Abstract
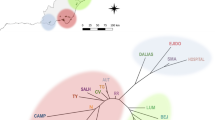
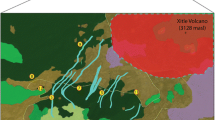
Tricholoma matsutake, a wild edible ectomycorrhizal mushroom, is revered for its distinguished flavor and iconic significance. Here, we test for landscape effects on T. matsutake gene flow and population structure in the Eastern Himalayas. Using single-nucleotide polymorphic (SNP) DNA markers, isolation by distance patterns were tested on eight populations within and between watersheds. We find that high, treeless ridgelines are effective barriers to gene flow, even at distances less than 65 km, whereas populations located within watersheds are structured at greater distances. Mantel tests demonstrated a significant positive correlation between Fst and a “landscape distance” measured as the shortest distance between population pairs below treeline r = 0.574, P = 0.002, whereas strict euclidian distances do not correlate. AMOVA analysis revealed significant partitioning with 91% of the genetic variance found within populations and 7% found between watersheds, indicative of sexually recombining populations with limited gene flow between watersheds. We show that landscape is an important determinant of air-dispersed ectomycorrhizal species population structure in heterogeneous landscapes.


Similar content being viewed by others
References
Agapow PM, Burt A (2001) Indices of multilocus linkage disequilibrium. Mol Ecol Notes 1:101–102. doi:10.1046/j.1471-8278.2000.00014.x
Baas-Becking LGM (1934) Geobiologie; of inleiding tot de milieukunde. WP Van Stockum and Zoon NV, The Netherlands
Bergemann SE, Miller SL (2002) Size, distribution, and persistence of genets in local populations of the late-stage ectomycorrhizal basidiomycete, Russula brevipes. New Phytol 156:313–320. doi:10.1046/j.1469-8137.2002.00507.x
Bergemann SE, Douhan GW, Garbelotto M, Miller SL (2006) No evidence of population structure across three isolated subpopulations of Russula brevipes in an oak/pine woodland. New Phytol 170:177–184. doi:10.1111/j.1469-8137.2006.01654.x
Chapela IH, Garbelotto M (2004) Phylogeography and evolution in matsutake and close allies inferred by analyses of ITS sequences and AFLPs. Mycologia 96:730–741. doi:10.2307/3762107
Gherbi H, Delaruelle C, Selosse MA, Martin F (1999) High genetic diversity in a population of the ectomycorrhizal basidiomycete Laccaria amethystina in a 150-year-old beech forest. Mol Ecol 8:2003–2013. doi:10.1046/j.1365-294x.1999.00801.x
Glenn TC, Schable NA (2005) Isolating Microsatellite DNA Loci. Methods Enzymol 395:202–222. doi:10.1016/S0076-6879(05)95013-1
Goudet J, Raymond M, de-Meeus T, Rousset F (1996) Testing differentiation in diploid populations. Genetics 144:1933–1940
Grubisha LC, Bergemann SE, Bruns TD (2007) Host islands within the California Northern Channel Islands create fine-scale genetic structure in two sympatric species of the symbiotic ectomycorrhizal fungus Rhizopogon. Mol Ecol 16:1811–1822. doi:10.1111/j.1365-294X.2007.03264.x
He J (2003) Cross-scale institutional linkages of commercial matsutake mushroom management and marketing: a preliminary study of an NTFP in Zhongdian County, Yunnan. Yunnan Science and Technology Press, China
Hewitt GM (1996) Some genetic consequences of ice ages, and their role, in divergence and speciation. Biol J Linn Soc Lond 58:247–276
Hoegberg N, Holdenrieder O, Stenlid J (1999) Population structure of the wood decay fungus Fomitopsis pinicola. Heredity 83:354–360. doi:10.1038/sj.hdy.6885970
