
Auboeuf D., Honig A., Berget S.M., O’Malley B.W. (2002) Coordinate regulation of transcription and splicing by steroid receptor coregulanors. Science 298:416–419
Dowhan D.H., Hong E.P., Auboeuf D., Dennis A.P., Wilson M.M., Berget S.M., O’Malley B.W. (2005) Steroid hormone receptor coactivation and alternative RNA splicing by U2AF65-related proteins CAPERalpha and CAPERbeta. Mol. Cell 17: 429–439
Lynch K.W. (2004) Consequences of regulated pre-mRNA splicing in the immune system. Nat. Rev. Immunol. 4: 931–940
Buratti E., Baralle M., Baralle F.E. (2006) Defective splicing, disease and therapy: searching for master checkpoints in exon definition. Nucl. Acids Res. 34: 3494–3510
Matter N., Herrlich P. and Konig H. (2002) Signal-dependent regulation of splicing via phosphorylation of Sam68. Nature 420: 691–695
Matter N., Marx M., Weg-Remers S., Ponta H., Herrlich P. and Konig H. (2000) Heterogeneous ribonucleoprotein A1 is part of an exon-specific splice-silencing complex by oncogenic signaling pathways. J. Biol. Chem. 275: 35353–35360
Cheng C., Sharp P.A. (2006) Regulation of CD44 alternative splicing by SRm160 and its potential role in tumor cell invasion. Mol. Cell. Biol. 26: 362–370
Weg-Remers S., Ponta H., Herrlich P., Konig H. (2002) Antagonistic signalling pathways regulate alternative splicing of CD44 in T cells. Ann. N. Y. Acad. Sci. 973:112–115
Gui J.F., Lane W.S., Fu X.D. (1994) A serine kinase regulates intracellular localization of splicing factors in the cell cycle. Nature 369: 678–682
Colwill K., Pawson T., Andrews B., Prasad J., Manley J.L., Bell J.C., Duncan P.I. (1996) The Clk/Sty protein kinase phosphorylates SR splicing factors and regulates their intranuclear distribution. EMBO J. 15: 265–275
Rossi F., Labourier E., Forne T., Divita G., Derancourt J., Riou J.F., Antoine E., Cathala G., Brunel C., Tazi J. (1996) Specific phosphorylation of SR proteins by mammalian DNA topoisomerase I. Nature 381: 80–82
Ko T.K., Kelly E., Pines J. (2001) CrkRS: a novel conserved Cdc2-related protein kinase that colocalises with SC35 speckles. J. Cell Sci. 114: 2591–2603
Dellaire G., Makarov E.M., Cowger J.J., Longman D., Sutherland H.G., Luhrmann R., Torchia J., Bickmore W.A. (2002) Mammalian PRP4 kinase copurifies and interacts with components of both the U5 snRNP and the N-CoR deacetylase complexes. Mol. Cell. Biol. 22:5141–5156
Hu D., Mayeda A., Trembley J.H., Lahti J.M., Kidd V.J. (2002) CDK11 complexes promote pre-mRNA splicing. J. Biol. Chem. 278: 8623–8629
Xiao S.H., Manley J.L. (1998) Phosphorylation-dephosphorylation differentially affects activities of splicing factor ASF/SF2. EMBO J. 17: 6359–6367
Prasad J., Colwill K., Pawson T., Manley J.L. (1999) The protein kinase Clk/Sty directly modulates SR protein activity: both hyper- and hypophosphorylation inhibit splicing. Mol. Cell. Biol. 19:6991–7000
Kamachi M., Le T.M., Kim S.J., Geiger M.E., Anderson P., Utz P.J. (2002) Human autoimmune sera as molecular probes for the identification of an autoantigen kinase signaling pathway. J. Exp. Med. 196:1213–1225
Mikolajczyk M., Nelson M.A., Regulation of stability of cyclin-dependent kinase CDK11p110 and a caspase-processed form, CDK11p46, by Hsp90. Biochem. J. 384: 461–467, 2004.
