Abstract
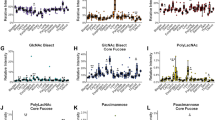
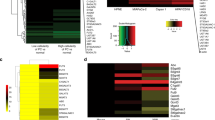
Human lung cancer is a major cause of cancer mortality worldwide. Advances in pathophysiologic understanding and novel biomarkers for diagnosis and treatment are significant tasks. We have undertaken a comprehensive glycoproteomic analysis of human lung adenocarcinoma tissues. Glycoproteins from paired lung adenocarcinoma and normal tissues were enriched by the lectins Con A, WGA, and AIL. 2-D PAGE revealed 30 differentially expressed protein spots, and 15 proteins were identified by MS/MS, including 8 up- (A1AT, ALDOA, ANXA1, CALR, ENOA, PDIA1, PSB1 and SODM) and 7 down-regulated (ANXA3, CAH2, FETUA, HBB, PRDX2, RAGE and VIME) proteins in lung cancer. By reverse-transcription PCR, nine proteins showed positive correlation between mRNA and glycoprotein expression. Vimentin and fetuin A (α2-HS-glycoprotein) were selected for further investigation. While for vimentin there was little correlation between total protein and mRNA abundance, expression of WGA-captured glycosylated vimentin protein was frequently decreased in cancer. Glycoarray analysis suggested that vimentins from normal and cancerous lung tissue differ in their contents of sialic acid and terminal GlcNAc. For fetuin A, both total protein and mRNA abundance showed concordant decrease in cancer. WGA- and AIL-binding glycosylated fetuin A was also consistently decreased in cancer. Glycoarray analysis suggested that high mannose glycan structures on fetuin A were only detectable in cancer but not normal tissue. The intriguing expression patterns of different isoforms of glycosylated vimentin and fetuin A in lung cancer illustrate the complexities and benefits of in-depth glycoproteomic analysis. In particular, the discovery of differentially glycosylated protein isoforms in lung adenocarcinoma may represent avenues towards new functional biomarkers for diagnosis, treatment guidance, and response monitoring.



Similar content being viewed by others
Abbreviations
- Con A:
-
Concanavalin A
- WGA:
-
Wheat germ agglutinin
- AIL:
-
Amylase inhibitor-like protein (jacalin)
- CALR:
-
Calreticulin
- VIME:
-
Vimentin
- FETUA:
-
Fetuin A (α2-HS-glycoprotein)
- A1AT:
-
α1-Antitrypsin
- PDIA1:
-
Protein disulfide isomerase A1
- PRDX2:
-
Peroxiredoxin-2
- ANXA1:
-
Annexin A1
- ANXA3:
-
Annexin A3
- RAGE:
-
Receptor for advanced glycosylation end products
- SODM:
-
Mitochondrial superoxide dismutase
- CAH2:
-
Carbonic anhydrase 2
- ENOA:
-
α-Enolase
- HBB:
-
Hemoglobin subunit β
- PSB1:
-
Proteasome subunit β type 1
- ALDOA:
-
Fructose-bisphosphate aldolase A
References
Pilobello KT, Mahal LK (2007) Curr Opin Chem Biol 11(3):300–305
von der Lieth CW, Lutteke T, Frank M (2006) Biochim Biophys Acta 1760(4):568–577
Zaia J (2004) Mass Spectrom Rev 23(3):161–227
Apweiler R, Hermjakob H, Sharon N (1999) Biochim Biophys Acta 1473(1):4–8
Kornfeld R, Kornfeld S (1985) Annu Rev Biochem 54:631–664
Van den Steen P, Rudd PM, Dwek RA, Opdenakker G (1998) Crit Rev Biochem Mol Biol 33(3):151–208
Helenius A, Aebi M (2001) Science 291(5512):2364–2369
Lowe JB (2001) Cell 104(6):809–812
Varki A (1993) Glycobiology 3(2):97–130
Ohtsubo K, Marth JD (2006) Cell 126(5):855–867
Bironaite D, Nesland JM, Dalen H, Risberg B, Bryne M (2000) Tumour Biol 21(3):165–175
Dennis JW, Granovsky M, Warren CE (1999) Bioessays 21(5):412–421