Hosford D, Pilz D, Molina R, Amaranthus R (1997) Ecology and management of the commercially harvested american matsutake mushroom. In: [USDA general technical report No. PNW-GTR 412]. USDA, USA
James TY, Vilgalys R (2001) Abundance and diversity of Schizophyllum commune spore clouds in the Caribbean detected by selective sampling. Mol Ecol 10:471–479. doi:10.1046/j.1365-294x.2001.01224.x
Lian C, Hogetsu T, Matsushita N, Guerin-Laguette A, Suzuki K, Yamada A (2003) Development of microsatellite markers from an ectomycorrhizal fungus, Tricholoma matsutake, by an ISSR-suppression-PCR method. Mycorrhiza 13:27–31
Lian C, Narimatsu M, Nara K, Hogetsu T (2006) Tricholoma matsutake in a natural Pinus densiflora forest: correspondence between above-and below-ground genets, association with multiple host trees and alteration of existing ectomycorrhizal communities. New Phytol 171:825–836. doi:10.1111/j.1469-8137.2006.01801.x
Manel S, Schwartz MK, Luikart G, Taberlet P (2003) Landscape genetics: combining landscape ecology and population genetics. Trends Ecol Evol 18:189–197. doi:10.1016/S0169-5347(03)00008-9
Miehe G, Miehe S, Vogel J, Co S, La D (2007) Highest treeline in the northern hemisphere found in southern Tibet. Mt Res Dev 27:169–173. doi:10.1659/mrd.0792
Morkkynen T, Weissenberg KV, Pappine A (1997) Estimation of dispersal gradients of S-and P-type basidiospores of Heterobasidion Annosum. Eur J Forest Pathol 27:291–300. doi:10.1111/j.1439-0329.1997.tb01083.x
Murata H, Ohta A, Yamada A, Narimatsu M, Futamura N (2005) Genetic mosaics in the massive persisting rhizosphere colony “shiro” of the ectomycorrhizal basidiomycete Tricholoma matsutake. Mycorrhiza 15:505–512. doi:10.1007/s00572-005-0358-1
Parrent JL, Garbelotto M, Gilbert GS (2004) Population genetic structure of the polypore Datronia caperata in fragmented mangrove forests. Mycol Res 108:403–410. doi:10.1017/S0953756204009773
Peakall ROD, Smouse PE (2006) Genalex 6: genetic analysis in excel population genetic software for teaching and research. Mol Ecol Notes 6:288–295. doi:10.1111/j.1471-8286.2005.01155.x
Redhead S (1997) The pine mushroom industry in Canada and the United States: Why it Exists and Where it is Going. In: Palm MA, Chapela IH, (eds) Mycology in Sustainable Development: Expanding Concepts and Vanishing Borders, pp 11–46
Rotem J, Aust HJ (1991) The effect of ultraviolet and solar radiation and temperature on survival of fungal propagules. J Phytopathol 133:76–84. doi:10.1111/j.1439-0434.1991.tb00139.x
Rousset F (1997) Genetic differentiation and estimation of gene flow from F-statistics under isolation by distance. Genetics 145:1219–1228
Rousset F, Raymond M (1995) GENEPOP (version 1.2): population genetics software for exact tests and ecumenicism. J Hered 86:248–249
Roy M, Dubois M-P, Proffit M, Vincenot L, Desmarais E, Selosse M-A (2008) Evidence from population genetics that the ectomycorrhizal basidiomycete Laccaria amethystina is an actual multihost symbiont. Mol Ecol 17:2825–2838. doi:10.1111/j.1365-294X.2008.03790.x
Salick J, Anderson JD, Woo J, Sherman RE, Cili N, Yin XZ, Na A, Dorje S (2004) Tibetan ethnobotany and gradient analysis: menri (Medicine Mountains), Eastern Himalayas. In: The millennium ecosystem assessment. Bridging scales and epistemologies: linking local knowledge and global science in multi-scale assessments. Alexandria, Egypt