Graveley B.R. (2000) Sorting out the complexity of SR protein functions. RNA 6:1196–1211
Patel N.A., Kaneko S., Apostolatos H.S., Bae S.S., Watson J.E., Davidowitz K., Chappell D.S., Birnbaum M.J., Cheng J.Q., Cooper D.R. (2005) Molecular and genetic studies imply Akt-mediated signaling promotes protein kinase CbetaII alternative splicing via phosphorylation of serine/arginine-rich splicing factor SRp40. J. Biol. Chem. 280:14302–14309
Chalfant C.E., Ogretmen B., Galadari S., Kroesen B.J., Pettus B.J., Hannun Y.A. (2001) FAS activation induces dephosphorylation of SR proteins: dependence on the de novo generation of ceramide and activation of protein phosphatase 1. J. Biol. Chem. 276: 44848–44855
Shin C., Manley J.L. (2004) Dephosphorylated SRp38 acts as a splicing repressor in response to heat shock. Nature 427: 553–558
Bedford M.T., Richard S. (2005) Arginine methylation an emerging regulator of protein function. Mol. Cell 18: 263–272
Nichols R.C., Wang X.W., Tang J., Hamilton B.J., High F.A., Herschman H.R. and Rigby W.F. (2000) The RGG domain in hnRNP A2 affects subcellular localization. Exp. Cell Res. 256: 522–532
Boisvert F.M., Cote J., Boulanger M.C., Cleroux P., Bachand F., Autexier C., Richard S. (2002) Symmetrical dimethylarginine methylation is required for the localization of SMN in Cajal bodies and pre-mRNA splicing. J. Cell Biol. 159: 957–969
Xu C., Henry M.F. (2004) Nuclear export of hnRNP Hrp1p and nuclear export of hnRNP Npl3p are linked and influenced by the methylation state of Npl3p. Mol. Cell. Biol. 24: 10742–10756
Cheng D., Côté J., Shaaban S., Bedford M.T. (2007) The arginine methyltransferase CARM1 regulates the coupling of transcription and mRNA processing. Mol. Cell 25:71–83
Bellare P., Kutach A.K., Rines A.K., Guthrie C., Sontheimer E.J. (2006) Ubiquitin binding by a variant Jab1/MPN domain in the essential pre-mRNA splicing factor Prp8p. RNA 12:292–302
Berro R., Kehn K., de la Fuente C., Pumfery A., Adair R., Wade J., Colberg-Poley A.M., Hiscott J., Kashanchi F. (2006) Acetylated Tat regulates human immunodeficiency virus type 1 splicing through its interaction with the splicing regulator p32. J. Virol. 80:3189–3204
Li T., Evdokimov E., Shen R.F., Chao C.C., Tekle E., Wang T., Stadtman E.R., Yang D.C., Chock P.B. (2004) Sumoylation of heterogeneous nuclear ribonucleoproteins, zinc finger proteins, and nuclear pore complex proteins: a proteomic analysis. Proc. Natl. Acad. Sci. USA 101:8551–8556
Daoud R., Mies G., Smialowska A., Olah L., Hossmann K.A., Stamm S. (2002) Ischemia induces a translocation of the splicing factor tra2-beta 1 and changes alternative splicing patterns in the brain. J. Neurosci. 22: 5889–1899
van der Houven van Oordt W., Diaz-Meco M.T., Lozano J., Krainer A.R., Moscat J., Caceres J.F. (2000) The MKK[3/6]-p38-signaling cascade alters the subcellular distribution of hnRNP A1 and modulates alternative splicing regulation. J. Cell Biol. 149:307–316
Blaustein M., Pelisch F., Tanos T., Munoz M.J., Wengier D., Quadrana L., Sanford J.R., Muschietti J.P., Kornblihtt A.R., Caceres J.F., Coso O.A., Srebrow A. (2005) Concerted regulation of nuclear and cytoplasmic activities of SR proteins by AKT. Nat. Struct. Mol. Biol. 12: 1037–1044
Edenfeld G., Volohonsky G., Krukkert K., Naffin E., Lammel U., Grimm A., Engelen D., Reuveny A., Volk T., Klambt C. (2006) The splicing factor crooked nect associates with the RNA-binding protein HOW to control glial cell maturation in Drosophila. Neuron 52:969–980
Kornblihtt A.R., de la Mata M., Fededa J.P., Munoz M.J., Nogues G. (2004) Multiple links between transcription and splicing. RNA 10: 1489–1498
Ge H., Si Y., Wolffe A.P. (1998) A novel transcriptional coactivatior, p52, functionally interacts with the essential splicing factor ASF/SF2. Mol. Cell 2:751–759
Lai M.C., Teh B.H., Tarn W.Y. (1999) A human papillomavirus E2 transcriptional activator. The interactions with cellular splicing factors and potential function in pre-mRNA processing. J. Biol. Chem. 274:11832–11841
Monsalve M., Wu Z., Adelmant G., Puigserver P., Fan M., Spiegelman B.M. (2000) Direct coupling of transcription and mRNA processing through the thermogenic coactivator PGC-1. Mol. Cell 6:307–316
Auboeuf D., Dowhan D.H., Li X., Larkin K., Ko L., Berget S.M. and O’Malley B.W. (2004) CoAA, a nuclear receptor coactivator protein at the interface of transcriptional coactivation and RNA splicing. Mol. Cell. Biol. 24:442–453
Chen H.H., Wang Y.C., Fann M.J. (2006) Identification and characterization of the CDK12/cyclin L1 complex involved in alternative splicing regulation. Mol. Cell. Biol. 26:2736–2745
Sgambato V., Minassian R., Nairn A.C., Hyman S.E. (2003) Regulation of ania-6 splice variants by distinct signaling pathways in striatal neurons. J. Neurochem. 86:153–164
Nissim-Rafinia M., Kerem B. (2002) Splicing regulation as a potential genetic modifier. Trends Genet. 18:123–127
Bracco L., Kearsey J. (2003) The relevance of alternative RNA splicing to pharmacogenomics. Trends Biotech. 21:346–353
Hagiwara M. (2005) Alternative splicing: a new drug target of the post-genome era. Biochim. Biophys. Acta. 1754:324–331
Acknowledgments
The author thanks Wen-Cheng Chang and Dr. Hung-Hsi Chen for scientific comments on the manuscript and Dr. Tim C. Taylor for editing the manuscript.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Tarn, WY. Cellular signals modulate alternative splicing. J Biomed Sci 14, 517–522 (2007). https://doi.org/10.1007/s11373-007-9161-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11373-007-9161-7