An HJ, Miyamoto S, Lancaster KS, Kirmiz C, Li B, Lam KS, Leiserowitz GS, Lebrilla CB (2006) J Proteome Res 5(7):1626–1635
Kirmiz C, Li B, An HJ, Clowers BH, Chew HK, Lam KS, Ferrige A, Alecio R, Borowsky AD, Sulaimon S, Lebrilla CB, Miyamoto S (2007) Mol Cell Proteomics 6(1):43–55
Lehne G (2000) Curr Drug Targets 1(1):85–99
Varki A (1999) Glycosylation changes in cancer. In: Varki A, Cummings R, Esko JD, Freeze H, Hart G, Marth JD (eds) Essentials of glycobiology. Cold Spring Harbor Laboratory Press, Cold Spring Harbor
Qiu R, Regnier FE (2005) Anal Chem 77(9):2802–2809
Debray H, Decout D, Strecker G, Spik G, Montreuil J (1981) Eur J Biochem 117(1):41–55
Goldstein IJ, Poretz RD (1986) Isolation, physicochemical characterization and carbohydrate binding specificity of lectins. In: Liener IE, Sharon N, Goldstein IJ (eds) The lectins—properties, functions and applications in biology and medicine. Academic Press, Orlando
Gu C, Oyama T, Osaki T, Li J, Takenoyama M, Izumi H, Sugio K, Kohno K, Yasumoto K (2004) Br J Cancer 90(2):436–442
Lopez-Ferrer A, Barranco C, de Bolos C (2002) Am J Clin Pathol 118(5):749–755
Rho JH, Qin S, Wang JY, Roehrl MH (2008) J Proteome Res 7(7):2959–2972
Shevchenko A, Wilm M, Vorm O, Mann M (1996) Anal Chem 68(5):850–858
Peng J, Gygi SP (2001) J Mass Spectrom 36(10):1083–1091
Edge AS, Spiro RG (1987) J Biol Chem 262(33):16135–16141
Hayase T, Rice KG, Dziegielewska KM, Kuhlenschmidt M, Reilly T, Lee YC (1992) Biochemistry 31(20):4915–4921
Kolarich D, Weber A, Turecek PL, Schwarz HP, Altmann F (2006) Proteomics 6(11):3369–3380
Kueper T, Grune T, Prahl S, Lenz H, Welge V, Biernoth T, Vogt Y, Muhr GM, Gaemlich A, Jung T, Boemke G, Elsasser HP, Wittern KP, Wenck H, Stab F, Blatt T (2007) J Biol Chem 282(32):23427–23436
Roberts NB, Green BN, Morris M (1997) Clin Chem 43(5):771–778
Srikrishna G, Huttunen HJ, Johansson L, Weigle B, Yamaguchi Y, Rauvala H, Freeze HH (2002) J Neurochem 80(6):998–1008
Julenius K, Molgaard A, Gupta R, Brunak S (2005) Glycobiology 15(2):153–164
Stevens VJ, Vlassara H, Abati A, Cerami A (1977) J Biol Chem 252(9):2998–3002
Hong SH, Misek DE, Wang H, Puravs E, Giordano TJ, Greenson JK, Brenner DE, Simeone DM, Logsdon CD, Hanash SM (2004) Cancer Res 64(15):5504–5510
Steinert PM, Roop DR (1988) Annu Rev Biochem 57:593–625
Gilles C, Polette M, Mestdagt M, Nawrocki-Raby B, Ruggeri P, Birembaut P, Foidart JM (2003) Cancer Res 63(10):2658–2664
Thiery JP (2002) Nat Rev Cancer 2(6):442–454
Ngan CY, Yamamoto H, Seshimo I, Tsujino T, Man-i M, Ikeda JI, Konishi K, Takemasa I, Ikeda M, Sekimoto M, Matsuura N, Monden M (2007) Br J Cancer 96(6):986–992
Swallow CJ, Partridge EA, Macmillan JC, Tajirian T, DiGuglielmo GM, Hay K, Szweras M, Jahnen-Dechent W, Wrana JL, Redston M, Gallinger S, Dennis JW (2004) Cancer Res 64(18):6402–6409
Matsuzaki K, Okazaki K (2006) J Gastroenterol 41(4):295–303
Watzlawick H, Walsh MT, Yoshioka Y, Schmid K, Brossmer R (1992) Biochemistry 31(48):12198–12203
Lang GA, Yeaman GR (2001) J Autoimmun 16(2):151–161
Carrell RW, Jeppsson JO, Laurell CB, Brennan SO, Owen MC, Vaughan L, Boswell DR (1982) Nature 298(5872):329–334
Vaughan L, Lorier MA, Carrell RW (1982) Biochim Biophys Acta 701(3):339–345
Goulet F, Moore KG, Sartorelli AC (1992) Biochem Biophys Res Commun 188(2):554–558
Wallner BP, Mattaliano RJ, Hession C, Cate RL, Tizard R, Sinclair LK, Foeller C, Chow EP, Browing JL, Ramachandran KL et al (1986) Nature 320(6057):77–81