Saville BJ, Yoell H, Anderson JB (1996) Genetic exchange and recombination in populations of the root-infecting fungus Armillaria gallica. Mol Ecol 5:485–497. doi:10.1111/j.1365-294X.1996.tb00341.x
Sha T, Zhang HB, Ding HS, Li ZJ, Cheng LZ, Zhao ZW, Zhang YP (2007) Genetic diversity of Tricholoma matsutake in Yunnan Province. Chin Sci Bull 52:1212–1216. doi:10.1007/s11434-007-0194-0
Smits N, Fargues J, Rougier M, Goujet R, Itier B (1996) Effects of temperature and solar radiation interactions on the survival of quiescent conidia of the entomopathogenic hyphomycete Paecilomyces fumosoroseus (Wize) Brown and Smith. Mycopathologia 135:163–170. doi:10.1007/BF00632338
Stenlid J (1985) Population structure of Heterobasidion annosum as determined by somatic incompatibility, sexual incompatibility, and isoenzyme patterns. Can J Bot 63:2268–2273
Storfer A, Murphy MA, Evans JS, Goldberg CS, Robinson S, Spear SF, Dezzani R, Delmelle E, Vierling L, Waits LP (2007) Putting the ‘landscape’ in landscape genetics. Heredity 98:128–142. doi:10.1038/sj.hdy.6800917
Thrall PH, Burdon JJ, Bever JD (2002) Local adaptation in the Linum marginale–Melampsora lini host-pathogen interaction. Evol Int J Org Evol 56:1340–1351
White TJ, Bruns T, Lee S, Taylor J (1990) Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. PCR protocols: a guide to methods and applications 18:315–322
Wright S (1943) Isolation by distance. Genetics 28:114–138
Xu J, Guo H, Yang ZL (2007) Single nucleotide polymorphisms in the ectomycorrhizal mushroom Tricholoma matsutake. Microbiology 153:2002. doi:10.1099/mic.0.2006/005686-0
Xu J, Sha TAO, Li Y, Zhao Z, Yang Z (2008) Recombination and genetic differentiation among natural populations of the ectomycorrhizal mushroom Tricholoma matsutake from southwestern China. Mol Ecol 17:1238–1247. doi:10.1111/j.1365-294X.2007.03665.x
Ye S, Dhillon S, Ke X, Collins AR, Day INM (2001) An efficient procedure for genotyping single nucleotide polymorphisms. Nucleic Acids Res 29:e88. doi:10.1093/nar/29.17.e88
Yun W, Hall IR, Evans LA (1997) Ectomycorrhizal fungi with edible fruiting bodies. 1. Tricholoma matsutake and related fungi. Econ Bot 51:311–327
Zhang Q, Wu S (2000) Mountain geoecology and sustainable development of the Tibetan Plateau. Kluwer Academic, Norwell
Zhou Z, Miwa M, Hogetsu T (1999) Analysis of genetic structure of a Suillus grevillei population in a Larix kaempferi stand by polymorphism of inter-simple sequence repeat (ISSR). New Phytol 144:55–63. doi:10.1046/j.1469-8137.1999.00504.x
Acknowledgments
This work was supported by grants from the BP Conservation Programme, the Richard E. Schultes award from the Society for Economic Botany, and NSF Grant No. DGE-0232016 to Dr. Kenneth Y. Kaneshiro of the Center for Conservation Research and Training at the University of Hawaii at Manoa. We are grateful to Cui Yi, Cina Wongji and Cilin Qipin for their assistance in the field, and the people and municipal governments of the villages in which these studies were conducted. We are also grateful for the consideration and input of two anonymous reviewers.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Amend, A., Garbelotto, M., Fang, Z. et al. Isolation by landscape in populations of a prized edible mushroom Tricholoma matsutake . Conserv Genet 11, 795–802 (2010). https://doi.org/10.1007/s10592-009-9894-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10592-009-9894-0