Abstract
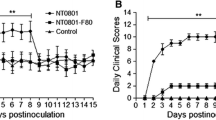
Developing a vaccine that can differentiate infected and vaccinated animals (DIVA) is a new challenge in the design of a vaccine for porcine reproductive and respiratory syndrome virus (PRRSV). Nonstructural protein 2 (nsp2) is the single largest viral product, and it has multiple roles in polypeptide processing and replication complex formation. Using reverse genetics and an infectious PRRSV cDNA clone, we constructed several deletion mutants in the non-essential region of nsp2. One mutant, which has a 131 amino acid deletion within a relatively conserved region of nsp2, was recovered and found to produce a viable virus. The deleted region was replaced with a peptide tag encoding eight amino acids. A recombinant virus containing the 131 amino acid deletion was found to produce normal virus yields in MARC-145 cells and porcine alveolar macrophages (PAM); however, gross and micro-histopathology showed that the virus was less virulent in pigs. The 131 amino acid peptide was expressed as a recombinant protein and used to coat enzyme-linked immunosorbent assay (ELISA) plates. This peptide was recognized by sera from pigs infected with wild-type virus, but not by sera from pigs infected with the deletion mutant. The results from this study show that nsp2 is an important target for the development of marker vaccines and for virus attenuation.






Similar content being viewed by others
References
E.J. Snijder, J.J. Meulenberg, J. Gen. Virol. 79, 961–979 (1998)
R. Allende, T.L. Lewis, Z. Lu, D.L. Rock, G.F. Kutish, A. Ali, A.R. Doster, F.A. Osorio, J. Gen. Virol. 80, 307–315 (1999)
C.J. Nelsen, M.P. Murtaugh, K.S. Faaberg, J. Virol. 73, 270–280 (1999)
M.H. Verheije, M.V. Kroese, P.J. Rottier, J.J. Meulenberg, J. Gen. Virol. 82, 2607–2614 (2001)
S. Wootton, D. Yoo, D. Rogan, Arch. Virol. 145, 2297–2323 (2000). doi:https://doi.org/10.1007/s007050070022
M. de Lima, A.K. Pattnaik, E.F. Flores, F.A. Osorio, Virology 353, 410–421 (2006). doi:https://doi.org/10.1016/j.virol.2006.05.036
J. Han, Y. Wang, K.S. Faaberg, Virus Res. 122, 175–182 (2006). doi:https://doi.org/10.1016/j.virusres.2006.06.003
S. Shen, J. Kwang, W. Liu, D.X. Liu, Arch. Virol. 145, 871–883 (2000). doi:https://doi.org/10.1007/s007050050680
K. Tian, X. Yu, T. Zhao, Y. Feng, Z. Cao, C. Wang, Y. Hu, X. Chen, D. Hu, X. Tian, D. Liu, S. Zhang, X. Deng, Y. Ding, L. Yang, Y. Zhang, H. Xiao, M. Qiao, B. Wang, L. Hou, X. Wang, X. Yang, L. Kang, M. Sun, P. Jin, S. Wang, Y. Kitamura, J. Yan, G.F. Gao, PLoS ONE 13, e526 (2007). doi:https://doi.org/10.1371/journal.pone.0000526
M. Yoshi, T. Okinaga, A. Miyazaki, K. Kato, H. Ikeba, H. Tsunemitsu, Arch. Virol. 153, 1323–1334 (2008). doi:https://doi.org/10.1007/s00705-008-0098-6
Y. Fang, R.R.R. Rowland, M. Roof, J.K. Lunney, J. Christopher-Hennings, E.A. Nelson, J. Virol. 80, 11447–11455 (2006). doi:https://doi.org/10.1128/JVI.01032-06
J. Han, G. Liu, Y. Wang, K.S. Faaberg, J. Virol. 81, 9878–9890 (2007). doi:https://doi.org/10.1128/JVI.00562-07
D.Y. Kim, J.G. Calvert, K.O. Chang, K. Horlen, M. Kerrigan, R.R.R. Rowland, Virus Res. 128, 106–114 (2007). doi:https://doi.org/10.1016/j.virusres.2007.04.019
A.M. Catanzariti, T.A. Soboleva, D.A. Jans, P.G. Board, R.T. Baker, Protein Sci. 13, 1331–1339 (2004). doi:https://doi.org/10.1110/ps.04618904
Y. Fang, P. Schneider, W.P. Zhang, K.S. Faaberg, E.A. Nelson, R.R. Rowland, Arch. Virol. 152, 1009–1117 (2007). doi:https://doi.org/10.1007/s00705-007-0936-y
T. Storgaard, M. Oleksiewicz, A. Botner, Arch. Virol. 144, 2389–2401 (1999). doi:https://doi.org/10.1007/s007050050652
Y.J. Choi, S.I. Yun, S.Y. Kang, Y.M. Lee, J. Virol. 80, 723–736 (2006). doi:https://doi.org/10.1128/JVI.80.2.723-736.2006
C. Lee, J.G. Calvert, S.K. Welch, D. Yoo, Virology 331, 47–62 (2005). doi:https://doi.org/10.1016/j.virol.2004.10.026
J.J. Meulenberg, J.N. Bos-de Ruijter, R. van de Graaf, G. Wensvoort, R.J. Moormann, J. Virol. 72, 380–387 (1998)
H.M. Truong, Z. Lu, G.F. Kutish, J. Galeota, F.A. Osorio, A.K. Pattnaik, Virology 325, 308–319 (2004). doi:https://doi.org/10.1016/j.virol.2004.04.046
H.S. Nielsen, G. liu, J. Nielsen, M.B. Oleksiewicz, A. Botner, T. Storgaard, K.S. Faaberg, J. Virol. 77, 3702–3711 (2003). doi:https://doi.org/10.1128/JVI.77.6.3702-3711.2003
N. Frias-Staheli, N.V. Giannakopoulos, M. Kikkert, S.L. Taylor, A. Bridgen, J. Paragas, J.A. Richt, R.R. Rowland, C.S. Schmaljohn, D.J. Lenschow, E.J. Snijder, A. García-Sastre, H.W. Virgin IV, Cell Host Microbe 13, 404–416 (2007). doi:https://doi.org/10.1016/j.chom.2007.09.014
E.J. Snijder, H. van Tol, N. Roos, K.W. Pedersen, J. Gen. Virol. 82, 985–994 (2001)
B.P. Cormack, R. Valdivia, S. Falkow, Gene 173, 33–38 (1996). doi:https://doi.org/10.1016/0378-1119(95)00685-0
M. Ormö, A.B. Cubitt, K. Kallio, L.A. Gross, R.Y. Tsien, S.J. Remington, Science 273, 1392–1395 (1996). doi:https://doi.org/10.1126/science.273.5280.1392
Acknowledgments
The project was supported by National Pork Board Project #06-135, the National Research initiative of the USDA Cooperative State Research, Education and Extension Service, grant number 2003-35204-13704.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Kim, DY., Kaiser, T.J., Horlen, K. et al. Insertion and deletion in a non-essential region of the nonstructural protein 2 (nsp2) of porcine reproductive and respiratory syndrome (PRRS) virus: effects on virulence and immunogenicity. Virus Genes 38, 118–128 (2009). https://doi.org/10.1007/s11262-008-0303-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11262-008-0303-4