Brichory FM, Misek DE, Yim AM, Krause MC, Giordano TJ, Beer DG, Hanash SM (2001) Proc Natl Acad Sci USA 98(17):9824–9829
Acknowledgments
We acknowledge financial support from the NIAID/NIH. We thank Drs. Bin Ye, Brian Liu, and Sam Mok of Brigham and Women’s Hospital for use of proteomic equipment. We thank the Taplin Biological Mass Spectrometry Center of Harvard Medical School for expert advice.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Supplementary Table 1
Detailed information on the MS/MS sequencing of the identified protein spots. (DOCX 68 kb)
Supplementary Fig. 1
2-D PAGE profiles of lectin-captured proteins from patient 4. (PDF 382 kb)
Supplementary Fig. 2
Magnified paired views of the 16 protein spots (arrowheads) from representative patient samples sequenced by MS/MS (N, normal lung; C, cancer). Spots 13 and 22 yielded the same protein identity (RAGE). (PDF 430 kb)
Supplementary Fig. 3
Comparison of mRNA expression levels between paired normal and lung adenocarcinoma tissues for 11 selected glycoproteins. RT-PCR analysis was performed on matched normal/tumor pairs from 7 patients (N/D, not detected). Data for the remaining 3 glycoproteins are shown in Fig. 2. RPL13A (ribosomal protein L13A) was used as a control transcript. The tumor to normal (T:N) ratio of each sample was normalized relative to the corresponding control transcript ratio. Amplification of ANXA3 (Fig. 1, spot 9) mRNA was not successful despite attempts with two sets of PCR primers. The average mRNA ratios (T:N) are shown in Table 1. (PDF 263 kb)
Supplementary Fig. 4
Confirmation of glycosylation and comparison of total protein abundance of vimentin and fetuin A. Presence of N-glycosylation of vimentin (A) and fetuin A (B) was demonstrated by PNGase F digestion. Unfractionated total soluble proteins were digested with PNGase F ((-), protein samples before PNGase F treatment; P, PNGase F-treated samples). Vimentin showed several protein bands. The most dominant band was selected for mobility shift calculation. Fetuin A showed two bands, both of which were shifted after deglycosylation to a similar extent. The calculated molecular weights (in kDa) of the vimentin bands are: A, 50.6; A’, 49.2; B, 50.6; B’, 49.7; C, 50.7; C’, 49.0; D, 50.6; D’, 49.5. The calculated molecular weights (in kDa) of the fetuin A bands are: E, 48.5; E’, 46.0; e, 44.1; e’, 42.3; F, 48.2; F’, 46.0; G, 48.6; G’, 46.3; H, 48.6; H’, 46.3. (C) Western blot comparison of total vimentin and fetuin A proteins. Unfractionated lung tissue protein extracts were used. The total intensities of all isoforms were used for comparison. Normalized expression ratios (T:N) are shown below the protein bands using (-actin the as an internal control. Ratios of >1, 1, or <1 describe increased, unchanged, or decreased expression in cancer, respectively. (PDF 300 kb)
Supplementary Fig. 5
Representative lectin glycoarray images. The fingerprint patterns of glycosylated vimentin (top) or fetuin A (bottom) isolated from cancer tissue protein extracts are shown. (PDF 236 kb)
Rights and permissions
About this article
Cite this article
Rho, JH., Roehrl, M.H.A. & Wang, J.Y. Glycoproteomic Analysis of Human Lung Adenocarcinomas Using Glycoarrays and Tandem Mass Spectrometry: Differential Expression and Glycosylation Patterns of Vimentin and Fetuin A Isoforms. Protein J 28, 148–160 (2009). https://doi.org/10.1007/s10930-009-9177-0
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10930-009-9177-0